Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_006218.2 | Alternative | 3724 nt | 158–3364 |
| NM_006218.3 | Alternative | 9104 nt | 158–3364 |
| NM_006218.4 | MANE Select | 9259 nt | 324–3530 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenFor these reasons, this variant has been classified as Pathogenic. Advanced modeling of protein sequence and biophysical properties (such as structural, functional, and spatial information, amino acid conservation, physicochemical variation, residue mobility, and thermodynamic stability) has been performed at Invitae for this missense variant, however the output from this modeling did not meet the statistical confidence thresholds required to predict the impact of this variant on PIK3CA protein function. ClinVar contains an entry for this variant (Variation ID: 31944). This missense change has been observed in individual(s) with PIK3CA-related disorders (PMID: 25599672, 26851524, 27631024). In at least one individual the variant was observed to be de novo. This variant is not present in population databases (gnomAD no frequency). This sequence change replaces glutamic acid, which is acidic and polar, with lysine, which is basic and polar, at codon 542 of the PIK3CA protein (p.Glu542Lys).
"This variant has been reported in ClinVar as Pathogenic (9 clinical laboratories) and as Likely pathogenic (2 clinical laboratories) and as Pathogenic by ClinGen Brain Malformations Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open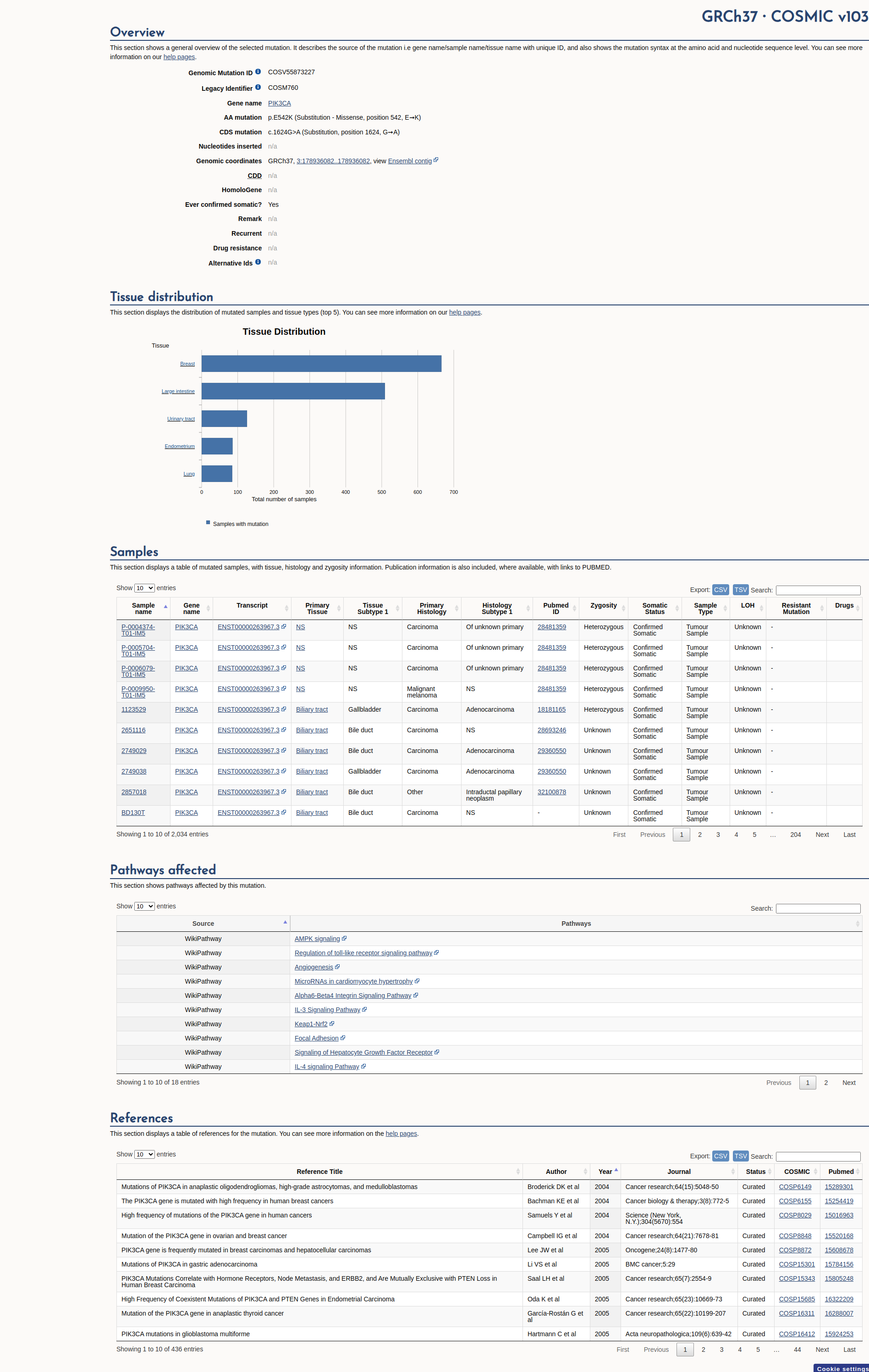
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PIK3CAE542KPIK3CAE542KSomaticNCBI Gene:5290|Show additional gene information Variant OverviewPIK3CA, the catalytic subunit of PI3-kinase, is frequently mutated in a diverse range of cancers including breast, endometrial and cervical cancers.The PIK3CA E542K mutation is known to be oncogenic.Hide mutation effect description The PIK3CA E542K mutation is located in the helical domain in exon 10 of the protein. This mutation has been found in breast and colon cancer and glioblastoma (PMID: 17376864, 31996845). Expression of this mutation in chicken embryonic fibroblasts (CEFs), Ba/F3 cells and MCF10A breast cells demonstrated that it is activating as measured by increased kinase activity, downstream pathway activation, cytokine- and factor-independent proliferation, anchorage-independent growth and tumor growth in xenograft models compared to wildtype PIK3CA (PMID: 17376864, 26627007, 16432179). Expression of this mutation in a transgenic mouse model was not sufficient to drive glioblastoma formation (PMID: 31996845). Mutations at this position are predicted to abrogate p85-mediated inhibition of catalytic activity, thus likely resulting in constitutive activation of PIK3CA enzymatic activity (PMID: 20593314). In a phase III trial for ER+, HER2− advanced breast cancer that had previously progressed on endocrine therapy, patients were given either palbociclib (CDK4/6 inhibitor) plus fulvestrant (ESR1 antagonist) or placebo plus fulvestrant. PIK3CA alterations, including the E542K mutation, were found at a significantly higher percentage in end-of-treatment samples as compared to pre-treatment samples in patients from both treatment arms, suggesting that PIK3CA alterations may play a role in resistance to fulvestrant. E542K was found more frequently than any other PIK3CA alteration in this study (PMID: 30206110). Expression of this mutation in a patient-derived xenograft model of lung squamous cell carcinoma demonstrated that it was sensitive to the pan-PIK3CA inhibitor BKM120 and the PIK3CAalpha-specific inhibitor BYL719 compared to PDX models expressing wildtype PIK3CA (PMID: 30093452). JAX-CKB: PIK3CA E542K is a hotspot mutation that lies within the PIK helical domain of the Pik3ca protein (UniProt.org). E542K results in increased PI3K activity and phosphorylation of Akt (PMID: 15647370), increased cell invasion (PMID: 26627007), and is transforming in cultured cells (PMID: 15647370, PMID: 26627007, PMID: 29533785, PMID: 17376864).


Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -76 bp |
| Donor Loss (DL) | 0.0 | 40 bp |
| Acceptor Gain (AG) | 0.0 | 15 bp |
| Donor Gain (DG) | 0.0 | 468 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)