Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_024675.4 | MANE Select | 4008 nt | 154–3714 |
| NM_024675.3 | RefSeq Select | 4069 nt | 201–3761 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: PALB2 c.3054G>C (p.Glu1018Asp) results in a conservative amino acid change located in the Partner and localiser of BRCA2, WD40 domain (IPR031920) of the encoded protein sequence. Three of five in-silico tools predict a damaging effect of the variant on protein function. The variant allele was found at a frequency of 0.00037 in 281404 control chromosomes, predominantly at a frequency of 0.0052 within the East Asian subpopulation in the gnomAD database. The observed variant frequency within East Asian control individuals in the gnomAD database is approximately 33 fold of the estimated maximal expected allele frequency for a pathogenic variant in PALB2 causing Hereditary Breast and Ovarian Cancer phenotype (0.00016), strongly suggesting that the variant is a benign polymorphism found primarily in populations of East Asian origin. c.3054G>C has been reported in the literature predominantly in individuals of East Asian origin affected with Hereditary Breast and Ovarian Cancer (Casadei_2011, Tischkowitz_2012, Thompson_2015, Phuah_2013, Nguyen_Dumont_2015, Kraus_2017, Li_2015, Yang_2017). These report(s) do not provide unequivocal conclusions about association of the variant with Hereditary Breast and Ovarian Cancer. Co-occurrences with other pathogenic variant(s) have been reported in our internal testing database (BRCA1 c.2728delC, p.Gln910fsX90), providing supporting evidence for a benign role. To our knowledge, no experimental evidence demonstrating an impact on protein function has been reported. Seven clinical diagnostic laboratories have submitted clinical-significance assessments for this variant to ClinVar after 2014 without evidence for independent evaluation. Four of these classified the variant as Benign/Likely benign and three classified it as a VUS. Based on the evidence outlined above, the variant was classified as benign.
Curators: Marc Tischkowitz, Arleen D. Auerbach. Submitters to LOVD: Marc Tischkowitz, Yukihide Momozawa.
This alteration is classified as likely benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
"This variant has been reported in ClinVar as Benign (6 clinical laboratories) and as Likely benign (7 clinical laboratories) and as Uncertain significance (2 clinical laboratories) and as Benign by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open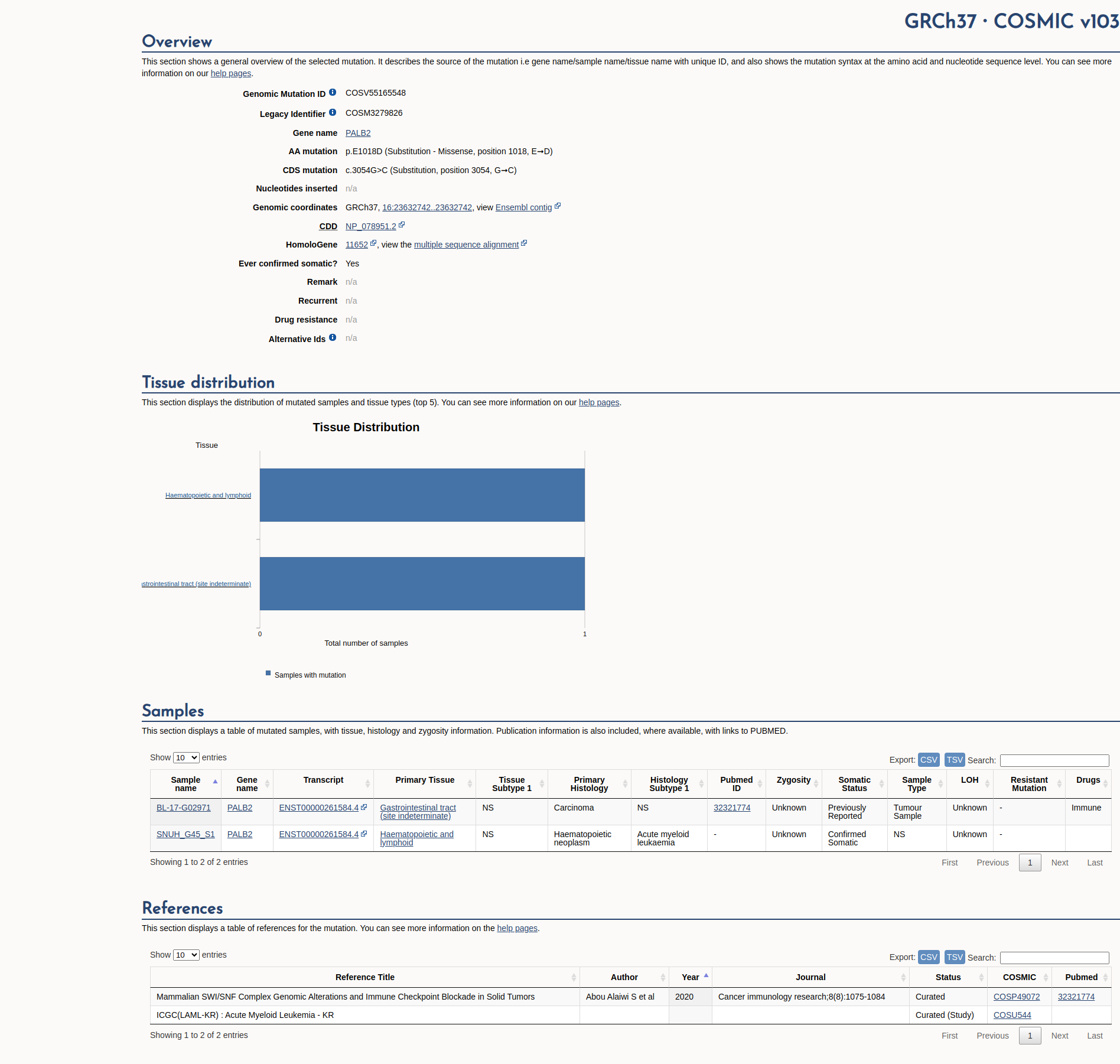
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PALB2E1018DPALB2E1018DSomaticNCBI Gene:79728|Show additional gene information Variant OverviewPALB2, a scaffolding protein involved in DNA repair, is altered in various cancers.The PALB2 E1018D mutation has not specifically been reviewed by the OncoKB team, and therefore its biological significance is unknown. JAX-CKB: PALB2 E1018D lies within WD repeat 4 of the Palb2 protein (UniProt.org). E1018D results in DNA damage-induced cell cycle checkpoint maintenance and homology-directed DNA repair activity similar to wild-type in cultured cells (PMID: 31757951, PMID: 35853885), and therefore, is predicted to have no effect on Palb2 protein function.
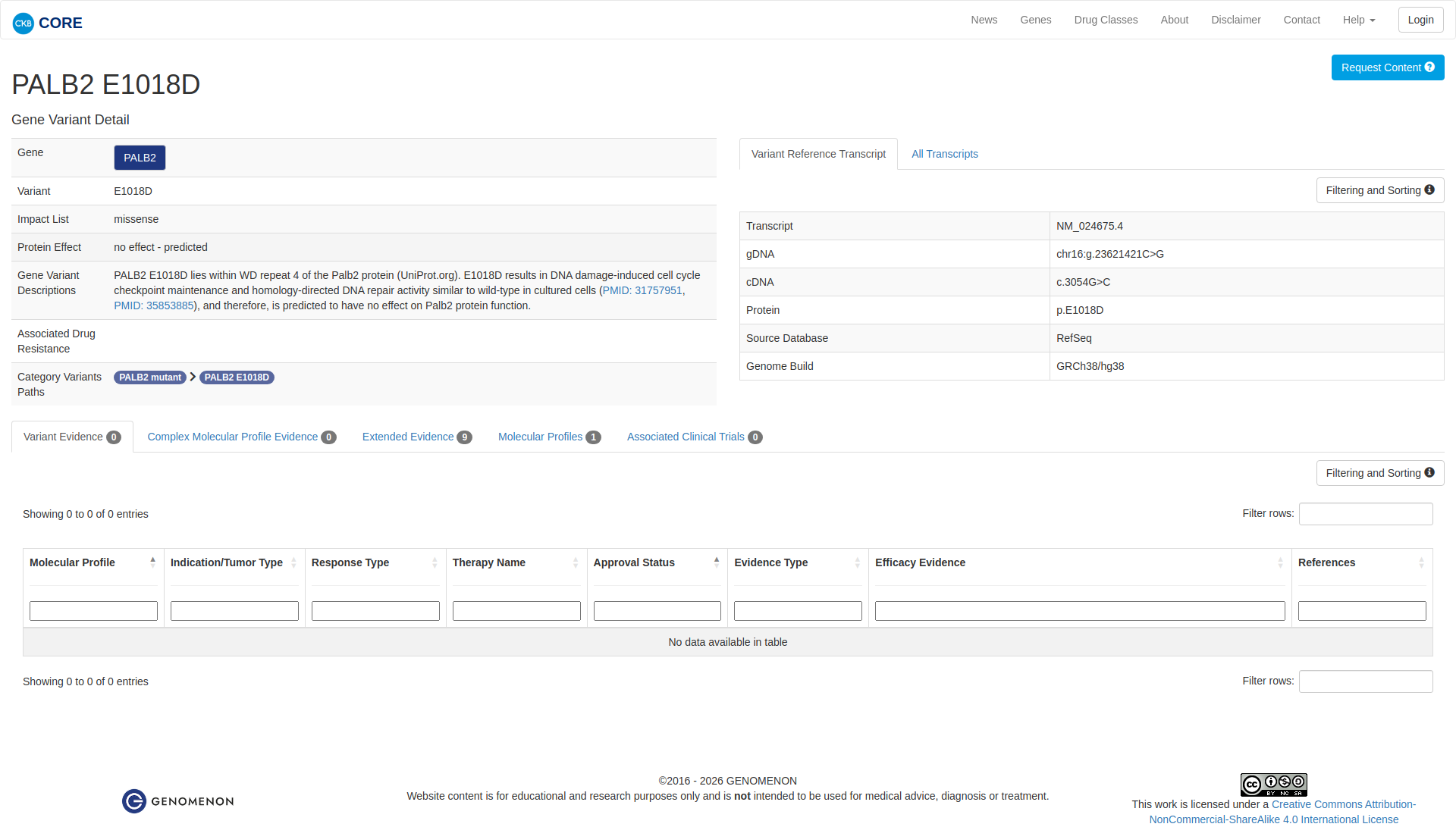
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 401 bp |
| Donor Loss (DL) | 0.0 | 308 bp |
| Acceptor Gain (AG) | 0.0 | 57 bp |
| Donor Gain (DG) | 0.01 | -28 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)