Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000059.4 | MANE Select | 11954 nt | 200–10456 |
| NM_000059.2 | Alternative | 11386 nt | 228–10484 |
| NM_000059.3 | RefSeq Select | 11386 nt | 228–10484 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe c.9117G>A pathogenic mutation (also known as p.P3039P), located in coding exon 22 of the BRCA2 gene, results from a G to A substitution at nucleotide position 9117. This nucleotide substitution does not change the proline at codon 3039. In silico splice site analysis predicts that this alteration will weaken the native splice donor site. However, this change occurs in the last base pair of coding exon 22 and has been shown to cause aberrant RNA transcripts due to exon skipping that results in a frameshift at codon 2985 and premature protein truncation (V2985Gfs*3) (Ambry internal data; Houdayer C et al. Hum. Mutat. 2012 Aug;33:1228-38; Acedo A et al. Hum. Mutat. 2015 Feb;36:210-21). This alteration has been reported in multiple families with hereditary breast and/or ovarian cancer (Hamann U et al. J. Med. Genet. 2002 Mar;39:E12; Spurdle AB et al. J. Clin. Oncol. 2008 Apr;26:1657-63; Fong PC et al. J. Clin. Oncol. 2010 May;28:2512-9; Novakovi S et al. Int. J. Oncol. 2012 Nov;41:1619-27; Takahashi M et al. Breast Cancer. 2017 Mar;24(2):336-340; Heramb C et al. Hered Cancer Clin Pract 2018 Jan;16:3; Arai M et al. J. Hum. Genet. 2018 Apr;63(4):447-457). This alteration has also been reported in a male with aggressive prostate cancer (Willems-Jones A et al. BJU Int. 2012 Dec;110:E1181-6) and in biological males with breast cancer (de Juan I et al. Fam. Cancer. 2015 Dec;14:505-13; Corman V et al. Endocr. Relat. Cancer. 2016 May;23:391-7). Of note, this mutation is also designated as 9345G>A in published literature. Based on the supporting evidence, this alteration is interpreted as a disease-causing mutation.
The BRCA2 c.9117G>A (p.Pro3039=) synonymous variant has been reported in the published literature individuals with breast cancer (PMIDs: 32658311 (2021), 29907814 (2018), 29176636 (2018), 28724667 (2017), 10638982 (2000)) and prostate cancer (PMIDs: 32853339 (2021), 28825054 (2017)). Experimental studies have shown that the variant causes the skipping of exon 23, resulting in premature termination in protein synthesis (PMID: 22505045 (2012), 22632462 (2012), 23451180 (2013), and 27060066 (2016)). The frequency of this variant in the general population, 0.000004 (1/248378 chromosomes (Genome Aggregation Database, http://gnomad.broadinstitute.org)), is consistent with pathogenicity. Analysis of this variant using software algorithms for the prediction of the effect of nucleotide changes on BRCA2 mRNA splicing yielded inconclusive findings. Based on the available information, this variant is classified as pathogenic.
This sequence change affects codon 3039 of the BRCA2 mRNA. It is a 'silent' change, meaning that it does not change the encoded amino acid sequence of the BRCA2 protein. RNA analysis indicates that this variant induces altered splicing and may result in an absent or altered protein product. The frequency data for this variant in the population databases is considered unreliable, as metrics indicate poor data quality at this position in the gnomAD database. This variant has been observed in individual(s) with breast, ovarian, or prostate cancer (PMID: 10638982, 17011978, 17148771, 22923021, 23035815, 26026974). This variant is also known as 9345G>A. ClinVar contains an entry for this variant (Variation ID: 38215). Based on a multifactorial likelihood algorithm using genetic, in silico, and/or statistical data, this variant has been determined to have a high probability of being pathogenic (PMID: 17924331, 21990134). Variants that disrupt the consensus splice site are a relatively common cause of aberrant splicing (PMID: 17576681, 9536098). Studies have shown that this variant results in skipping of exon 23, and produces a non-functional protein and/or introduces a premature termination codon (PMID: 22505045, 22632462, 23035815, 23451180, 25382762; internal data). For these reasons, this variant has been classified as Pathogenic.
Variant summary: BRCA2 c.9117G>A (p.Pro3039Pro) alters a conserved nucleotide located close to a canonical splice site and therefore could affect mRNA splicing, leading to a significantly altered protein sequence. Several computational tools predict a significant impact on normal splicing: Two predict that the variant abolishes a 5' splicing donor site. Multiple publications report experimental evidence that this variant affects mRNA splicing (e.g. Bonatti_2006, Acedo_2012, Houdayer_2012, Colombo_2013). The variant allele was found at a frequency of 4e-06 in 248378 control chromosomes. c.9117G>A has been reported in the literature in many individuals affected with Hereditary Breast And Ovarian Cancer Syndrome (e.g. Peelen_2000, Meindl_2002, Bonatti_2006, Novakovic_2012, Nakamura_2013, Corman_2016, Barrios_2017). These data indicate that the variant is very likely to be associated with disease. 17 other ClinVar submitters, including an expert panel (evaluation after 2014) cite the variant as pathogenic. Based on the evidence outlined above, the variant was classified as pathogenic.
"This variant has been reported in ClinVar as Pathogenic (32 clinical laboratories) and as pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen ENIGMA BRCA1 and BRCA2 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open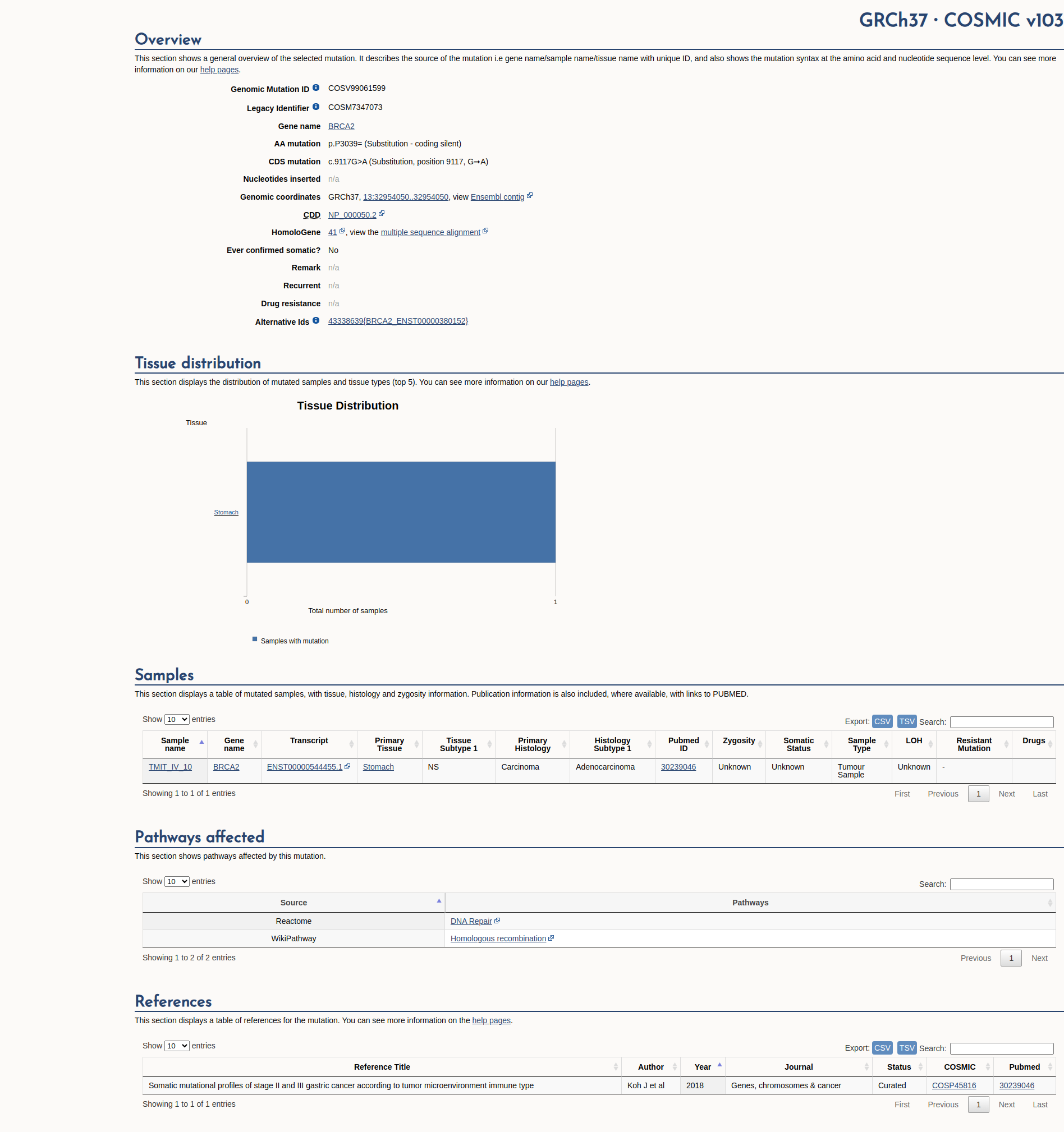
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRCA2P3039=BRCA2P3039=SomaticNCBI Gene:675|Show additional gene information Variant OverviewBRCA2, a tumor suppressor involved in the DNA damage response, is mutated in various cancer types.This is a synonymous mutation and is not annotated by OncoKB. JAX-CKB: No results found


Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.82 | -163 bp |
| Donor Loss (DL) | 0.89 | 0 bp |
| Acceptor Gain (AG) | 0.01 | 118 bp |
| Donor Gain (DG) | 0.02 | -65 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)