Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis variant is considered likely pathogenic. Functional studies indicate this variant impacts protein function [PMID: 28319063]. This variant is expected to disrupt protein structure [PMID: 19369211].
This missense variant replaces leucine with proline at codon 35 of the PALB2 protein. Computational prediction suggests that this variant may not impact protein structure and function (internally defined REVEL score threshold <= 0.5, PMID: 27666373). Functional studies have reported that this variant impacts PALB2 function in homolgy-directed DNA repair, BRCA1 binding, PARP inhibitor and cisplatin sensitivity, DNA damage response assays (PMID: 28319063, 31586400, 31757951, 31636395, 33964450, 35853885). This variant has been reported in two unrelated individuals affected with breast cancer including co-segregation with disease from mother to daughter in one family (PMID: 28319063, 32185139). This variant has not been identified in the general population by the Genome Aggregation Database (gnomAD). Although there is a suspicion that this variant may be associated with disease, additional studies are necessary to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
The p.L35P variant (also known as c.104T>C), located in coding exon 2 of the PALB2 gene, results from a T to C substitution at nucleotide position 104. The leucine at codon 35 is replaced by proline, an amino acid with similar properties. A functional assay demonstrated that p.L35P results in defects in activation and maintenance of the G2/M cell cycle checkpoint, which is involved in regulating response to DNA damage from ionizing radiation (Simhadri S et al. Oncogene, 2019 03;38:1585-1596). Additionally, functional assays have demonstrated that this alteration disrupts homlogy directed repair activity, confers sensitivity to PARP inhibitors, reduces RAD51 foci formation in response to DNA damage, and significantly diminshes complex formation with BRCA1 and BRCA2 (Wiltshire T et al. Genet. Med., 2019 Oct; Boonen RACM et al. Nat Commun, 2019 Nov;10:5296; Foo TK et al. Oncogene, 2017 07;36:4161-4170). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Since supporting evidence is still limited at this time, the clinical significance of this alteration remains unclear.
This sequence change replaces leucine, which is neutral and non-polar, with proline, which is neutral and non-polar, at codon 35 of the PALB2 protein (p.Leu35Pro). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with breast cancer (PMID: 28319063). It has also been observed to segregate with disease in related individuals. ClinVar contains an entry for this variant (Variation ID: 657328). An algorithm developed to predict the effect of missense changes on protein structure and function (PolyPhen-2) suggests that this variant is likely to be disruptive. Experimental studies have shown that this missense change affects PALB2 function (PMID: 28319063, 30337689, 31586400, 31636395, 31757951, 33169439, 33964450). In summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic.
"This variant has been reported in ClinVar as Likely pathogenic (2 clinical laboratories) and as Uncertain significance (2 clinical laboratories) and as Uncertain Significance by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PALB2L35PPALB2L35PSomaticNCBI Gene:79728|Show additional gene information Variant OverviewPALB2, a scaffolding protein involved in DNA repair, is altered in various cancers.The PALB2 L35P mutation is likely oncogenic.Hide mutation effect description The PALB2 L35P mutation lies on the N-terminal coiled-coil motif of the PALB2 protein responsible for BRCA1 binding. Ectopic expression of the L35P mutation in 293T cells reduced endogenous BRCA1 interactions, and significantly reduced homologous recombination efficiency when overexpressed in PALB2-depleted cells. In addition, expression of the L35P mutation in PALB2-deficient cells resulted in decreased ionizing radiation-induced foci formation compared to wild type PALB2, as well as increased sensitivity to DNA-damaging agents (PMID: 28319063). JAX-CKB: PALB2 L35P lies within the DNA-binding, and BRCA1 and RAD51-interacting regions of the Palb2 protein (UniProt.org). L35P confers a loss of function to the Palb2 protein as demonstrated by a loss of Brca1 binding, Rad51 foci formation, and homology-directed DNA repair activity in cultured cells, and failure to rescue PARP inhibitor sensitivity in PALB2-null cells (PMID: 28319063, PMID: 31586400, PMID: 31757951).

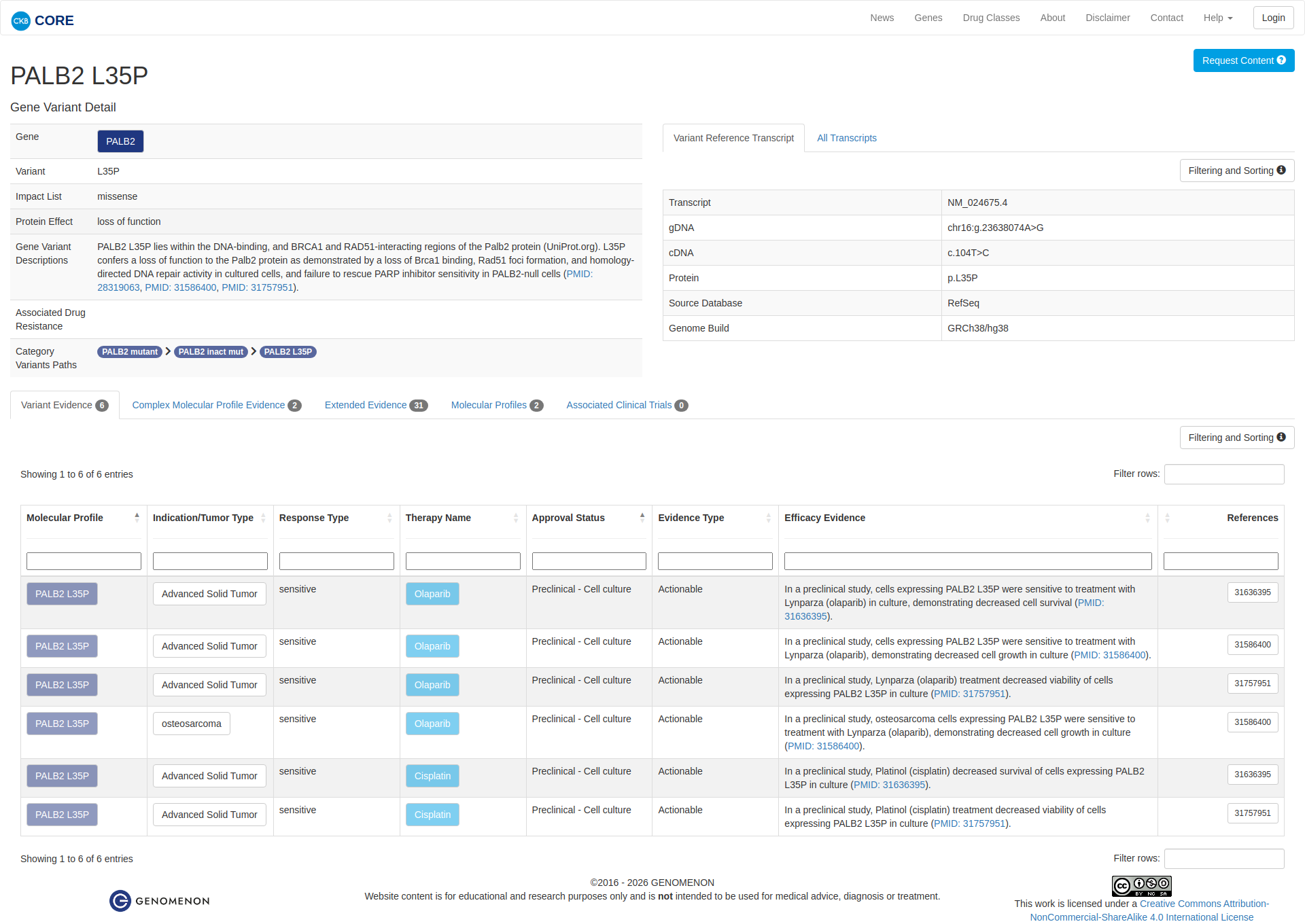
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.01 | -86 bp |
| Donor Loss (DL) | 0.0 | -4 bp |
| Acceptor Gain (AG) | 0.0 | -397 bp |
| Donor Gain (DG) | 0.0 | 399 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)