Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open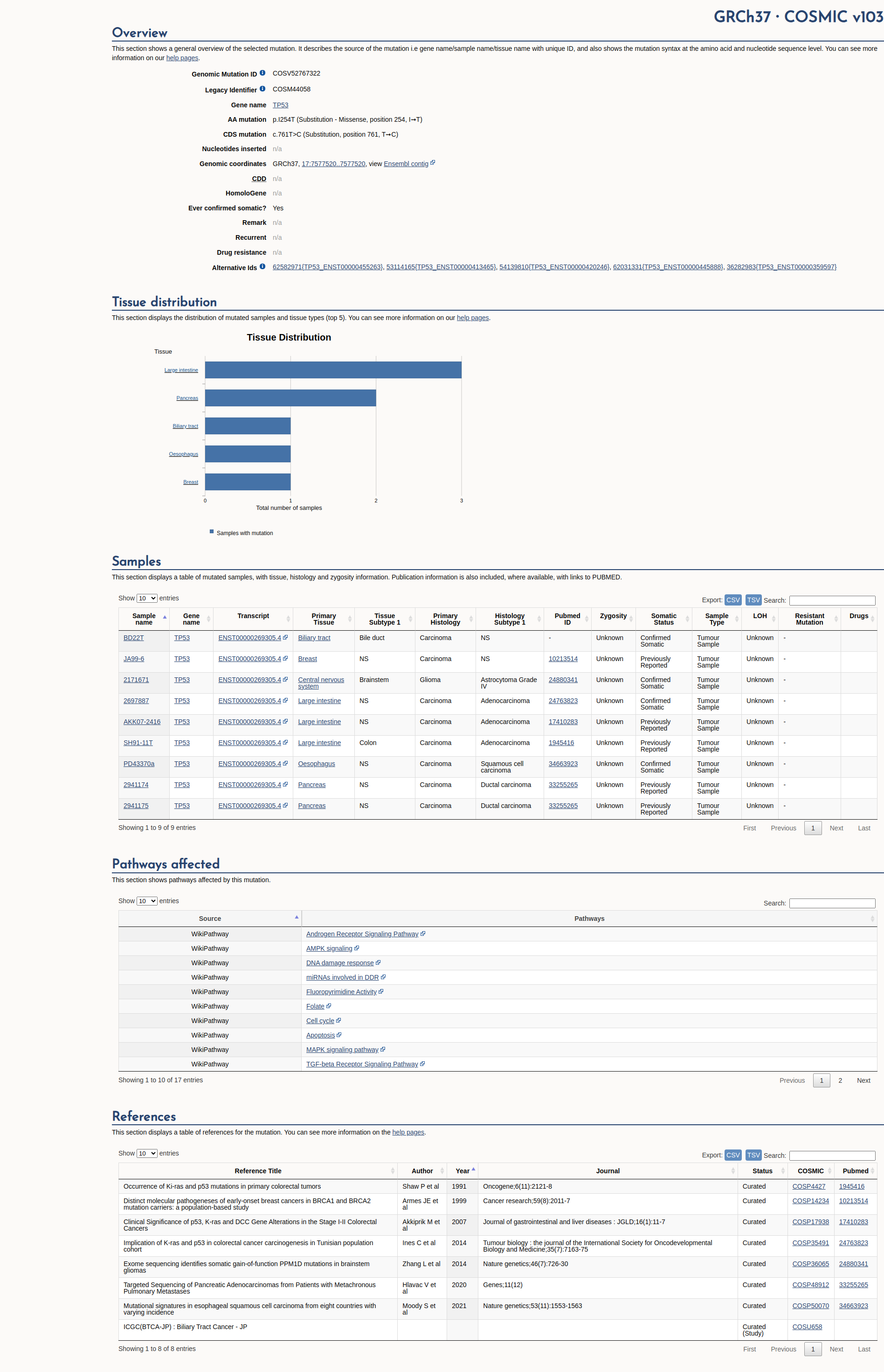
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: TP53I254TTP53I254TSomaticNCBI Gene:7157|Show additional gene information Variant OverviewTP53, a tumor suppressor in the DNA damage pathway, is the most frequently mutated gene in cancer.The TP53 I254T mutation is known to be oncogenic.Hide mutation effect description The TP53 I254T mutation is located in the protein's DNA binding domain. In vitro studies have demonstrated that this mutation is likely inactivating, as evidenced by the loss of p53-mediated transactivation and increased colony formation in the mutant compared to wildtype (PMID: 25584008). JAX-CKB: TP53 I254T lies within the DNA-binding domain of the Tp53 protein (PMID: 15510160). I254T results in decreased Tp53 transactivation activity and reduced ability to suppress colony formation in cell culture (PMID: 25584008).
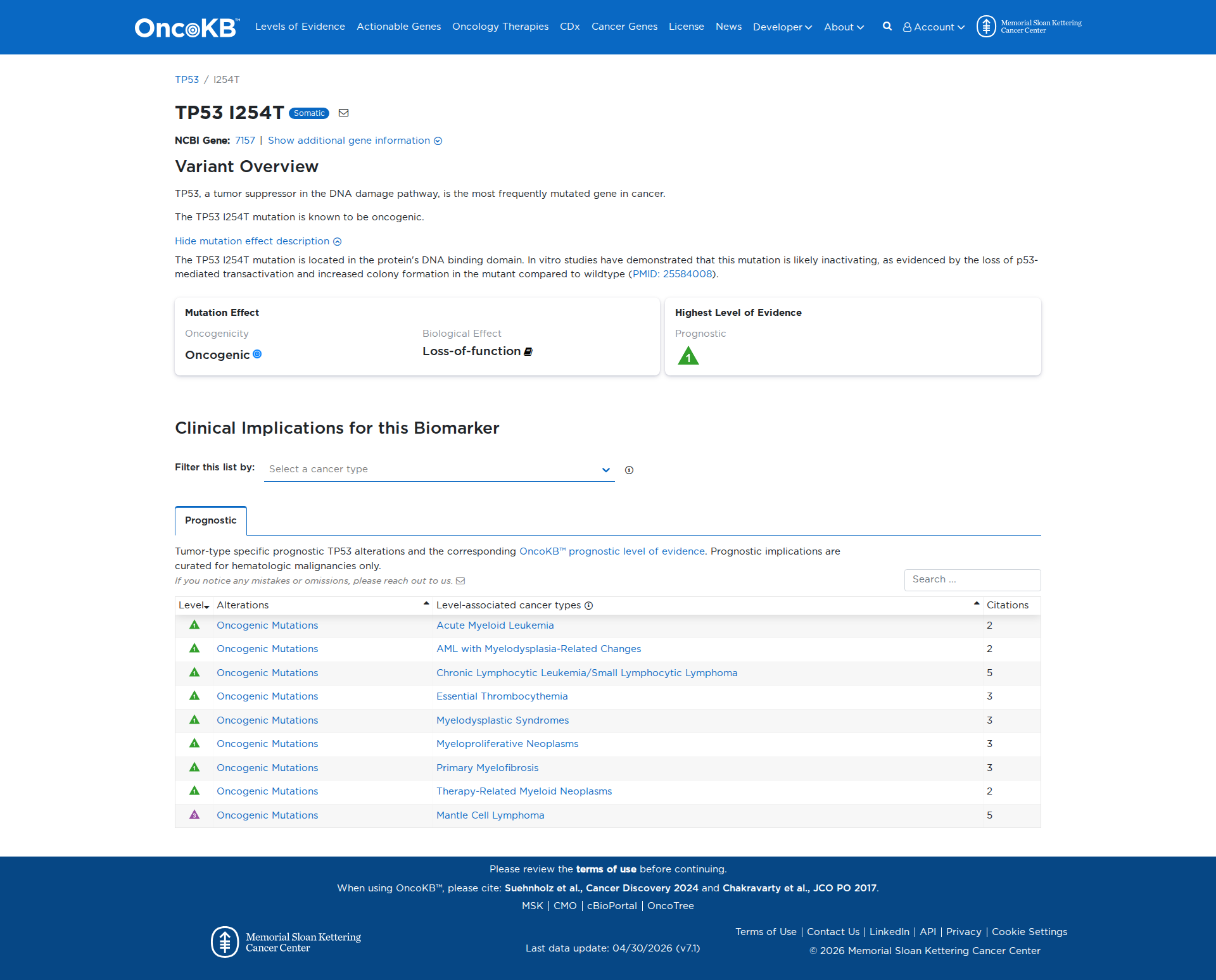
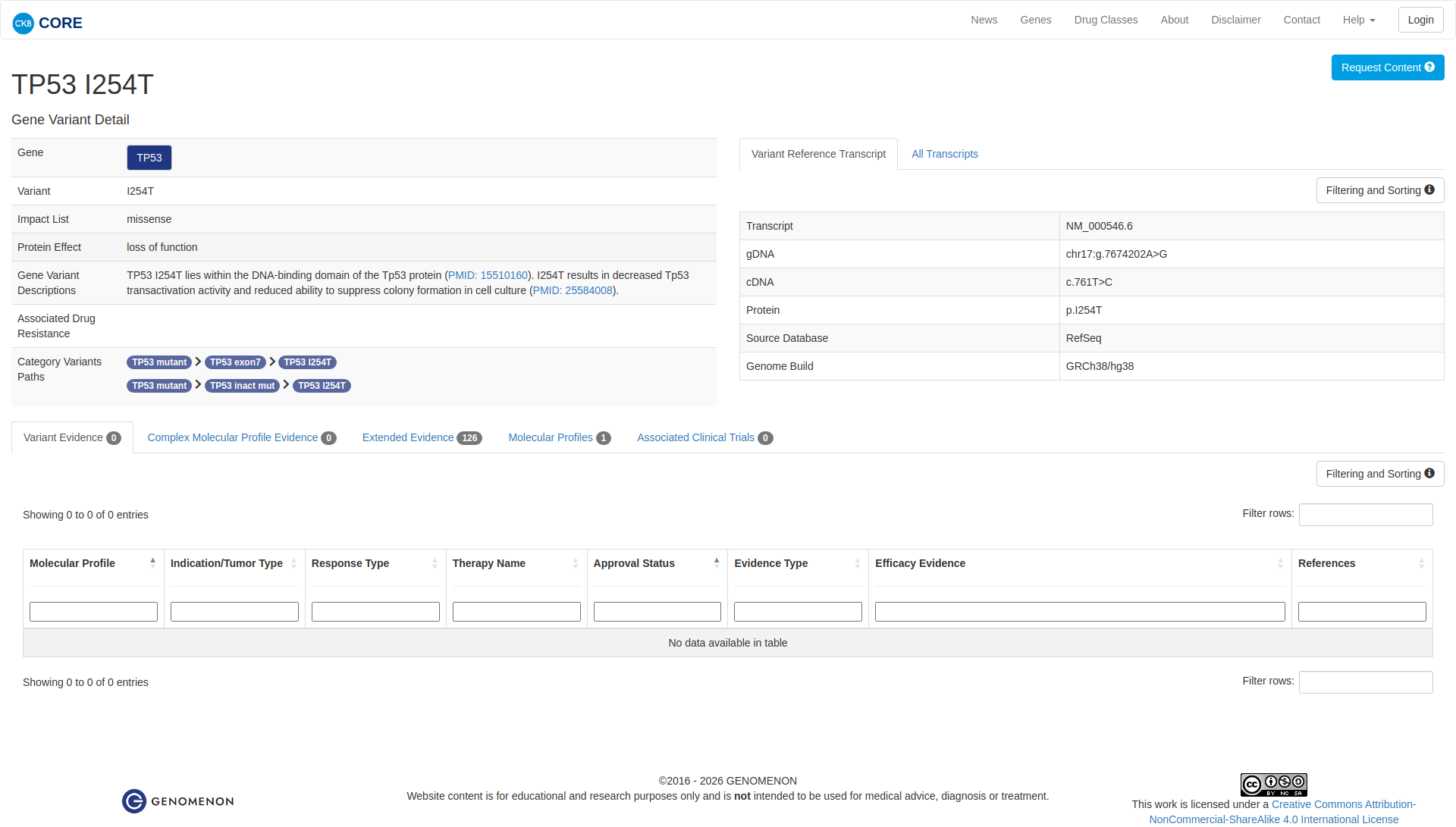
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 41 bp |
| Donor Loss (DL) | 0.0 | -21 bp |
| Acceptor Gain (AG) | 0.0 | -389 bp |
| Donor Gain (DG) | 0.0 | -311 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)