Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_006218.2 | Alternative | 3724 nt | 158–3364 |
| NM_006218.3 | Alternative | 9104 nt | 158–3364 |
| NM_006218.4 | MANE Select | 9259 nt | 324–3530 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThe c.1132T>C (p.C378R) alteration is located in exon 6 (coding exon 5) of the PIK3CA gene. This alteration results from a T to C substitution at nucleotide position 1132, causing the cysteine (C) at amino acid position 378 to be replaced by an arginine (R). This variant was not reported in population-based cohorts in the Genome Aggregation Database (gnomAD). This variant has been identified as a confirmed or suspected result of somatic mosaicism in affected tissue samples from individual(s) with vascular malformations, limb and/or digital anomalies, and adipose overgrowth; all features consistent with PIK3CA-related disorder (Parker, 2019; Delgado-Miguel, 2021; Paolacci, 2020; Su, 2022; Zerbib, 2024; McNulty, 2019). In at least one individual, it was determined to be de novo (Lunke, 2023). This amino acid position is highly conserved in available vertebrate species. This alteration is predicted to be deleterious by in silico analysis. Based on the available evidence, this alteration is classified as pathogenic.
"This variant has been reported in ClinVar as Likely pathogenic (1 clinical laboratories) and as Pathogenic (4 clinical laboratories)."
COSMIC Somatic Evidence
Open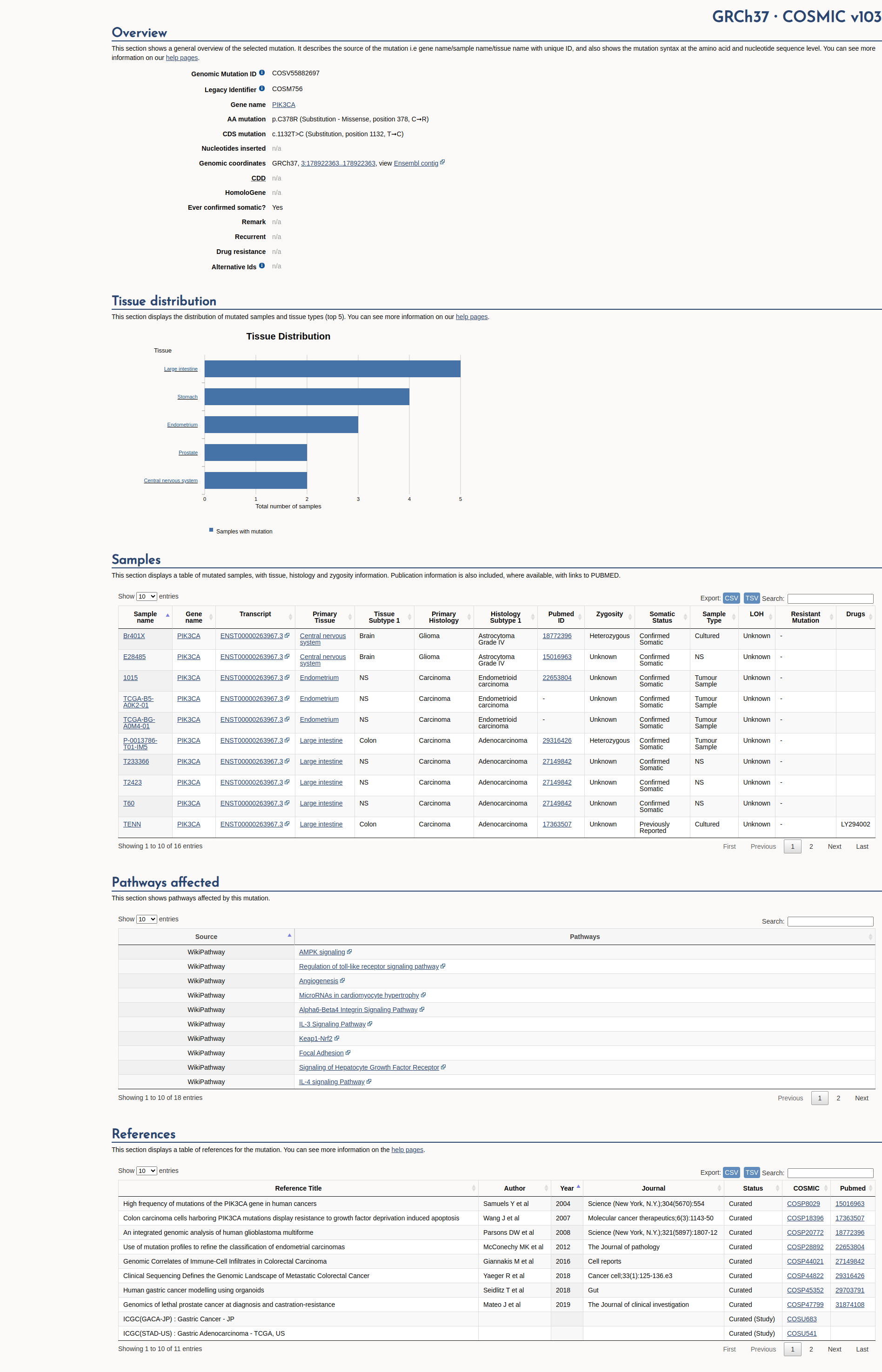
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: PIK3CAC378RPIK3CAC378RSomaticNCBI Gene:5290|Show additional gene information Variant OverviewPIK3CA, the catalytic subunit of PI3-kinase, is frequently mutated in a diverse range of cancers including breast, endometrial and cervical cancers.The PIK3CA C378R mutation is likely oncogenic.Hide mutation effect description The PIK3CA C378R mutation is located in the C2 domain in exon 6 of the protein. This mutation has been found in colon cancer (PMID: 17363507). A colon cancer cell line with this mutation displayed increased PI3K enzyme activity, increased activation of downstream PI3K targets and resistance to apoptosis compared to cancer cells with wildtype PIK3CA (PMID: 17363507). JAX-CKB: PIK3CA C378R lies within the C2 PI3K-type domain of the Pik3ca protein (UniProt.org). C378R results in constitutive activation of Pi3k and Akt signaling, and decreased apoptosis in cell culture (PMID: 17363507).
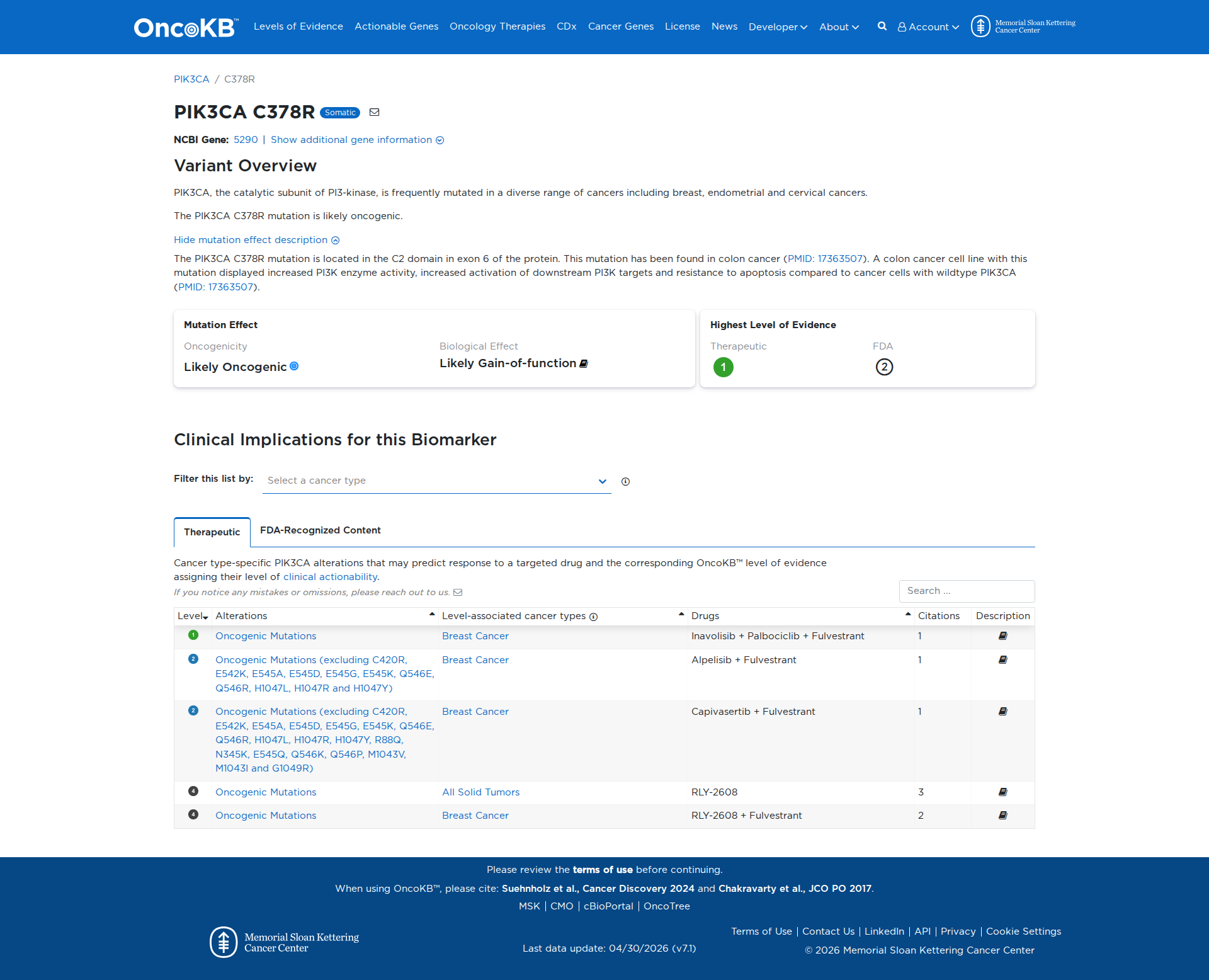

Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | 156 bp |
| Donor Loss (DL) | 0.0 | -57 bp |
| Acceptor Gain (AG) | 0.0 | -56 bp |
| Donor Gain (DG) | 0.0 | -383 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)