Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000051.3 | RefSeq Select | 13147 nt | 386–9556 |
| NM_000051.4 | MANE Select | 12915 nt | 151–9321 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenFor these reasons, this variant has been classified as Pathogenic. Loss-of-function variants in ATM are known to be pathogenic (PMID: 23807571, 25614872). This variant has not been reported in the literature in individuals with ATM-related conditions. This variant is not present in population databases (ExAC no frequency). This sequence change creates a premature translational stop signal (p.Gln2684Hisfs*8) in the ATM gene. It is expected to result in an absent or disrupted protein product.
"This variant has been reported in ClinVar as Pathogenic (3 clinical laboratories)."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: ATMQ2684Hfs*8ATMQ2684Hfs*8SomaticNCBI Gene:472|Show additional gene information Variant OverviewATM, a kinase involved in the DNA damage response, is mutated in various solid and hematologic malignancies.The ATM Q2684Hfs*8 is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in ATM is: ATM truncating mutations can produce several forms of C-terminally truncated ATM proteins. When found in the germline, these mutations result in ataxia-telangiectasia syndrome, which increases cancer predisposition, including a 20% to 30% lifetime risk of lymphoid, gastric, breast, central nervous system, skin, and other cancers (PMID: 27413114). Deletion of ATM in mouse models and cell lines demonstrates that it is oncogenic as measured by decreased DNA repair efficiency and increased cellular motility (PMID: 30553448, 30348496). JAX-CKB: No results found
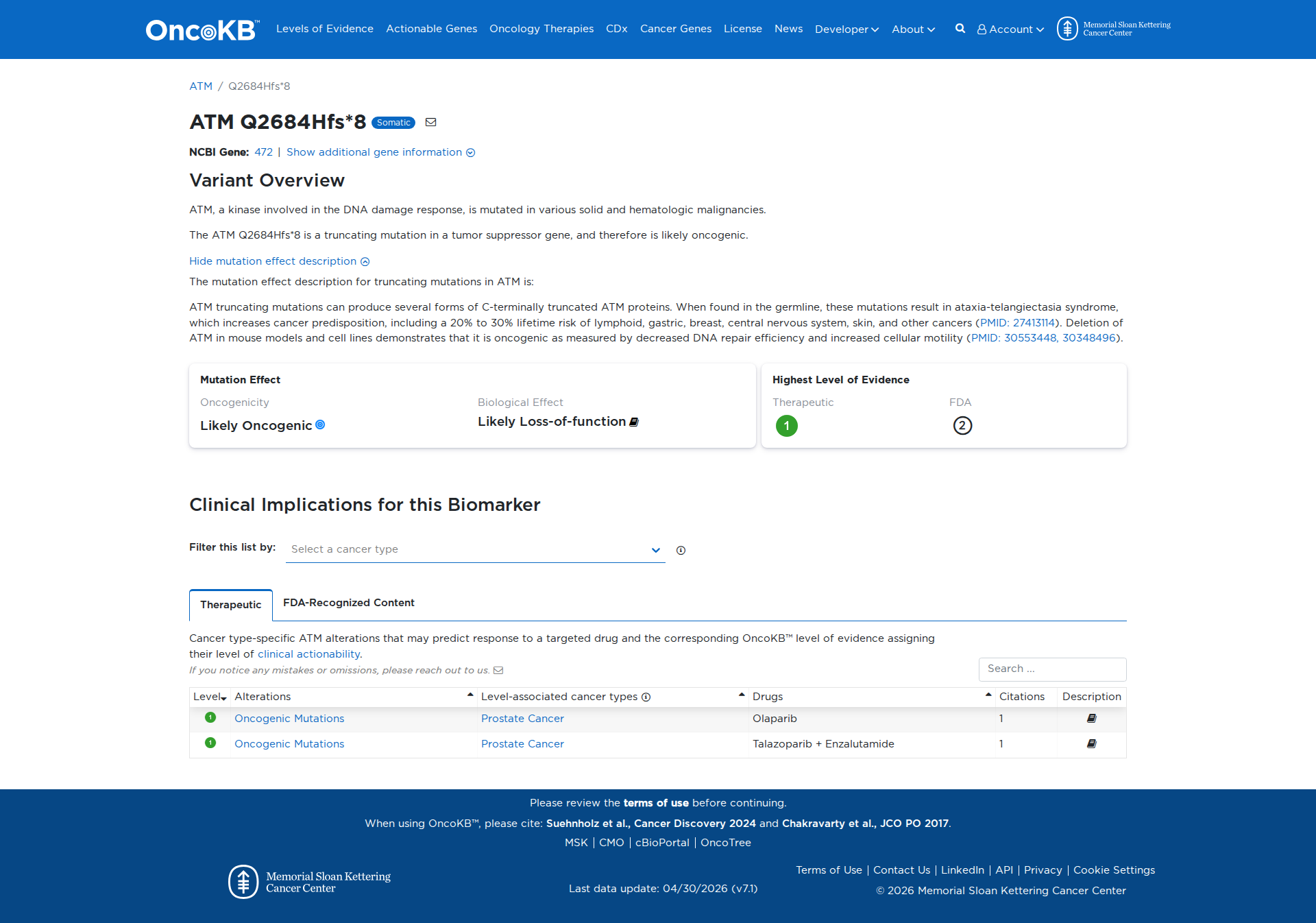

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 416 bp |
| Donor Loss (DL) | 0.0 | 34 bp |
| Acceptor Gain (AG) | 0.0 | -25 bp |
| Donor Gain (DG) | 0.01 | -9 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)