Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe p.Gly1121ValfsX3 variant in PALB2 has been reported in at least 8 individuals with breast cancer, including 2 siblings with a family history of PALB2-associated cancers (Blanco 2013 PMID: 23935836, Janatova 2013 PMID: 24136930, Castera 2014 PMID: 24549055, Antoniou 2014 PMID: 25099575, Thompson 2015 PMID: 26283626, Couch 2015 PMID: 25452441, Lee 2018 PMID: 29431189) and has also been reported by other clinical laboratories in ClinVar (Variation ID 126739). It was identified in 0.001% (1/67996) of European chromosomes by gnomAD (http://gnomad.broadinstitute.org, v.3.1). This variant is predicted to cause a frameshift, which alters the protein’s amino acid sequence beginning at position 1121 and leads to a premature termination codon 3 amino acids downstream. This alteration occurs within the last exon and is, therefore, likely to escape nonsense mediated decay (NMD) and result in a truncated protein that is missing ~5% of the coding region, with 62 amino acids removed. However, these terminal amino acids are part of the functionally critical WD40 domain that is necessary for PALB2 function, stability, and interaction with BRCA2 (Oliver 2009 PMID: 19609323). In addition, several downstream truncating variants, have been observed in individuals with PALB2-related cancers and Fanconi anemia, supporting the functional importance of the last exon. In summary, this variant meets criteria to be classified as likely pathogenic for autosomal dominant PALB2-associated breast cancer. ACMG/AMP Criteria applied: PS4_Moderate, PM2_Supporting, PVS1_Strong.
The PALB2 c.3362del (p.Gly1121Valfs*3) variant alters the translational reading frame of the PALB2 mRNA and causes the premature termination of PALB2 protein synthesis. This variant has been reported in the published literature in individuals affected with breast cancer (PMIDs: 34399810 (2021), 32885271 (2021)), or prostate cancer (PMID: 32338768 (2020)). Functional evidence suggests that this variant may impact protein function(PMID: 31636395 (2020)). The frequency of this variant in the general population, 0.000013 (1/74796 chromosomes (Genome Aggregation Database, http://gnomad.broadinstitute.org)), is consistent with pathogenicity. Based on the available information, this variant is classified as pathogenic.
This variant is predicted to result in loss of function through nonsense-mediated decay of the encoded transcript or premature truncation of the encoded protein in a gene in which loss of function is a known mechanism of disease (ACMG/AMP: PVS1; PMIDs:25099575, 33195396). Well-established functional studies have demonstrated this variant to have a damaging effect on protein function or splicing (ACMG/AMP: PS3_Supporting; PMID:33195396). This variant has been reported at an elevated frequency in affected individuals/in multiple affected individuals in the literature (ACMG/AMP: PS4_Moderate; PMIDs:23935836, 24136930, 24549055, 26283626). This variant is absent from or present at an exceedingly low frequency in gnomAD, a large-scale control population database (ACMG/AMP: PM2).
Variant summary: PALB2 c.3362delG (p.Gly1121ValfsX3) results in a premature termination codon, predicted to cause a truncation of the encoded protein, however, nonsense mediated decay is not expected to occur. The variant disrupts the Partner and localiser of BRCA2, WD40 domain which is involved in BRCA2 binding. A truncation downstream of this position has been classified as pathogenic by our laboratory (e.g. p.Tyr1183X). The variant allele was found at a frequency of 4.1e-06 in 246142 control chromosomes. c.3362delG has been reported in the literature in individuals affected with Hereditary Breast and Ovarian Cancer (e.g. Blanco_2013, Couch_2016, Janatova_2013, Thompson_2015). These data indicate that the variant is likely to be associated with disease. To our knowledge, no experimental evidence demonstrating an impact on protein function has been reported. The following publications have been ascertained in the context of this evaluation (PMID: 25452441, 23935836, 26283626, 26057125). ClinVar contains an entry for this variant (Variation ID: 126739). Based on the evidence outlined above, the variant was classified as pathogenic.
The c.3362delG pathogenic mutation, located in coding exon 13 of the PALB2 gene, results from a deletion of one nucleotide at nucleotide position 3362, causing a translational frameshift with a predicted alternate stop codon (p.G1121Vfs*3). This alteration occurs at the 3' terminus of thePALB2 gene, is not expected to trigger nonsense-mediated mRNA decay, and only impacts the last 66 amino acids of the protein. However, premature stop codons are typically deleterious in nature and the impacted region is critical for protein function (Ambry internal data). This mutation has been previously reported in numerous kindreds characterized by early-onset breast cancer, triple negative breast cancer, prostate and/or pancreatic cancer (Blanco A at al. PLoS ONE. 2013 Jul;8:e67538; Janatova M et al. Cancer Epidemiol. Biomarkers Prev. 2013 Dec;22:2323-32; Antoniou AC et al. N. Engl. J. Med. 2014 Aug;371:497-506; Castéra L et al. Eur. J. Hum. Genet. 2014 Nov;22:1305-13; Couch FJ et al. J. Clin. Oncol. 2015 Feb;33:304-11; Thompson ER et al. Breast Cancer Res. 2015 Aug;17:111; Thompson ER et al. J Clin Oncol, 2016 May;34:1455-9; Lee JEA et al. J. Pathol. 2018 May;245(1):53-60; Nguyen-Dumont T et al. Int J Cancer, 2020 10;147:2142-2149). This alteration was found to be functionally abnormal in a homology-directed DNA repair (HDR) assay (Wiltshire T et al. Genet Med, 2020 03;22:622-632). Based on the supporting evidence, this alteration is interpreted as a disease-causing mutation.
"This variant has been reported in ClinVar as Pathogenic (13 clinical laboratories) and as Likely Pathogenic (1 clinical laboratories) and as Likely pathogenic (2 clinical laboratories) and as pathogenic (1 clinical laboratories) and as Likely Pathogenic by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PALB2G1121Vfs*3PALB2G1121Vfs*3SomaticNCBI Gene:79728|Show additional gene information Variant OverviewPALB2, a scaffolding protein involved in DNA repair, is altered in various cancers.The PALB2 G1121Vfs*3 is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in PALB2 is: PALB2 truncating mutations can form several forms of C-terminally truncated PALB2 protein and are found in breast cancers. As such, PALB2 is considered a rare breast cancer susceptibility gene. In patient studies, PALB2 truncation mutations were shown to significantly increase the risk of breast cancer, with similar risks for estrogen receptor-positive and -negative breast cancer. PALB2 truncation mutations have also been found in cases of hereditary male breast cancer (PMID: 28858227, 18053174, 25529982, 29484706, 28279176, 28779002). JAX-CKB: No results found
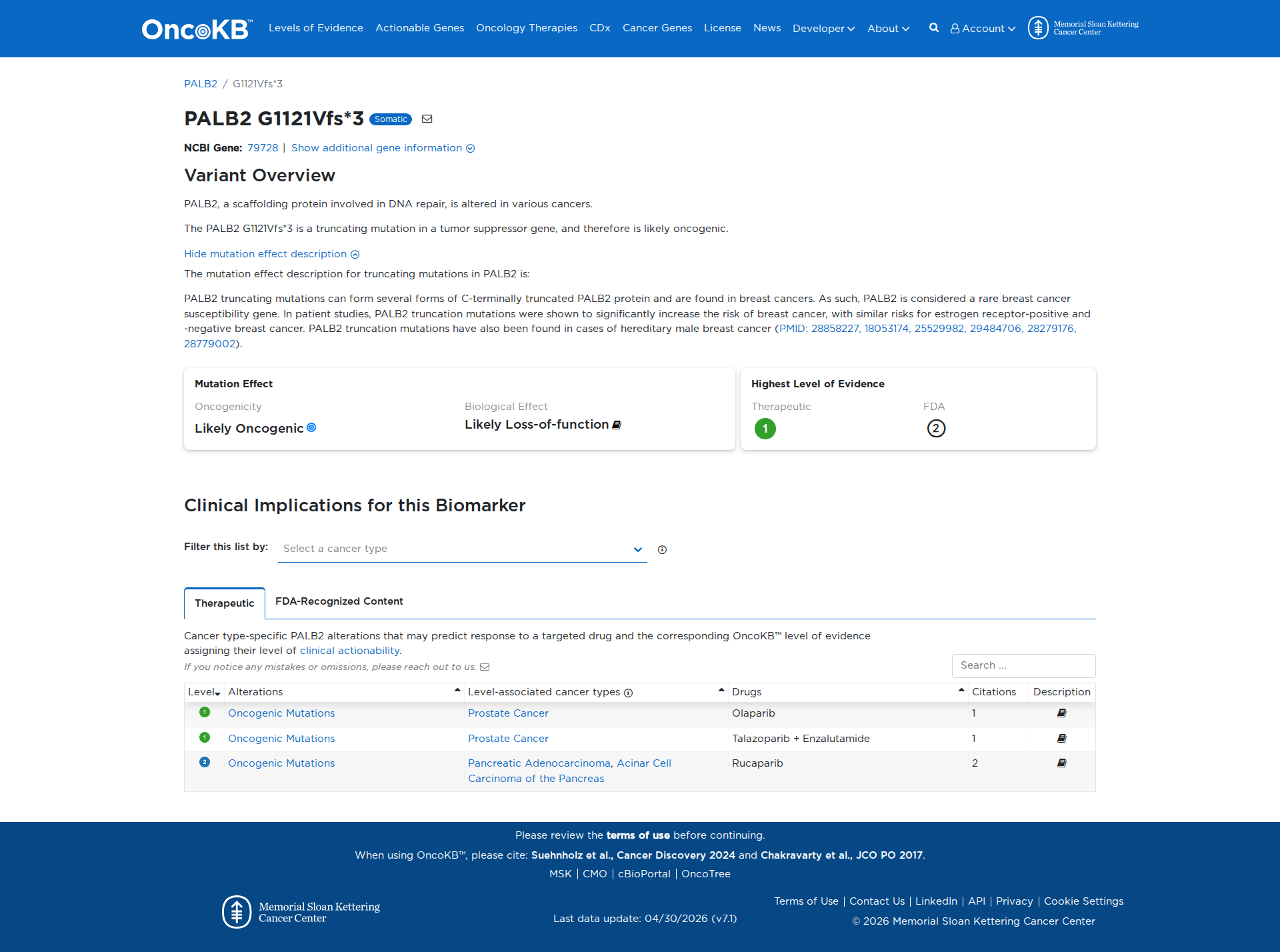
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 12 bp |
| Donor Loss (DL) | 0.0 | 2 bp |
| Acceptor Gain (AG) | 0.0 | 0 bp |
| Donor Gain (DG) | 0.0 | 143 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)