Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe p.R196L variant (also known as c.587G>T), located in coding exon 5 of the TP53 gene, results from a G to T substitution at nucleotide position 587. The arginine at codon 196 is replaced by leucine, an amino acid with dissimilar properties. Studies conducted in human cell lines indicate this alteration is proficient at growth suppression and has no dominant negative effect (Kotler E et al. Mol.Cell. 2018 Jul;71:178-190.e8; Giacomelli AO et al. Nat. Genet. 2018 Oct;50:1381-1387). This variant is in the DNA binding domain of the TP53 protein and is reported to have partially functional transactivation in yeast based assays (Kato S et al. Proc. Natl. Acad. Sci. USA. 2003 Jul;100:8424-9). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the available evidence, the clinical significance of this variant remains unclear.
This sequence change replaces arginine, which is basic and polar, with leucine, which is neutral and non-polar, at codon 196 of the TP53 protein (p.Arg196Leu). This variant is not present in population databases (gnomAD no frequency). This missense change has been observed in individual(s) with clinical features of Li-Fraumeni Syndrome (PMID: 33850299, 34308366). ClinVar contains an entry for this variant (Variation ID: 100814). Invitae Evidence Modeling incorporating data from in vitro experimental studies (PMID: 12826609, 29979965, 30224644) indicates that this missense variant is expected to disrupt TP53 function with a positive predictive value of 97.5%. Algorithms developed to predict the effect of sequence changes on RNA splicing suggest that this variant may disrupt the consensus splice site. In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
This missense variant replaces arginine with leucine at codon 196 of the TP53 protein. Computational prediction suggests that this variant may have deleterious impact on protein structure and function (internally defined REVEL score threshold >= 0.7, PMID: 27666373). Functional studies have shown that this variant has a neutral effect on TP53 protein function (PMID: 29979965, 30224644). To our knowledge, this variant has not been reported in individuals affected with hereditary cancer in the literature. This variant has not been identified in the general population by the Genome Aggregation Database (gnomAD). The available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
"This variant has been reported in ClinVar as Uncertain significance (5 clinical laboratories)."
COSMIC Somatic Evidence
Open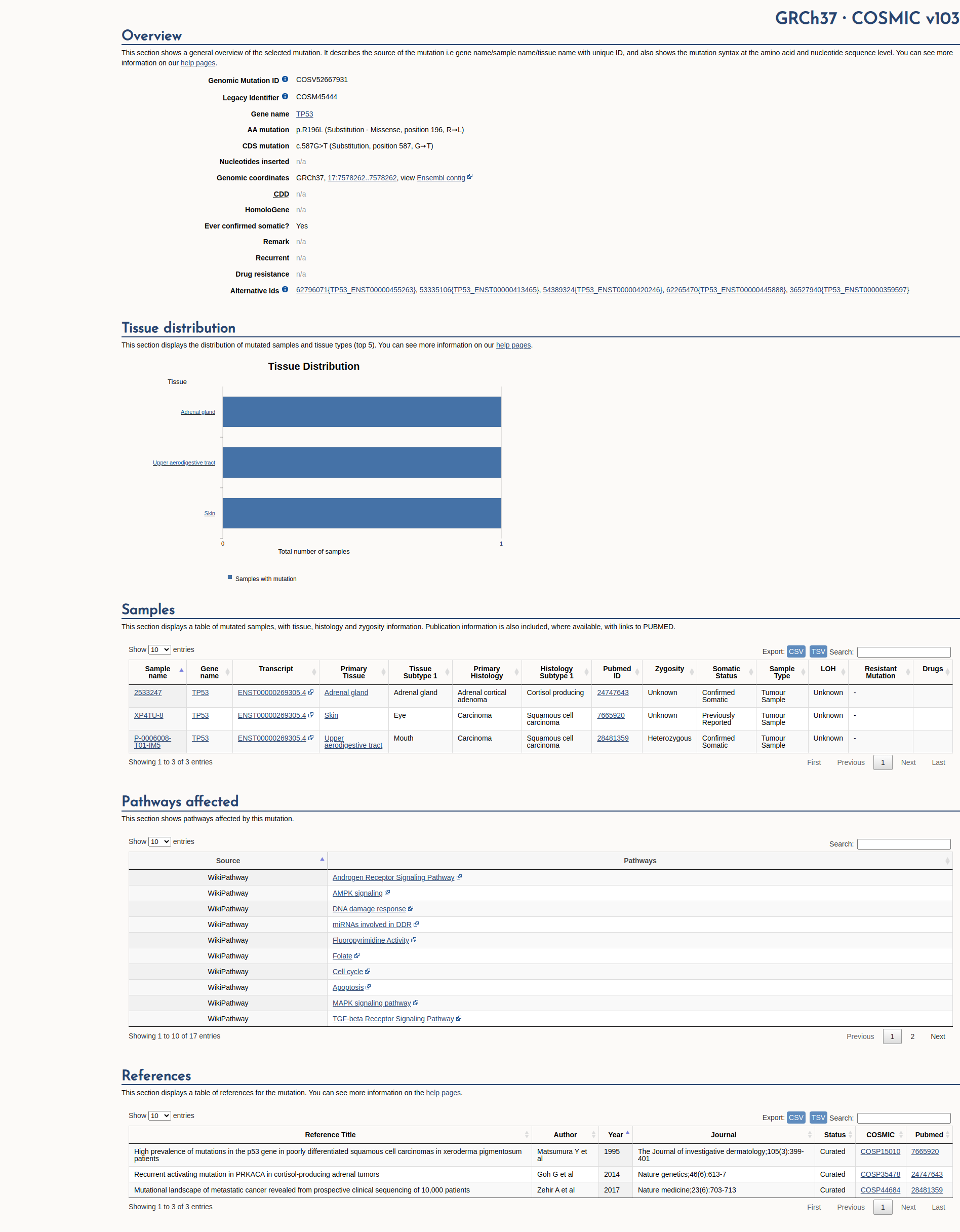
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: TP53R196LTP53R196LSomaticNCBI Gene:7157|Show additional gene information Variant OverviewTP53, a tumor suppressor in the DNA damage pathway, is the most frequently mutated gene in cancer.There is conflicting and/or weak data describing the biological significance of the TP53 R196L mutation.Hide mutation effect description The TP53 R196L mutation arises in the p53 DNA-binding domain of the protein. When this mutation is expressed in human Saos cells at different experimental temperature conditions, a difference in transactivational activity of this TP53 mutant has been observed (PMID: 14559903). However, there is no specific data demonstrating whether the mutation has decreased transactivational activity relative to wildtype p53 (PMID: 14559903). Nonetheless, an alternate mutation at this position, the TP53 R196G mutation, has been demonstrated to decrease p53 transactivation activity relative to wildtype p53 (PMID: 22710932). The R196L mutation is also a 3D hotspot mutation and therefore TP53 R196L is considered likely oncogenic (PMID: 28115009). JAX-CKB: TP53 R196L lies within the DNA-binding domain of the Tp53 protein (PMID: 21760703). R196L has been identified in the scientific literature (PMID: 25847421, PMID: 27432539, PMID: 33850299), but has not been biochemically characterized and therefore, its effect on Tp53 protein function is unknown (PubMed, Aug 2025).
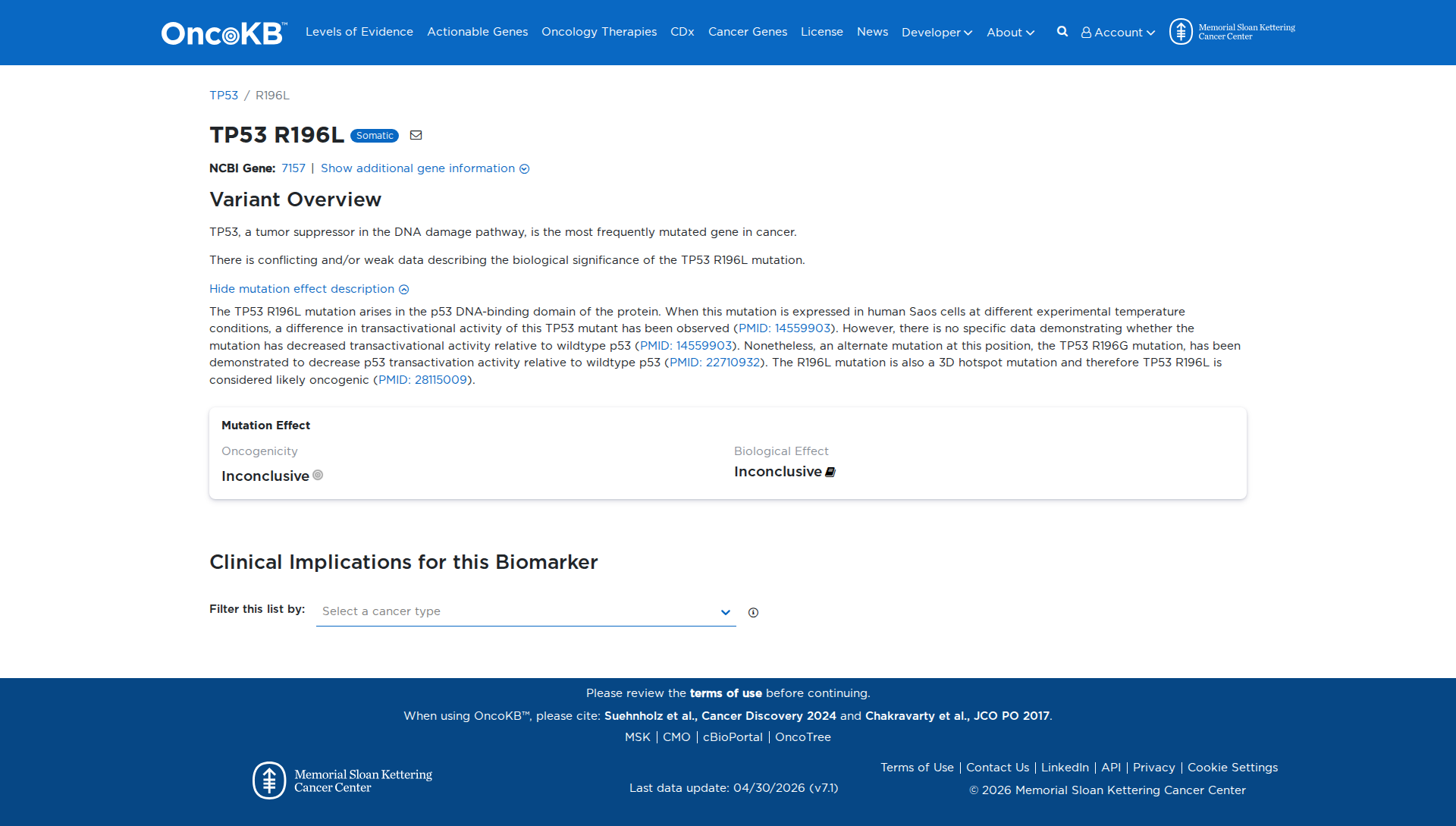

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.14 | 10 bp |
| Donor Loss (DL) | 0.08 | -90 bp |
| Acceptor Gain (AG) | 0.0 | -3 bp |
| Donor Gain (DG) | 0.0 | 392 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)