Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000051.3 | RefSeq Select | 13147 nt | 386–9556 |
| NM_000051.4 | MANE Select | 12915 nt | 151–9321 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe p.R23* pathogenic mutation (also known as c.67C>T), located in coding exon 1 of the ATM gene, results from a C to T substitution at nucleotide position 67. This changes the amino acid from an arginine to a stop codon within coding exon 1. This mutation has been reported in the tumor and germline of an individual diagnosed with mantle cell lymphoma (Camacho E et al. Blood. 2002 Jan;99(1):238-44), and in a patient with gastric cancer (Huang DS et al. Oncotarget. 2015 Dec;6(38):40953-8). In addition to the clinical data presented in the literature, this alteration is expected to result in loss of function by premature protein truncation or nonsense-mediated mRNA decay. As such, this alteration is interpreted as a disease-causing mutation.
This variant changes 1 nucleotide in exon 2 of the ATM gene, creating a premature translation stop signal. This variant is expected to result in an absent or non-functional protein product. This variant has been reported in the homozygous state or with a second pathogenic ATM variant in multiple individuals affected with ataxia-telangiectasia (PMID: 26896183, 33547824). This variant has also been reported in individuals affected with mantle cell lymphoma, gastric cancer, medulloblastoma, and breast cancer (PMID: 11756177, 26506520, 29753700, 31214711). This variant has been identified in 5/251364 chromosomes in the general population by the Genome Aggregation Database (gnomAD). Loss of ATM function is a known mechanism of disease (clinicalgenome.org). Based on the available evidence, this variant is classified as Pathogenic.
This sequence change creates a premature translational stop signal (p.Arg23*) in the ATM gene. It is expected to result in an absent or disrupted protein product. Loss-of-function variants in ATM are known to be pathogenic (PMID: 23807571, 25614872). This variant is present in population databases (rs746235533, gnomAD 0.01%). This premature translational stop signal has been observed in individual(s) with lymphoma, gastric cancer, and medulloblastoma (PMID: 11756177, 26506520, 29753700). ClinVar contains an entry for this variant (Variation ID: 232248). RNA analysis performed to evaluate the impact of this premature translational stop signal on mRNA splicing indicates it does not significantly alter splicing (internal data). For these reasons, this variant has been classified as Pathogenic.
Variant summary: ATM c.67C>T (p.Arg23X) results in a premature termination codon, predicted to cause a truncation of the encoded protein or absence of the protein due to nonsense mediated decay, which are commonly known mechanisms for disease. Truncations downstream of this position have been classified as pathogenic by our laboratory. The variant allele was found at a frequency of 2e-05 in 251364 control chromosomes (gnomAD). c.67C>T has been reported in the literature in homozygous and compound heterozygous individuals affected with Ataxia-Telangiectasia (examples: Amirfar_2021 and Jackson_2016). These data indicate that the variant is very likely associated with disease. Jackson_2016 demonstrated the homozygous patient's lymphoblastoid cell lines showing ATM protein was present, kinase activity was absent and radiosensitivity was high. Five clinical diagnostic laboratories have submitted clinical-significance assessments for this variant to ClinVar after 2014 and all classified the variant as pathogenic. Based on the evidence outlined above, the variant was classified as pathogenic.
"This variant has been reported in ClinVar as Pathogenic (13 clinical laboratories) and as Pathogenic by ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open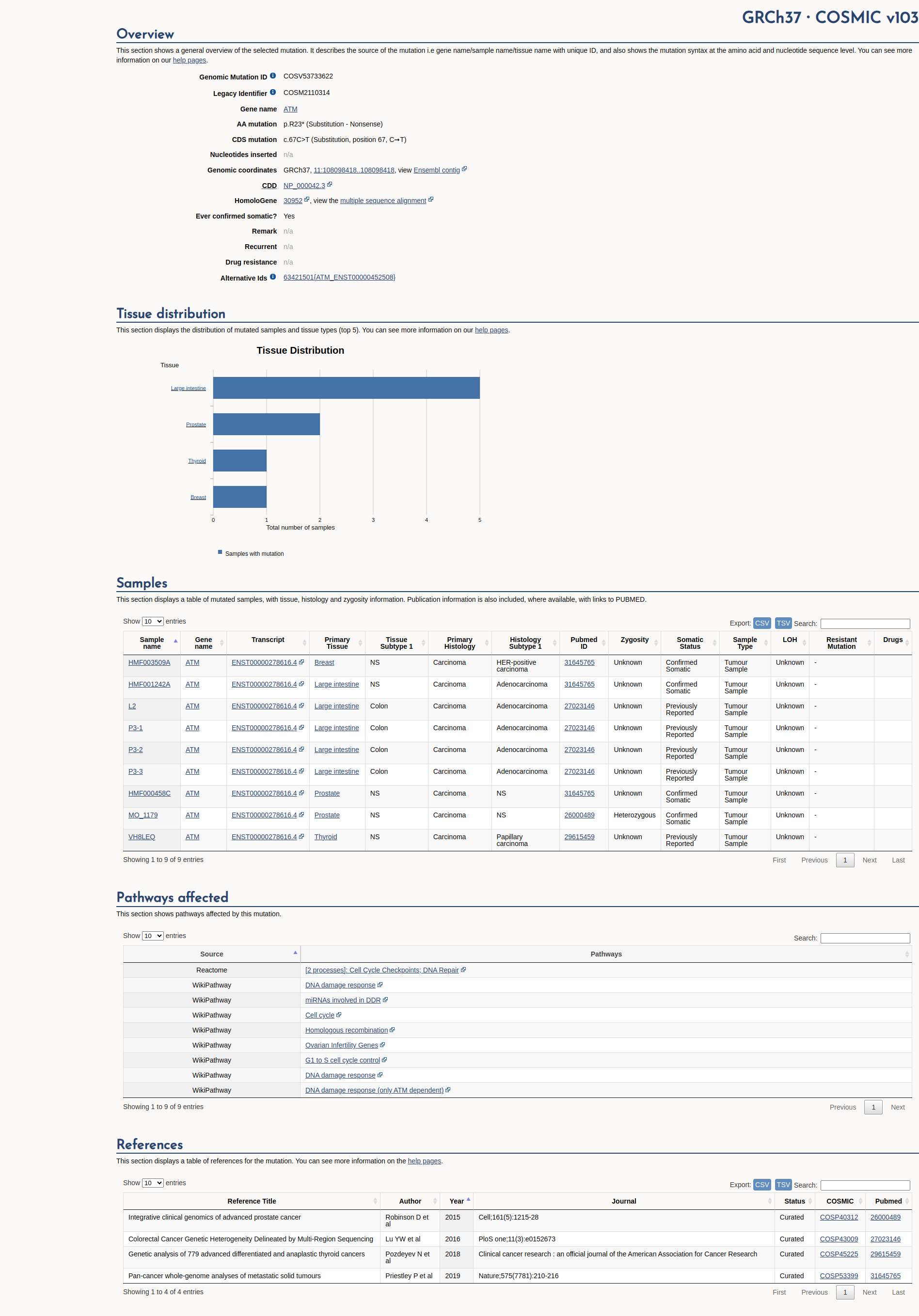
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: ATMR23*ATMR23*SomaticNCBI Gene:472|Show additional gene information Variant OverviewATM, a kinase involved in the DNA damage response, is mutated in various solid and hematologic malignancies.The ATM R23* is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in ATM is: ATM truncating mutations can produce several forms of C-terminally truncated ATM proteins. When found in the germline, these mutations result in ataxia-telangiectasia syndrome, which increases cancer predisposition, including a 20% to 30% lifetime risk of lymphoid, gastric, breast, central nervous system, skin, and other cancers (PMID: 27413114). Deletion of ATM in mouse models and cell lines demonstrates that it is oncogenic as measured by decreased DNA repair efficiency and increased cellular motility (PMID: 30553448, 30348496). JAX-CKB: No results found
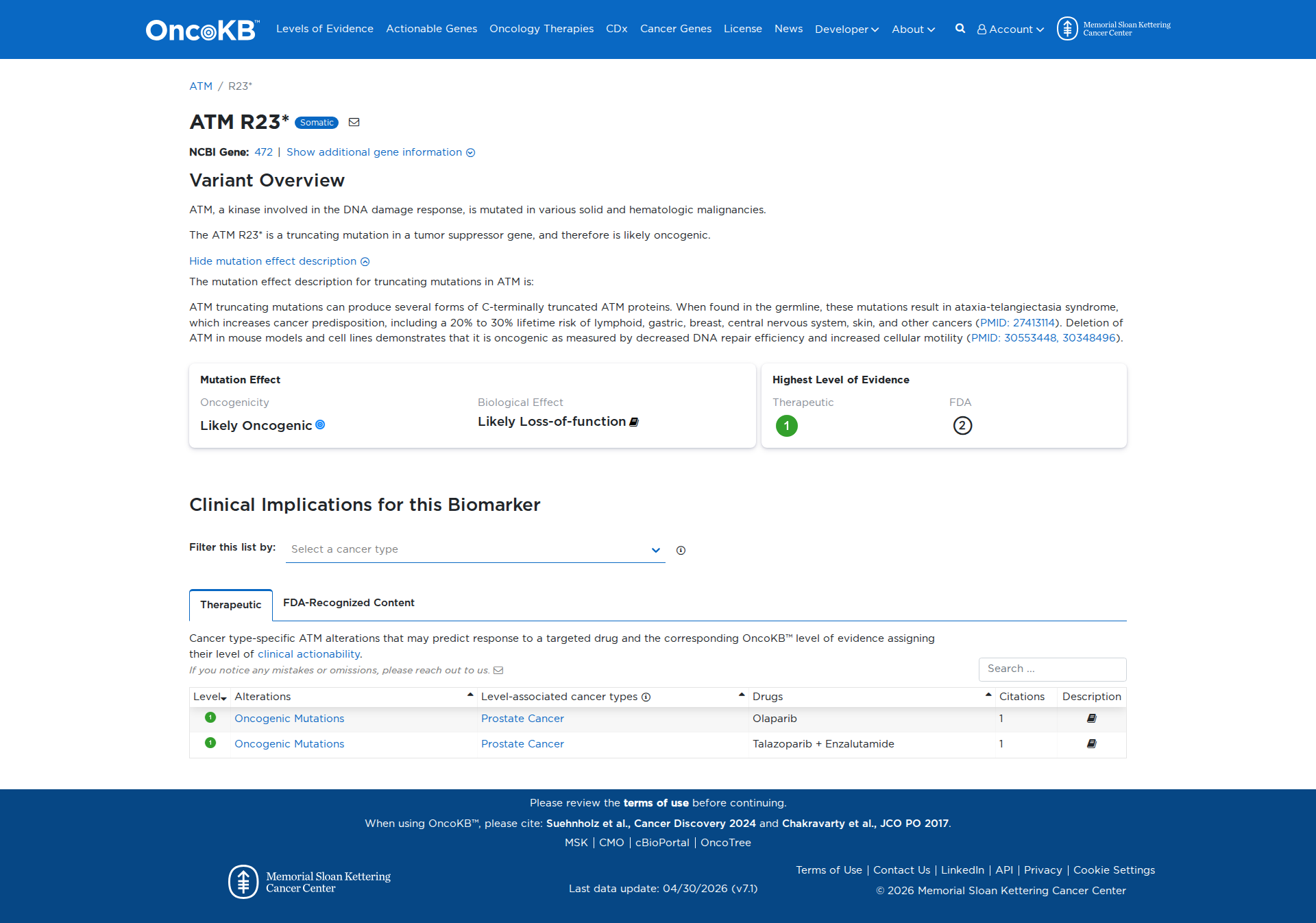
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.12 | -96 bp |
| Donor Loss (DL) | 0.16 | 5 bp |
| Acceptor Gain (AG) | 0.0 | 85 bp |
| Donor Gain (DG) | 0.0 | -96 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)