Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000059.4 | MANE Select | 11954 nt | 200–10456 |
| NM_000059.2 | Alternative | 11386 nt | 228–10484 |
| NM_000059.3 | RefSeq Select | 11386 nt | 228–10484 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis alteration is classified as likely benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
IARC class based on posterior probability from multifactorial likelihood analysis, thresholds for class as per Plon et al. 2008 (PMID: 18951446). Class 1 based on posterior probability = 0.000782
Variant summary: The BRCA2 c.7057G>C (p.Gly2353Arg) variant involves the alteration of a conserved nucleotide. 3/4 in silico tools predict a damaging outcome for this substitution (SNPs&GO not captured due to low reliability index). This variant was found in 8/122392 control chromosomes at a frequency of 0.0000654, which does not exceed the estimated maximal expected allele frequency of a pathogenic BRCA2 variant (0.0007503). The variant was found in HBOC spectrum patients however, without strong evidence for pathogenicity. In fact, the variant was observed in several patients to co-occur with (potentially) pathogenic BRCA1 and BRCA2 variants such as BRCA1 c.3193_3194insG (p.Asp1065?fs); BRCA2 c.4478_4481delAAAG (p.Glu1493_Ser1494?fs); BRCA2 c.476-2A>G; BRCA1 c.IVS5+1G>A (c.212+1G>A) BRCA1 c.3612delA (p.Ala1206ProfsX4), BRCA1 c.2346dupT indicating neutrality. Furthermore, Lindor_HM_2012 reviewed a multifactorial probability based model and calculated, odds in favor of causality was 0.03 and posterior probability of being deleterious 6.12104 and classified the variant as IARC Class I (Neutral) variant. Additionally, Alsop_2012 reports LOH of the variant within a breast tumor. Moreover, several clinical diagnostic laboratories classify variant as Benign/Likely benign via ClinVar. Considering all evidence, the variant was classified as Likely Benign.
"This variant has been reported in ClinVar as Likely benign (11 clinical laboratories) and as likely benign (1 clinical laboratories) and as Benign (7 clinical laboratories) and as Uncertain significance (4 clinical laboratories) and as Benign by Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) expert panel."
COSMIC Somatic Evidence
Open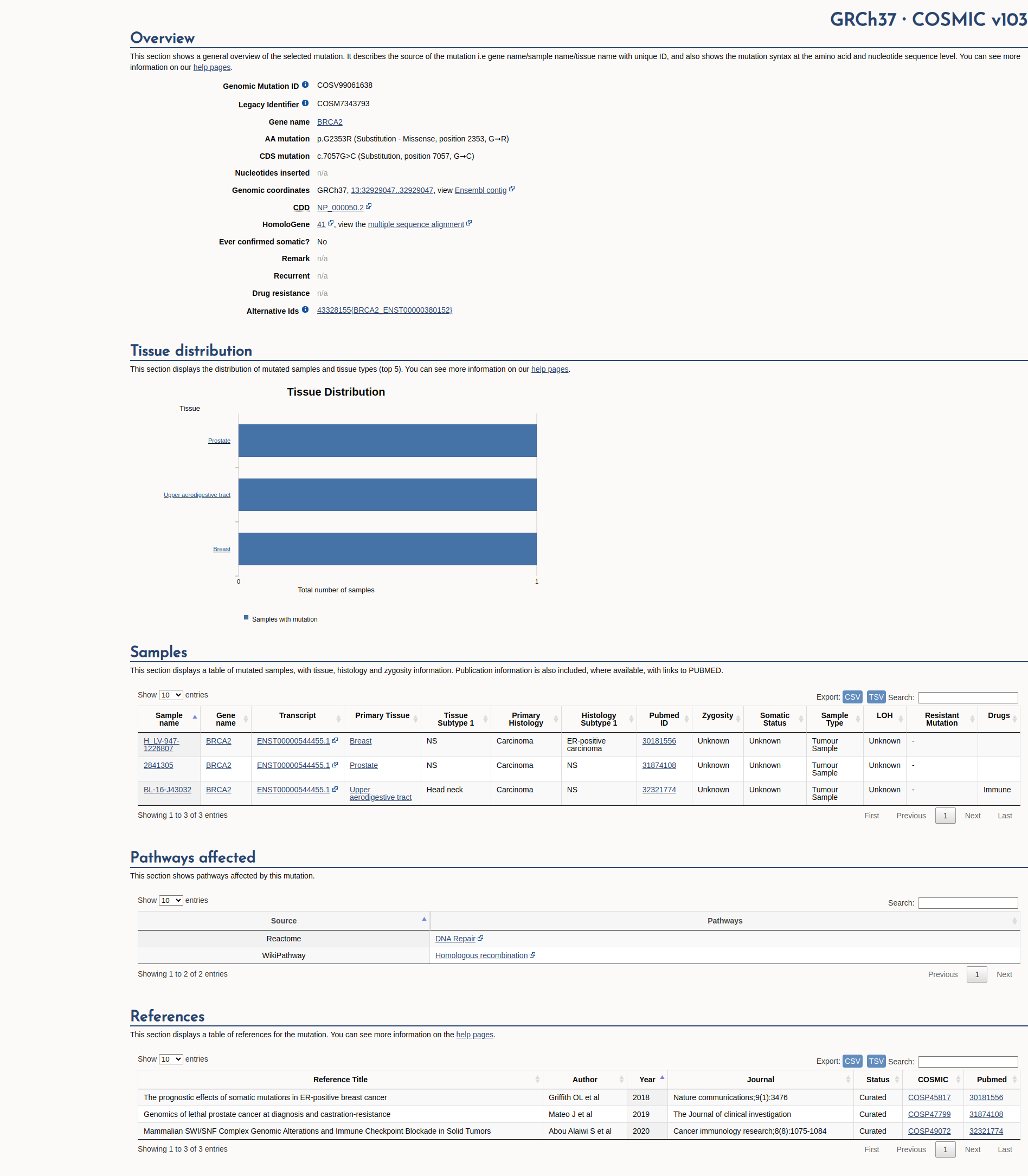
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRCA2G2353RBRCA2G2353RSomaticNCBI Gene:675|Show additional gene information Variant OverviewBRCA2, a tumor suppressor involved in the DNA damage response, is mutated in various cancer types.The BRCA2 G2353R mutation has not specifically been reviewed by the OncoKB team, and therefore its biological significance is unknown. JAX-CKB: No results found

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 197 bp |
| Donor Loss (DL) | 0.0 | 0 bp |
| Acceptor Gain (AG) | 0.01 | -49 bp |
| Donor Gain (DG) | 0.0 | 383 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)