Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_007294.4 | MANE Select | 7088 nt | 114–5705 |
| NM_007294.2 | Alternative | 7191 nt | 201–5792 |
| NM_007294.3 | RefSeq Select | 7224 nt | 233–5824 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenPP5, PM2, PVS1
The p.Asn1355LysfsX10 variant in BRCA1 has been reported in >100 individuals with BRCA1-associated cancers (Friedman 1994 PMID:7894493, Zhang 2011 PMID:21324516, George 2013 PMID:23633455, Cao 2013 PMID:23175448, Cunningham 2014 PMID:24504028, Leongamornlert 2014 PMID:24556621, Rashid 2016 PMID:27553291, Sun 2017 PMID:28724667, Maxwell 2017 PMID:28831036, Hirasawa 2017 PMID:29348823, Li 2018 PMID:30078507, Singh 2018 PMID:29470806, Wen 2018 PMID:28993434, Li 2019 PMID:29752822, Breast Information Core Database (BIC): https://research.nhgri.nih.gov/bic/). It has also been identified in 0.004% (3/68040) European chromosomes by gnomAD (http://gnomad.broadinstitute.org, v.3.1). This variant is predicted to cause a frameshift, which alters the protein’s amino acid sequence beginning at position 1355 and leads to a premature termination codon 10 amino acids downstream. This alteration is then predicted to lead to a truncated or absent protein. Loss of function of the BRCA1 gene is an established disease mechanism in autosomal dominant hereditary breast and ovarian cancer (HBOC). Additionally, this variant was classified as pathogenic on April 22, 2016 by the ClinGen-approved ENIGMA expert panel (Variation ID 17674). In summary, this variant meets criteria to be classified as pathogenic for autosomal dominant HBOC. ACMG/AMP Criteria applied: PS4, PM2_Supporting, PVS1.
The BRCA1 c.4065_4068del p.(Asn1355LysfsTer10) variant causes a shift in the protein reading frame that is predicted to result in premature termination of the protein. Loss of normal protein function through either protein truncation or nonsense-mediated mRNA decay is expected. This variant has been observed in individuals with breast and ovarian cancer in several studies in the literature (PMIDs: 35908255; 35918668; 35409996; 35220195; 34657373). The highest frequency of this allele in the Genome Aggregation Database is 0.000041 in the European (non-Finnish) population (version 4.0.0). Additionally, a different proven pathogenic variant resulting in premature protein termination and located in the same exon has been reported in individuals with hereditary breast and ovarian cancer (ClinVar). Based on the available evidence, the c.4065_4068del p.(Asn1355LysfsTer10) is classified as pathogenic for hereditary breast and ovarian cancer.
This submission and the accompanying classification are no longer maintained by the submitter. For more information on current observations and classification, please contact variantquestions@myriad.com.
The c.4065_4068delTCAA pathogenic mutation, located in coding exon 9 of the BRCA1 gene, results from a deletion of 4 nucleotides at nucleotide positions 4065 to 4068, causing a translational frameshift with a predicted alternate stop codon (p.N1355Kfs*10). This alteration has been reported in multiple families with hereditary breast and ovarian cancer from a variety of ethnic backgrounds (Shattuck-Eidens D et al. JAMA. 1995 Feb;273:535-41; Langston AA et al. N. Engl. J. Med. 1996 Jan;334:137-42; Spitzer E et al. Int. J. Cancer. 2000 Feb;85:474-81; Ahn SH et al. Cancer Lett. 2007 Jan;245:90-5; Cunningham J et al. Sci. Rep. 2014 Feb;4:4026; Rashid MU et al. BMC Cancer. 2016 Aug;16:673; Li JY et al. Int. J. Cancer 2019 Jan;144(2):281-289; Siraj AK et al. Hum. Mutat. 2019 Mar). This mutation was also identified in an individual from a cohort of 191 prostate cancer patients with at least three prostate cancer cases in their families; this individual's family was also noted to be affected with breast and colon cancer (Leongamornlet D et al. Br. J. Cancer. 2014 Mar;110:1663-72). Of note, this alteration is also designated as 4184del4 in published literature. In addition to the clinical data presented in the literature, this alteration is expected to result in loss of function by premature protein truncation or nonsense-mediated mRNA decay. As such, this alteration is interpreted as a disease-causing mutation.
"This variant has been reported in ClinVar as Pathogenic (51 clinical laboratories) and as Likely pathogenic (1 clinical laboratories) and as Pathogenic by Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) expert panel."
COSMIC Somatic Evidence
Open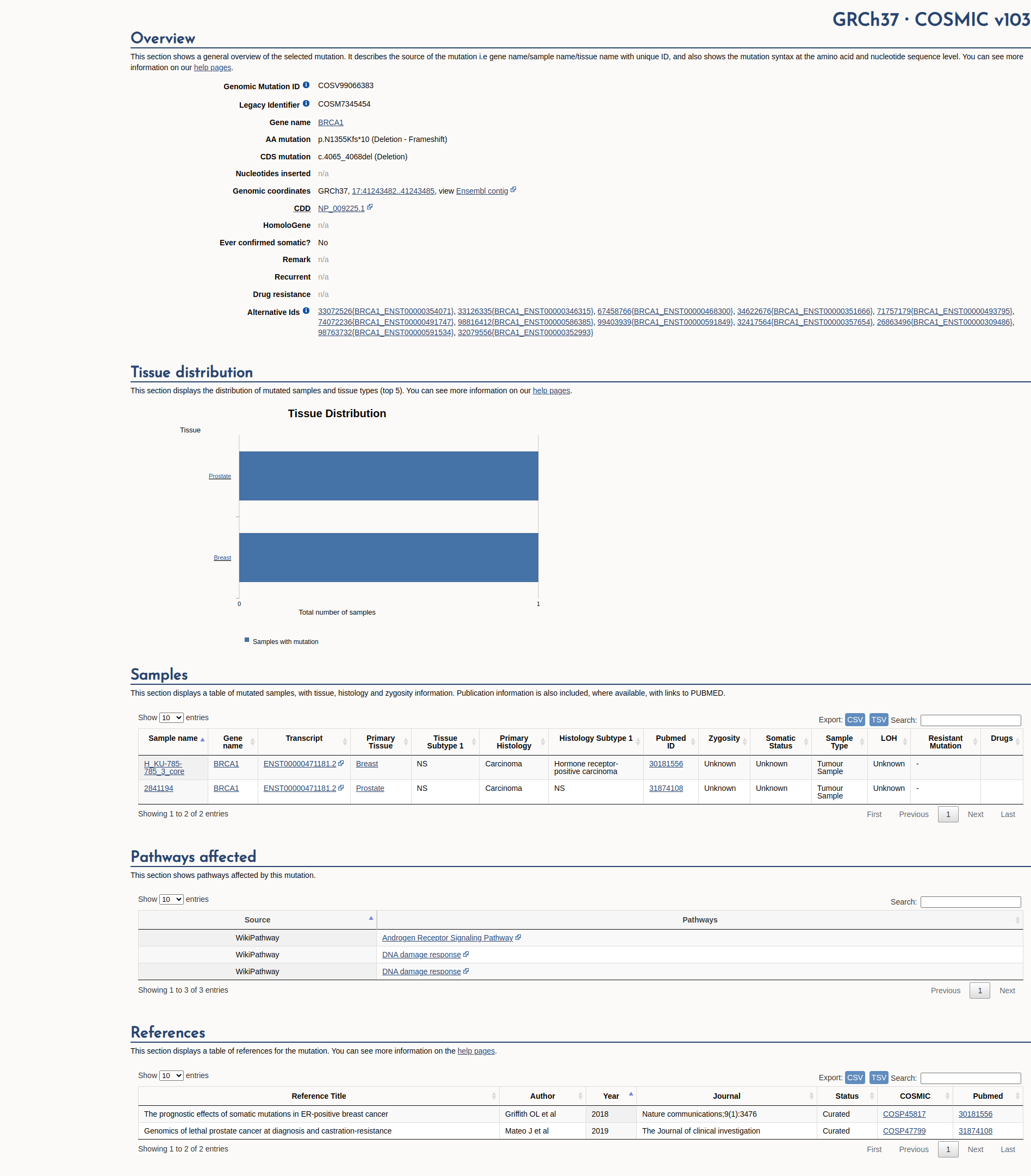
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: BRCA1N1355Kfs*10BRCA1N1355Kfs*10SomaticNCBI Gene:672|Show additional gene information Variant OverviewBRCA1, a tumor suppressor involved in the DNA damage response, is mutated in various cancer types.The BRCA1 N1355Kfs*10 is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in BRCA1 is: Truncating mutations in BRCA1 can lead to varying C-terminally truncated proteins that results in aberrant protein folding, contributing to loss of BRCA1 protein function. Human breast cancer cell lines that contain a truncating mutation in BRCA1 display elevated levels of aneuploidy and impaired DNA damage response. In addition, truncating mutations have been shown to induce aberrant protein localization, which may impact the interaction of important binding partners (PMID: 20608970). Mouse models of BRCA1 truncating mutations develop cancer, including mammary carcinomas, lymphomas and ovarian carcinomas (PMID: 11358863, 12483515, 12947386). JAX-CKB: No results found
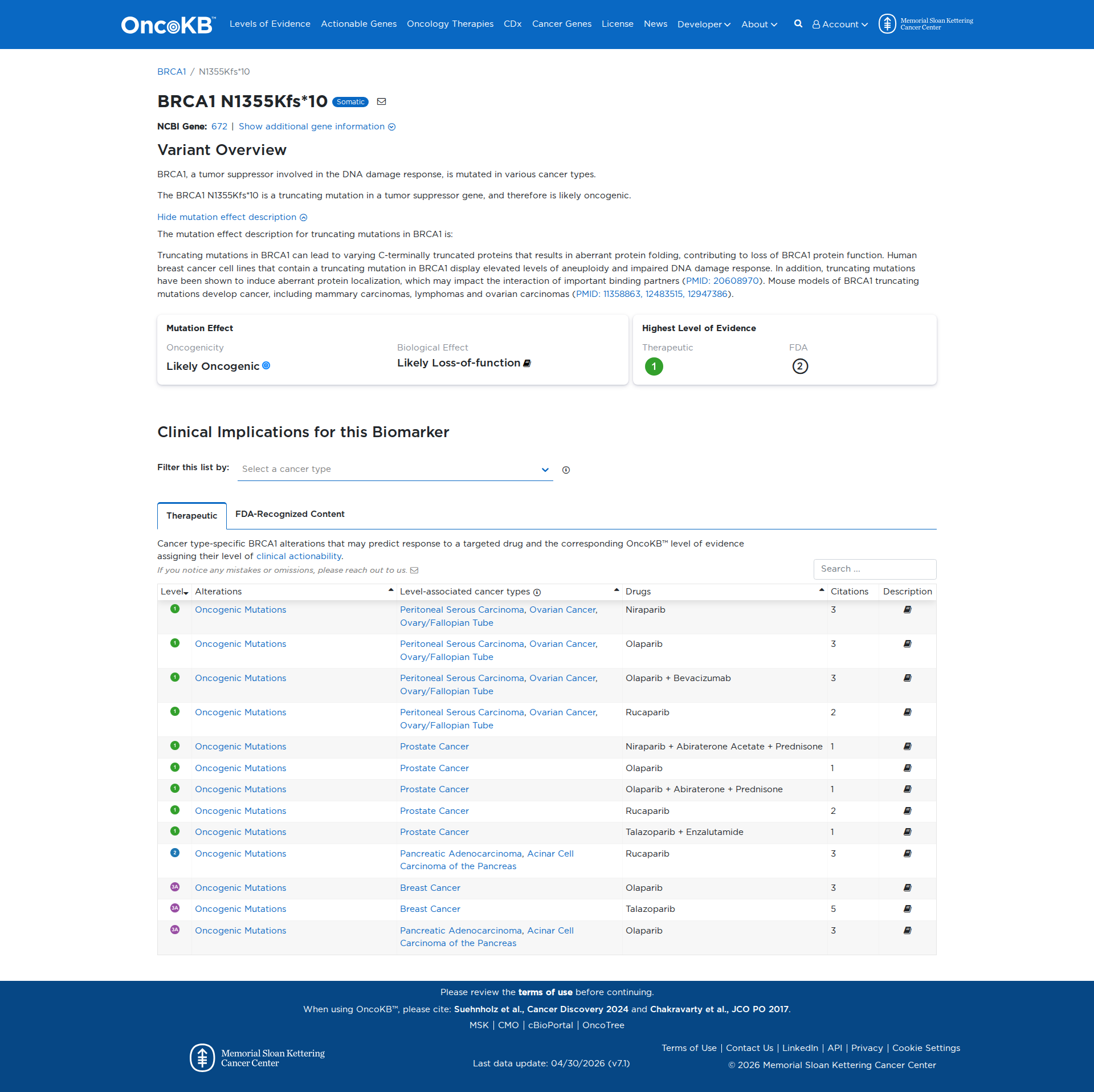
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.03 | 41 bp |
| Donor Loss (DL) | 0.14 | -27 bp |
| Acceptor Gain (AG) | 0.02 | 171 bp |
| Donor Gain (DG) | 0.0 | -40 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)