Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_001354609.2 | Alternative | 9687 nt | 227–2530 |
| NM_001354609.1 | Alternative | 9702 nt | 226–2529 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenIn summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic. This variant disrupts the p.Thr241 amino acid residue in BRAF. Other variant(s) that disrupt this residue have been determined to be pathogenic (PMID: 17704260, 18042262, 19206169, 28404629). This suggests that this residue is clinically significant, and that variants that disrupt this residue are likely to be disease-causing. Advanced modeling performed at Invitae incorporating data from internal and/or published experimental studies (Invitae) indicates that this missense variant is expected to disrupt BRAF function. ClinVar contains an entry for this variant (Variation ID: 44829). This missense change has been observed in individual(s) with clinical features of RASopathy spectrum disorders (Invitae). This variant is not present in population databases (gnomAD no frequency). This sequence change replaces threonine, which is neutral and polar, with lysine, which is basic and polar, at codon 241 of the BRAF protein (p.Thr241Lys).
"This variant has been reported in ClinVar as Pathogenic (1 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as Pathogenic by ClinGen RASopathy Variant Curation Expert Panel expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRAFT241KBRAFT241KSomaticNCBI Gene:673|Show additional gene information Variant OverviewBRAF, an intracellular kinase, is frequently mutated in melanoma, thyroid and lung cancers among others.The BRAF T241K mutation has not specifically been reviewed by the OncoKB team. However, BRAF T241P is likely oncogenic, and therefore BRAF T241K is considered likely oncogenic.Hide mutation effect description The BRAF T241K mutation has not specifically been reviewed by the OncoKB team. However, the mutation effect description for BRAF T241P, an alternate allele of BRAF T241K, is: The BRAF T241P mutation is located in the cysteine-rich domain in exon 6 of the protein. This mutation has been found in cardio-facio-cutaneous syndrome (CFC) (PMID: 19206169). Expression of this mutation in NIH-3T3 cells demonstrated that it is likely activating as measured by moderately increased activation of MEK and ERK and slightly increased colony formation compared to wildtype BRAF (PMID: 19206169). JAX-CKB: No results found
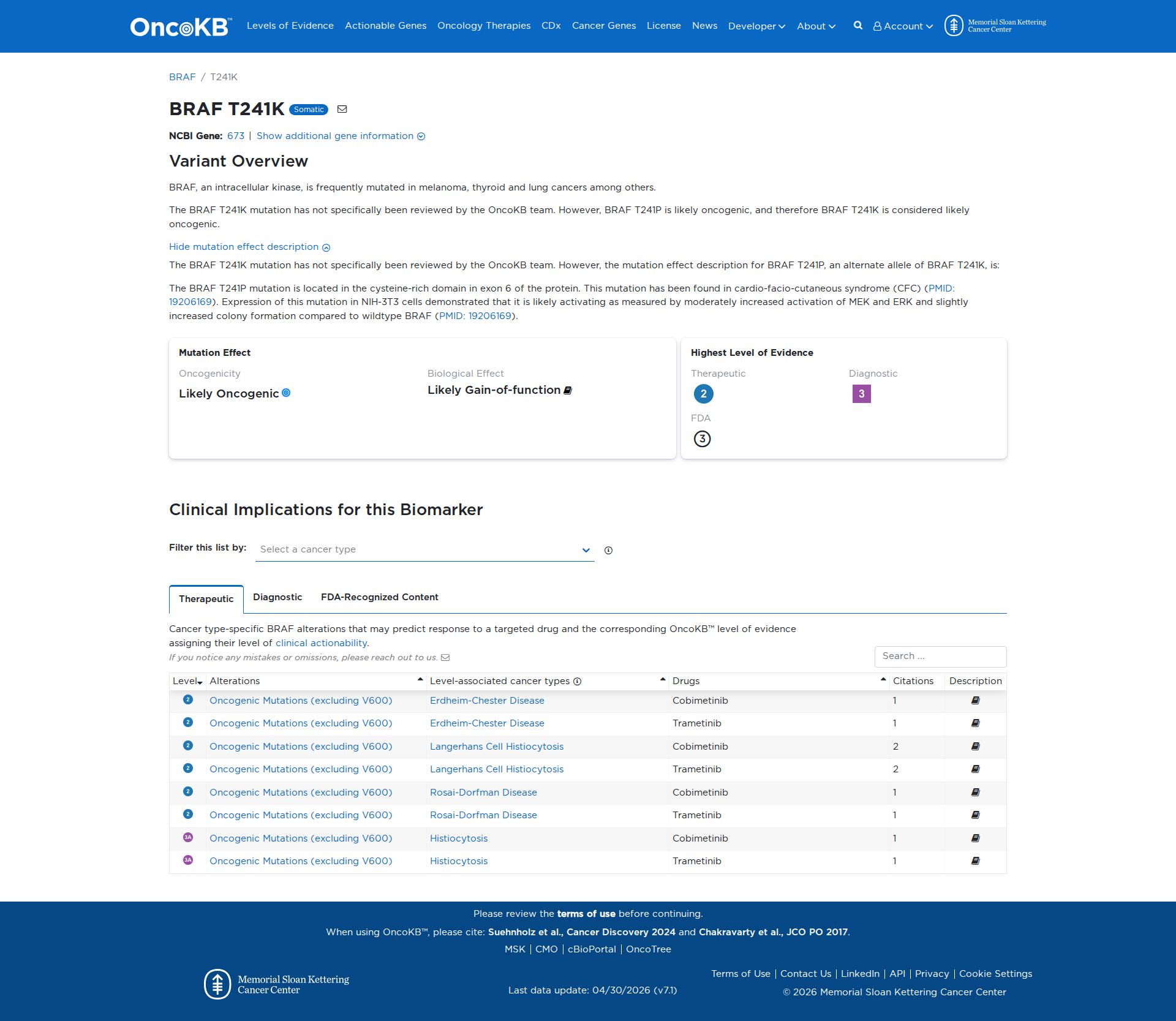

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -15 bp |
| Donor Loss (DL) | 0.0 | -138 bp |
| Acceptor Gain (AG) | 0.04 | -38 bp |
| Donor Gain (DG) | 0.0 | -142 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)