Genetic Information
Gene & Transcript Details
| ID | Status | Details |
|---|---|---|
| NM_000051.3 | RefSeq Select | 13147 nt | 386–9556 |
| NM_000051.4 | MANE Select | 12915 nt | 151–9321 |
Variant Details
Clinical & Population Data
Population Frequency
gnomADClinVar
OpenThis missense variant replaces cysteine with tyrosine at codon 977 of the ATM protein. Computational prediction suggests that this variant may have deleterious impact on protein structure and function (internally defined REVEL score threshold >= 0.7, PMID: 27666373). To our knowledge, functional studies have not been performed for this variant. This variant has been reported in the compound heterozygous state in at least one family affected with ataxia-telangiectasia (communication with external laboratories; ClinVar SCV001812983.2, SCV000282917.8). This variant has also been reported in an individual affected with breast cancer (PMID: 31317629). This variant has not been identified in the general population by the Genome Aggregation Database (gnomAD). The available evidence is insufficient to determine the role of this variant in disease conclusively. Therefore, this variant is classified as a Variant of Uncertain Significance.
The p.C977Y variant (also known as c.2930G>A), located in coding exon 19 of the ATM gene, results from a G to A substitution at nucleotide position 2930. The cysteine at codon 977 is replaced by tyrosine, an amino acid with highly dissimilar properties. This variant has been confirmed in trans with an ATM pathogenic mutation in multiple, related individuals with clinical features of ataxia telangiectasia (external correspondence from GeneDx). In addition, this variant co-segregated with disease in one family tested in an external laboratory. Another alteration at the same codon, p.C977R (c.2929T>C) has been described in an individual diagnosed with ataxia telangiectasia (Galatolo D et al. Int J Mol Sci, 2021 Aug;22(16):8490). This amino acid position is highly conserved in available vertebrate species. In addition, this alteration is predicted to be deleterious by in silico analysis. Based on the majority of available evidence to date, this variant is likely to be pathogenic.
This sequence change replaces cysteine, which is neutral and slightly polar, with tyrosine, which is neutral and polar, at codon 977 of the ATM protein (p.Cys977Tyr). The frequency data for this variant in the population databases is considered unreliable, as metrics indicate poor data quality at this position in the gnomAD database. This missense change has been observed in individual(s) with ataxia-telangiectasia and/or pancreatic cancer (PMID: 35047863; internal data). In at least one individual the data is consistent with being in trans (on the opposite chromosome) from a pathogenic variant. It has also been observed to segregate with disease in related individuals. ClinVar contains an entry for this variant (Variation ID: 233765). Invitae Evidence Modeling of protein sequence and biophysical properties (such as structural, functional, and spatial information, amino acid conservation, physicochemical variation, residue mobility, and thermodynamic stability) has been performed for this missense variant. However, the output from this modeling did not meet the statistical confidence thresholds required to predict the impact of this variant on ATM protein function. This variant disrupts the p.Cys977 amino acid residue in ATM. Other variant(s) that disrupt this residue have been observed in individuals with ATM-related conditions (PMID: 34445196), which suggests that this may be a clinically significant amino acid residue. In summary, the currently available evidence indicates that the variant is pathogenic, but additional data are needed to prove that conclusively. Therefore, this variant has been classified as Likely Pathogenic.
The ATM c.2930G>A (p.Cys977Tyr) variant has been reported in the published literature in individuals affected with breast cancer (PMID: 31317629 (2019)) and pancreatic cancer (PMID: 35047863 (2022)). This variant has been reported in individuals with clinical features of ataxia telangiectasia, and were compound heterozygous for the variant and a pathogenic or likely pathogenic variant (Invitae, personal communication regarding ClinVar ID 233765). Additionally, the variant was reported to segregate with disease in at least one family (Invitae, personal communication regarding ClinVar ID 233765). The frequency of this variant in the general population, 0.000034 (1/29448 chromosomes (Genome Aggregation Database, http://gnomad.broadinstitute.org)), is consistent with pathogenicity. Analysis of this variant using bioinformatics tools for the prediction of the effect of amino acid changes on protein structure and function yielded predictions that this variant is damaging. Based on the available information, this variant is classified as likely pathogenic.
Variant summary: ATM c.2930G>A (p.Cys977Tyr) results in a non-conservative amino acid change in the encoded protein sequence. Algorithms developed to predict the effect of missense changes on protein structure and function are either unavailable or do not agree on the potential impact of this missense change. The variant allele was found at a frequency of 2.5e-06 in 1602390 control chromosomes in the gnomAD database (v4.1 dataset). c.2930G>A has been observed in heterozygous state in individuals affected with ATM-related cancer phenotypes (e.g. Amendola_2019, Yu_2022), and in compound heterozygous state in individuals with ataxia-telangiectasia (Labcorp Genetics (formerly Invitae), internal data). These data indicate that the variant is likely to be associated with disease. To our knowledge, no experimental evidence demonstrating an impact on protein function has been reported. The following publications have been ascertained in the context of this evaluation (PMID: 31317629, 35047863). ClinVar contains an entry for this variant (Variation ID: 233765). Based on the evidence outlined above, the variant was classified as likely pathogenic.
"This variant has been reported in ClinVar as Likely pathogenic (4 clinical laboratories) and as Uncertain significance (1 clinical laboratories) and as likely pathogenic (1 clinical laboratories)."
COSMIC Somatic Evidence
Open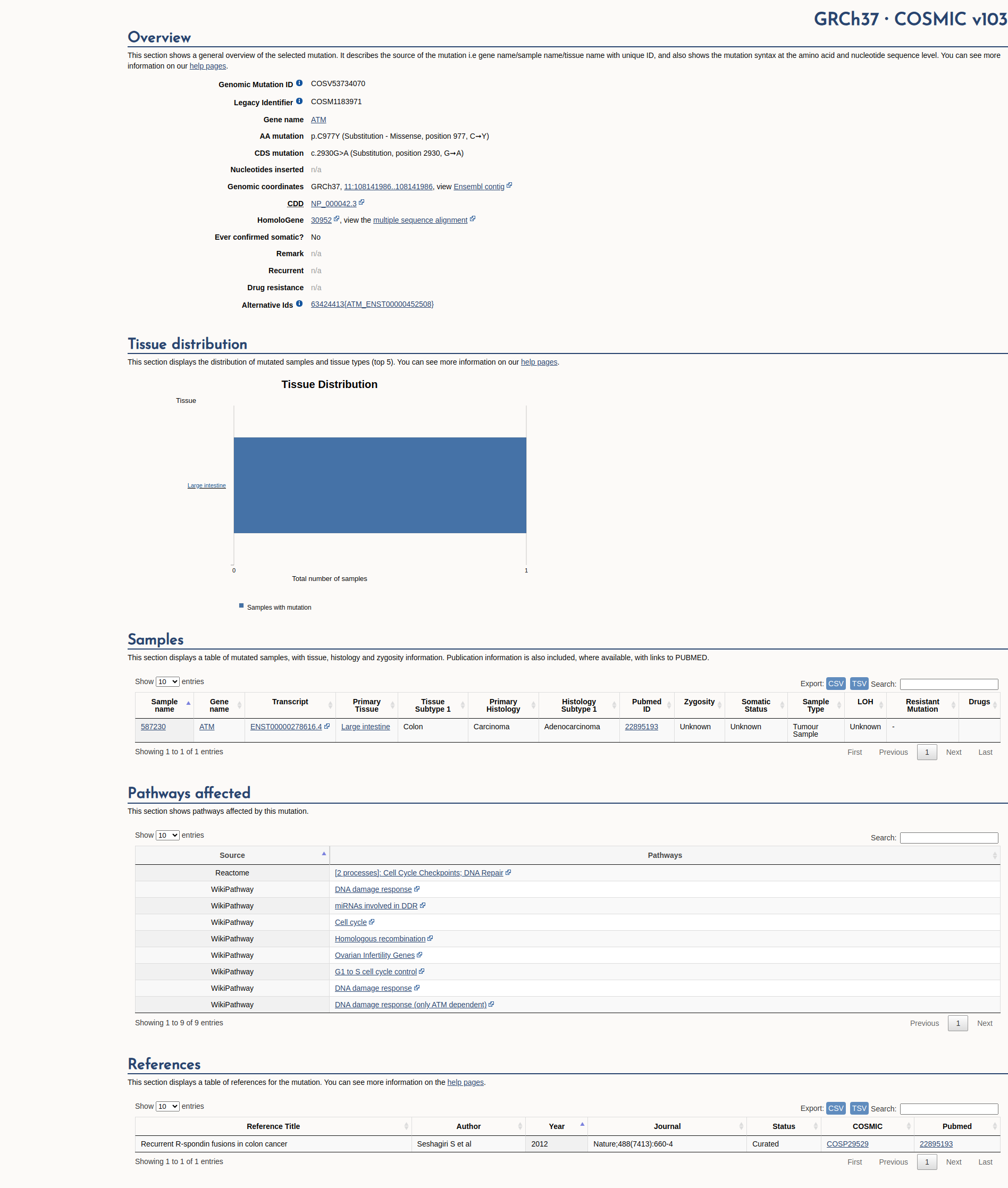
Functional Impact & Domains
Functional Domain
Error in OpenAI Consolidation. OncoKB: ATMC977YATMC977YSomaticNCBI Gene:472|Show additional gene information Variant OverviewATM, a kinase involved in the DNA damage response, is mutated in various solid and hematologic malignancies.The ATM C977Y mutation has not specifically been reviewed by the OncoKB team, and therefore its biological significance is unknown. JAX-CKB: No results found
Click on previews to view full database entries. External databases may require institutional access.
Computational Analysis
Pathogenicity Predictions
SpliceAISpliceAI Scores
Window: ±500bp| Effect Type | Score | Position |
|---|---|---|
| Acceptor Loss (AL) | 0.02 | 63 bp |
| Donor Loss (DL) | 0.0 | -288 bp |
| Acceptor Gain (AG) | 0.0 | -195 bp |
| Donor Gain (DG) | 0.01 | -113 bp |
VCEP Guidelines
Applied ACMG/AMP Criteria (VCEP Specific)
PM2 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
PP5 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)
BP4 (Unknown (Pre-LLM))
From pre-LLM assessment (LLM Failed)