Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000546.3 | Alternative | 2640 nt | 252–1433 |
| NM_000546.5 | RefSeq Select | 2591 nt | 203–1384 |
| NM_000546.6 | MANE Select | 2512 nt | 143–1324 |
| NM_000546.4 | Alternative | 2586 nt | 198–1379 |
| NM_000546.2 | Alternative | 2629 nt | 252–1433 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenVariant summary: The variant, TP53 c.344A>G (p.His115Arg) results in a non-conservative amino acid change located in the DNA-binding domain (IPR011615) of the encoded protein sequence. Three of five in-silico tools predict a damaging effect of the variant on protein function. The variant was absent in 245988 control chromosomes (gnomAD). The available data on variant occurrences in the general population are insufficient to allow any conclusion about variant significance. To our knowledge, no occurrence of c.344A>G in individuals affected with Li-Fraumeni Syndrome or related tumor phenotypes has been reported. Publications reported experimental evidence evaluating an impact on protein function, where one study demonstrated that the variant, p.H115R, does not affect the transactivation activity on eight different promoter elements (including p21) (data taken from the IARC Database, reference: Kato 2003), while another study demonstrated that the variant protein has a decreased oligonucleotide binding activity in an EMSA assay (where the probe contained the p53 response element from the p21 promoter) (Zupnick 2006). These data does not allow unequivocal conclusions about the variant effect. Three clinical diagnostic laboratories have submitted clinical-significance assessments for this variant to ClinVar after 2014 without evidence for independent evaluation. All laboratories classified the variant as uncertain significance. Based on the evidence outlined above, the variant was classified as uncertain significance.
This alteration is classified as likely benign based on a combination of the following: seen in unaffected individuals, population frequency, intact protein function, lack of segregation with disease, co-occurrence, RNA analysis, in silico models, amino acid conservation, lack of disease association in case-control studies, and/or the mechanism of disease or impacted region is inconsistent with a known cause of pathogenicity.
This missense variant replaces histidine with arginine at codon 115 of the TP53 protein. Computational prediction suggests that this variant may not impact protein structure and function (internally defined REVEL score threshold <= 0.5, PMID: 27666373). Functional studies have shown the mutant protein to exhibit normal transactivation activity in a yeast transactivation assay (PMID 12826609 and IARC database) and to be functional in human cell growth assays (PMID: 29979965, 30224644). The mutant protein has shown a partially reduced binding affinity for the p53 response element from the p21 promoter (PMID: 16687402). This variant has not been reported in individuals affected with hereditary cancer in the literature. This variant has not been identified in the general population by the Genome Aggregation Database (gnomAD). The available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
This sequence change replaces histidine, which is basic and polar, with arginine, which is basic and polar, at codon 115 of the TP53 protein (p.His115Arg). This variant is not present in population databases (gnomAD no frequency). This variant has not been reported in the literature in individuals affected with TP53-related conditions. ClinVar contains an entry for this variant (Variation ID: 182925). Invitae Evidence Modeling incorporating data from in vitro experimental studies (PMID: 12826609, 29979965, 30224644) indicates that this missense variant is not expected to disrupt TP53 function with a negative predictive value of 97.5%. Experimental studies have shown that this missense change does not substantially affect TP53 function (PMID: 12826609, 29979965, 30224644). In summary, the available evidence is currently insufficient to determine the role of this variant in disease. Therefore, it has been classified as a Variant of Uncertain Significance.
This missense variant replaces histidine with arginine at codon 115 of the TP53 protein. Computational prediction suggests that this variant may not impact protein structure and function Functional studies have shown that this variant to exhibit normal transactivation activity in a yeast transactivation assay and to be functional in human cell growth and proliferation assays (PMID: 12826609, 29979965, 30224644). To our knowledge, this variant has not been reported in individuals affected with TP53-related disorders in the literature. This variant has not been identified in the general population by the Genome Aggregation Database (gnomAD). The available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
"This variant has been reported in ClinVar as Uncertain significance (3 clinical laboratories) and as Likely benign (2 clinical laboratories) and as Uncertain Significance (1 clinical laboratories) and as Likely Benign by ClinGen TP53 Variant Curation Expert Panel, ClinGen expert panel."
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: TP53H115RTP53H115RSomaticNCBI Gene:7157|Show additional gene information Variant OverviewTP53, a tumor suppressor in the DNA damage pathway, is the most frequently mutated gene in cancer.The TP53 H115R mutation is likely oncogenic.Hide mutation effect description The mutation effect description for DNA binding domain missense mutations in TP53 is: The DNA-binding domain (DBD) of TP53 allows for contact with target DNA sequences to transactivate downstream genes. Mutations in the DBD result in conformational changes of the protein, altering contact of TP53 with its target DNA sequences and thereby altering transcriptional function (PMID: 8023157, 11900253). Given that TP53 directs the transcription of proteins that enable apoptosis, the inactivation of TP53 results in cells harboring damaged DNA and overall genomic instability (PMID: 11900253, 11900253). JAX-CKB: No results found
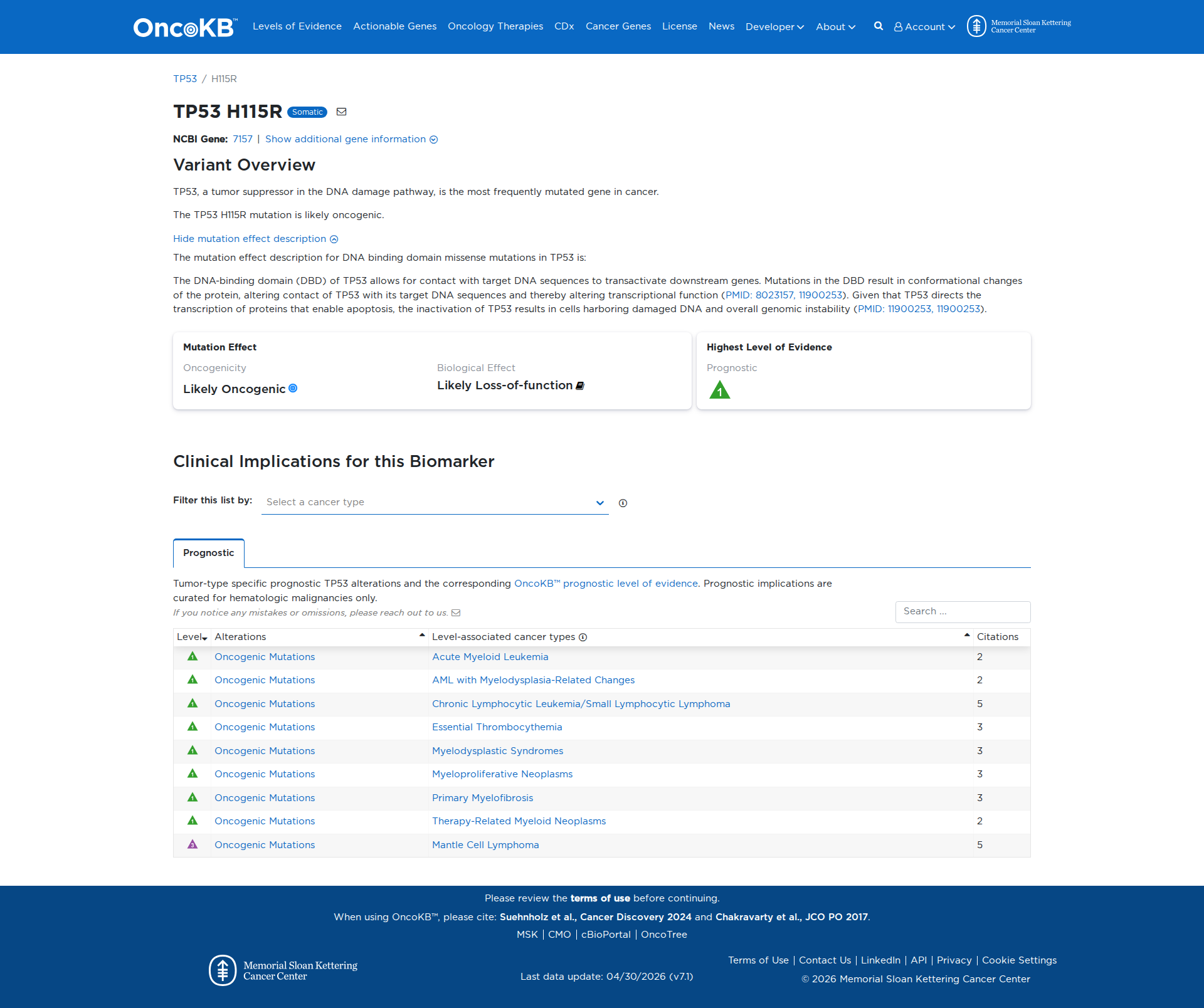

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 228 bp |
| Donor Loss (DL) | 0.02 | 169 bp |
| Acceptor Gain (AG) | 0.03 | 43 bp |
| Donor Gain (DG) | 0.0 | -31 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)