Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_001127208.2 | RefSeq Select | 9796 nt | 488–6496 |
| NM_001127208.3 | MANE Select | 9589 nt | 297–6305 |
| NM_001127208.1 | Alternative | 9677 nt | 387–6395 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
Open""
COSMIC Somatic Evidence
Open
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: TET2Q976*TET2Q976*SomaticNCBI Gene:54790|Show additional gene information Variant OverviewTET2, a tumor suppressor and DNA demethylase, is frequently mutated in hematologic malignancies.The TET2 Q976* is a truncating mutation in a tumor suppressor gene, and therefore is likely oncogenic.Hide mutation effect description The mutation effect description for truncating mutations in TET2 is: Truncating mutations of TET2 disrupt the C-terminal catalytic domain of the protein, predicted to cause gene inactivation and are considered oncogenic events (PMID: 21057493, 24315485). Consequently, these alterations abrogate TET2 enzymatic function to generate 5-hydroxymethylcytosine (5-hmC) (PMID: 21057493). JAX-CKB: No results found
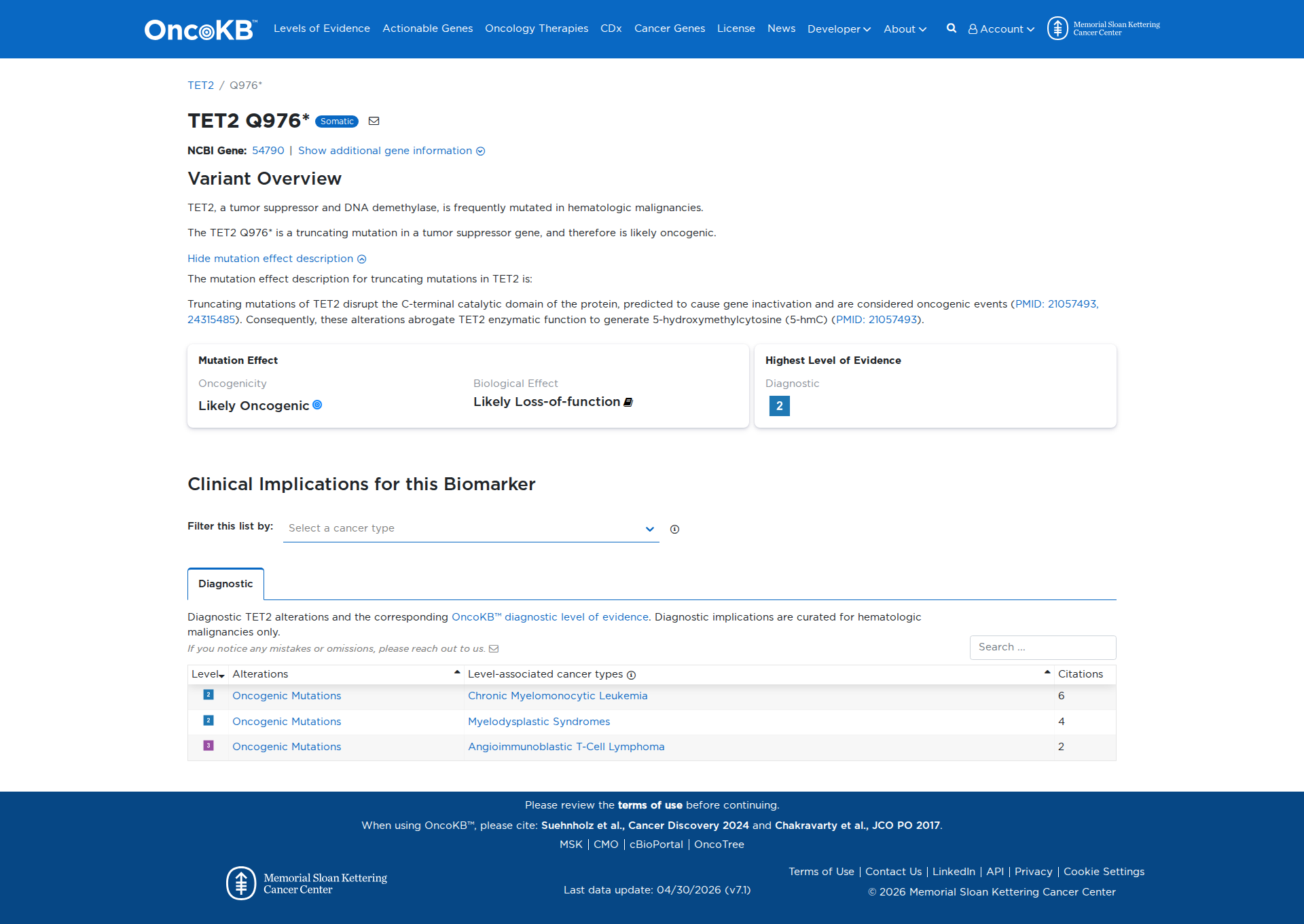
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.0 | -51 bp |
| Donor Loss (DL) | 0.01 | 483 bp |
| Acceptor Gain (AG) | 0.01 | -283 bp |
| Donor Gain (DG) | 0.01 | -173 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PVS1 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PS3 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)