The PTEN-specific PVS1 decision tree supports PVS1_Strong for this canonical splice acceptor variant because it affects the exon 9 acceptor of NM_000314.8, and preserved-frame loss of entire exon 9 is explicitly treated as a critical altered region in PTEN.
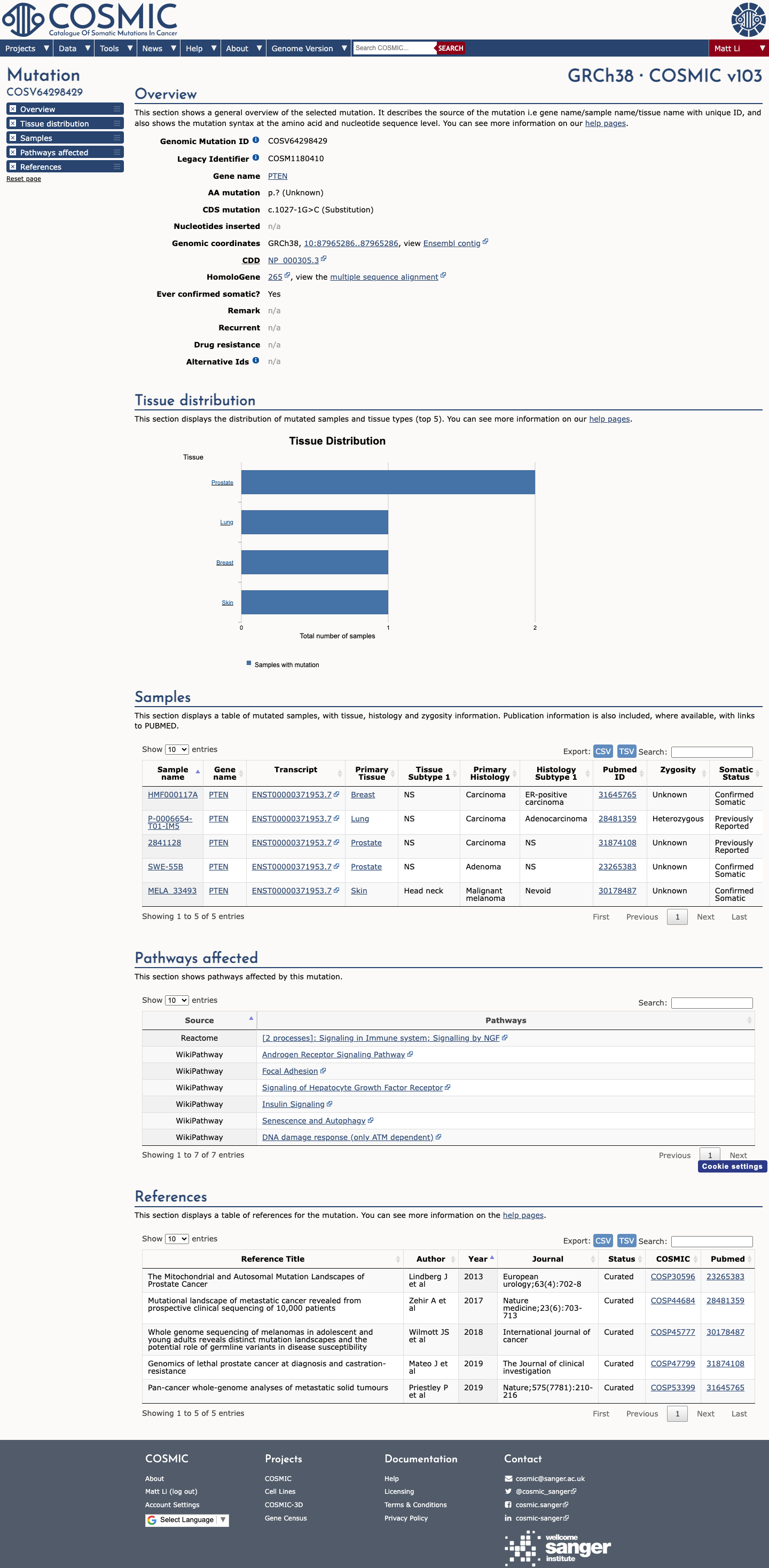
PS1_Strong is met because ClinVar contains a same-nucleotide pathogenic splicing variant at NM_000314.8:c.1027-1G>A.
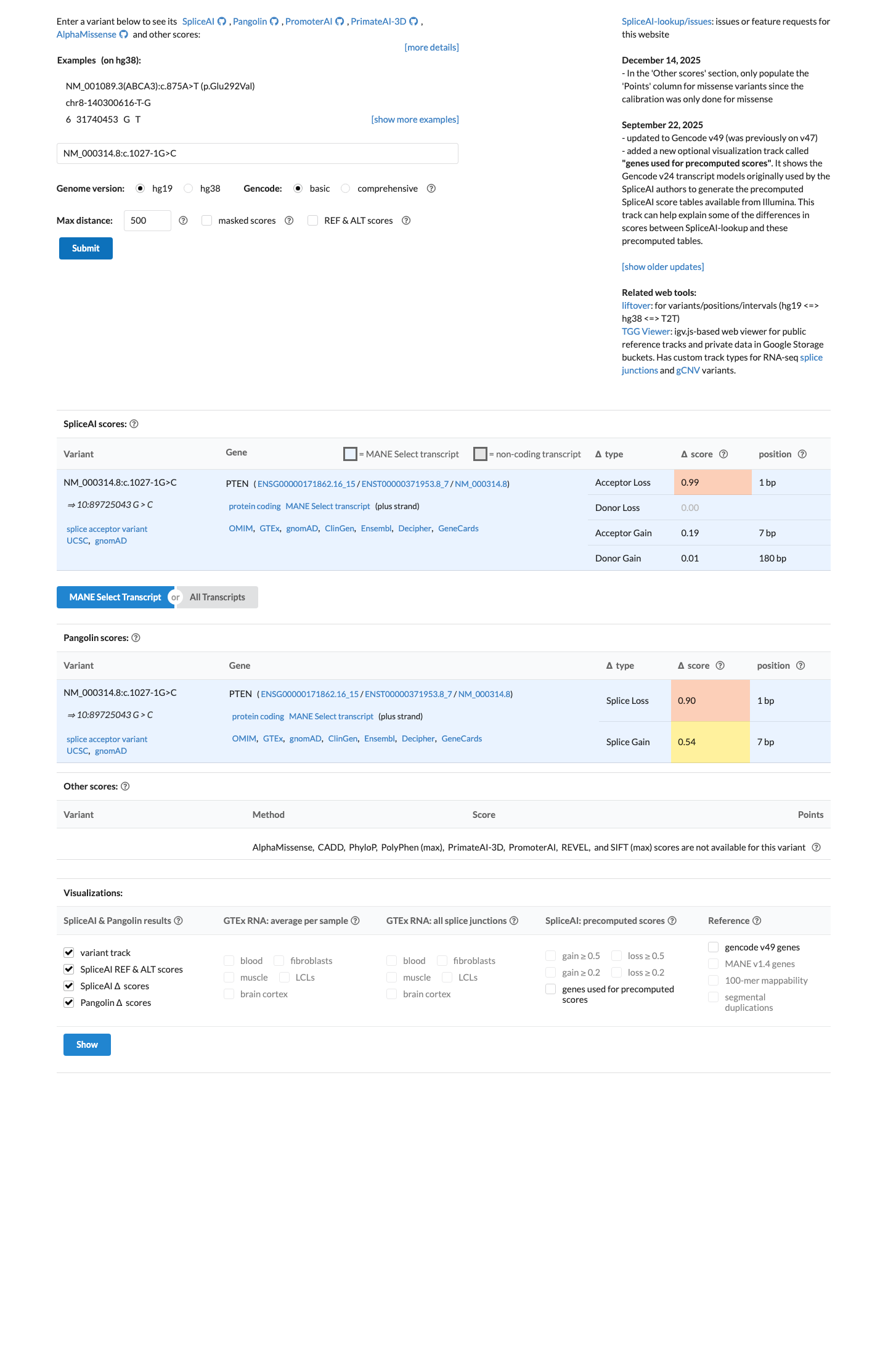
clinvar ↗The queried variant shows equal predicted splice acceptor loss to the known pathogenic same-position variant by SpliceAI (DS_AL 0.99), satisfying the PTEN same-position splicing use of PS1.
spliceai ↗PM2_Supporting is met because the variant is absent from both gnomAD v2.1 and gnomAD v4.1.
gnomad_v4 ↗No assembled RNA assay, de novo series, segregation dataset, or PTEN PS4 phenotype-scored case series was available to upgrade beyond the current evidence combination.
cspec ↗