Classification rationale
1
The PALB2 VCEP specification defines BA1 at gnomAD v4 grpmax filtering AF >0.1%, BS1 at >0.01%, and BP1 for all missense variants.
cspec ↗2
In the assembled workspace, gnomAD v4 reports grpmax_faf 0.00602134 (~0.602%), which exceeds both the BA1 and BS1 PALB2 thresholds by a wide margin.
gnomad_v4 ↗3
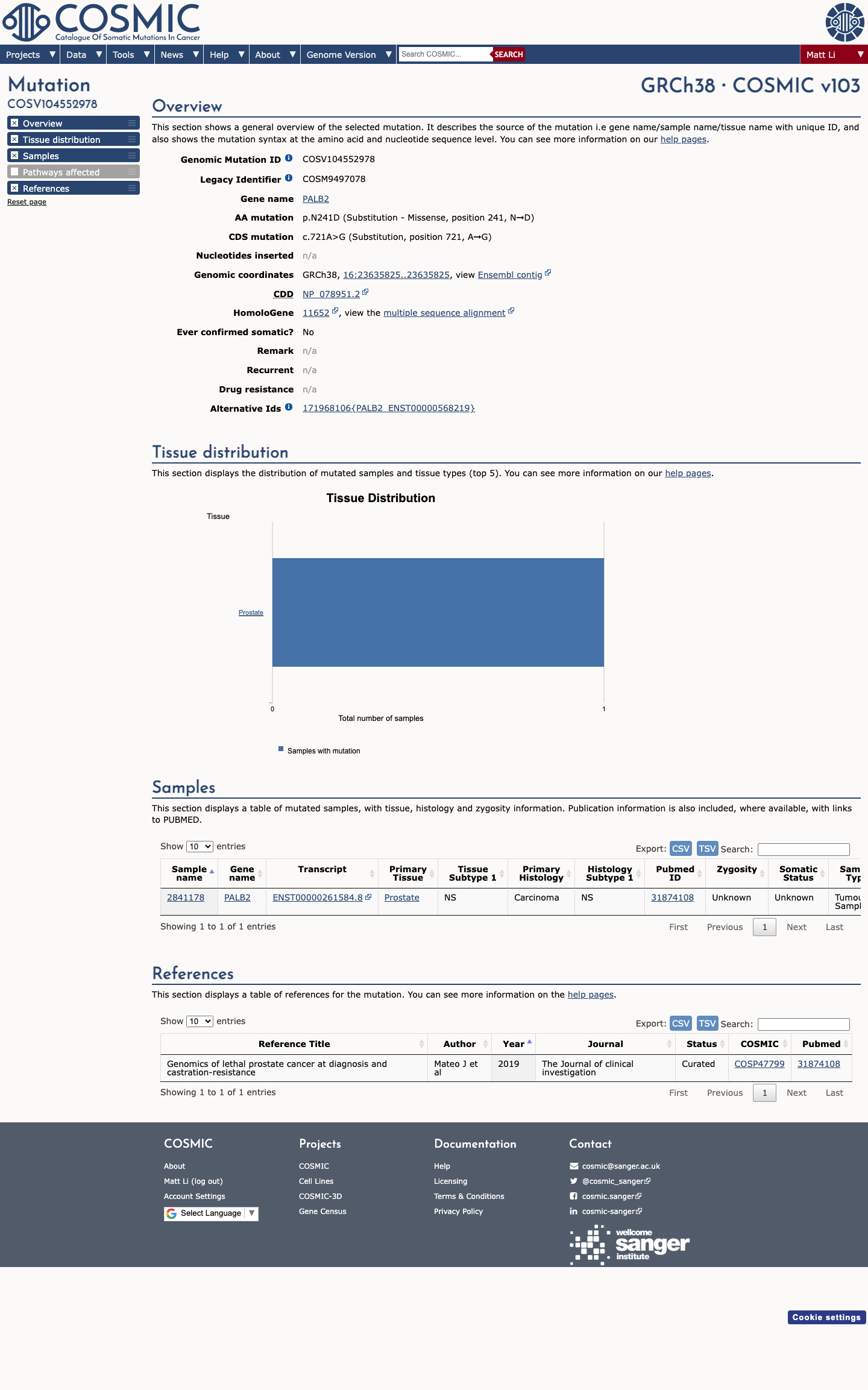
The variant is a missense substitution p.(Asn241Asp)/p.(N241D), so PALB2-specific BP1 also applies; ClinVar expert-panel benign classification is concordant but was not needed as an ACMG criterion because BA1 alone is sufficient for a benign call.
clinvar ↗