Classification rationale
1
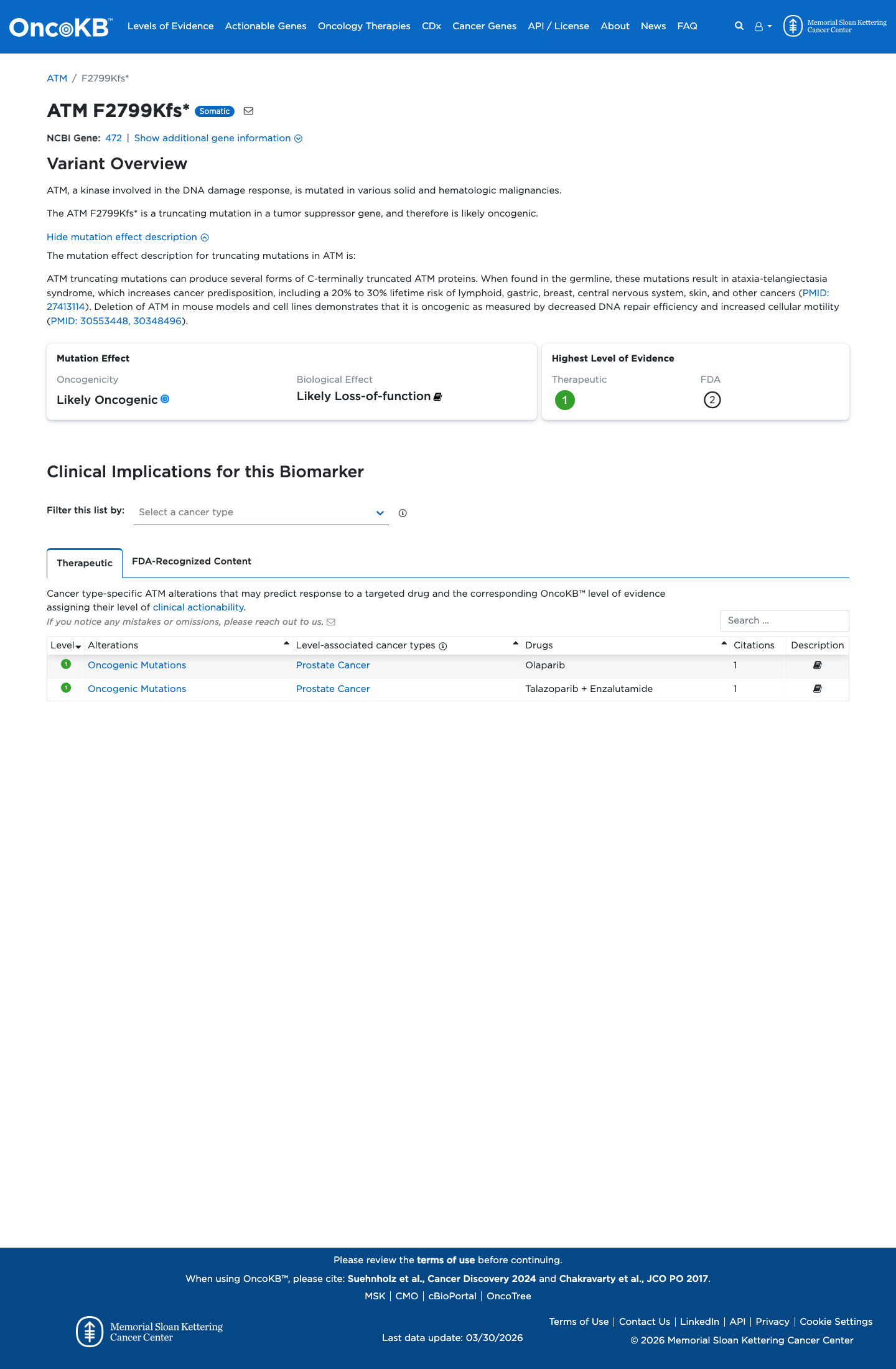
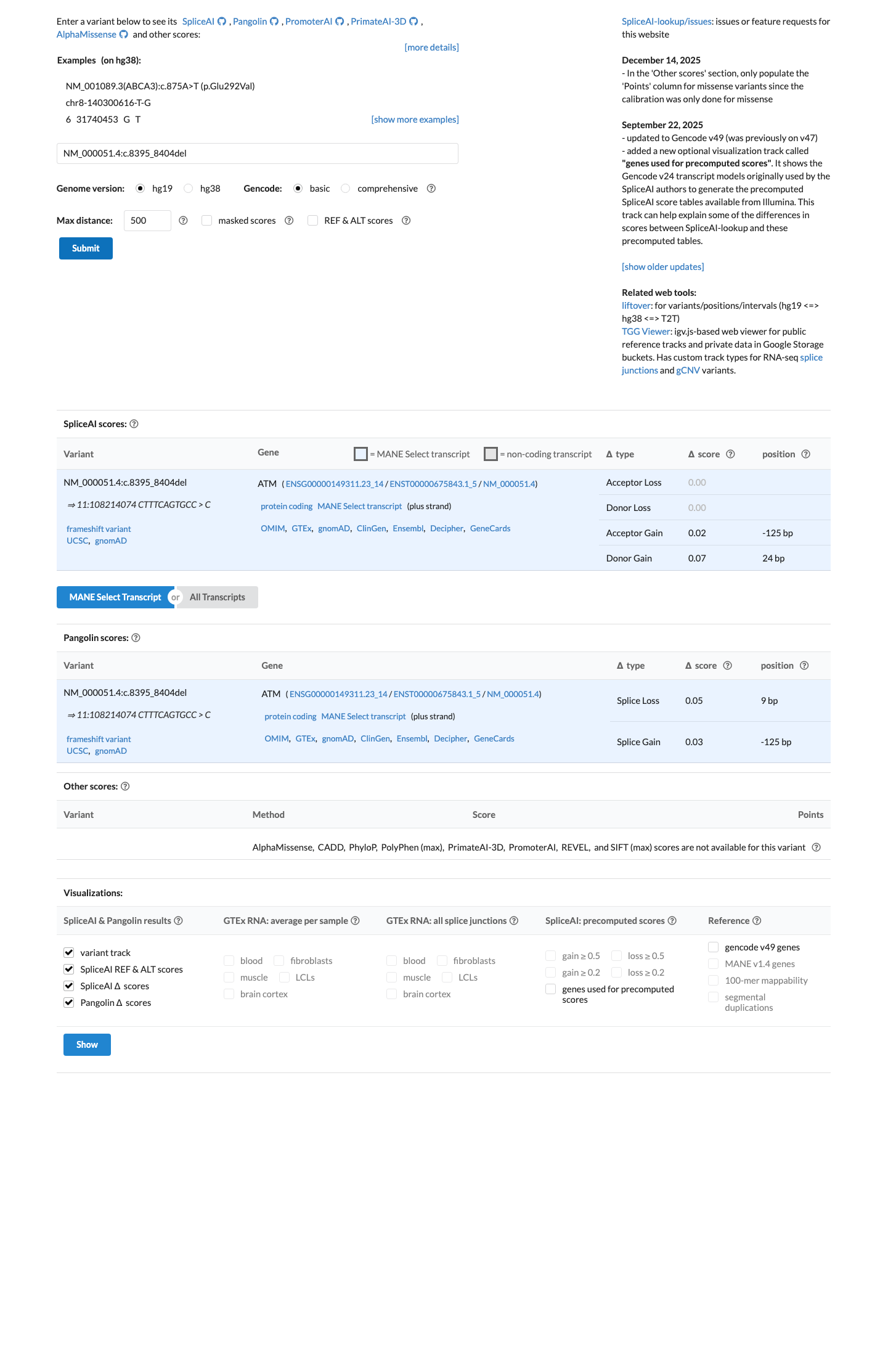
ATM HBOP VCEP v1.5 supports PVS1 for null variants through the ATM PVS1 decision tree, and this variant is an exonic frameshift with predicted consequence p.(Phe2799LysfsTer4)/p.(F2799Kfs*4).
cspec ↗2
ATM-specific PM5_Supporting applies to truncating variants with premature termination codons upstream of p.Arg3047; this variant truncates at codon 2802, upstream of that boundary.
cspec ↗3
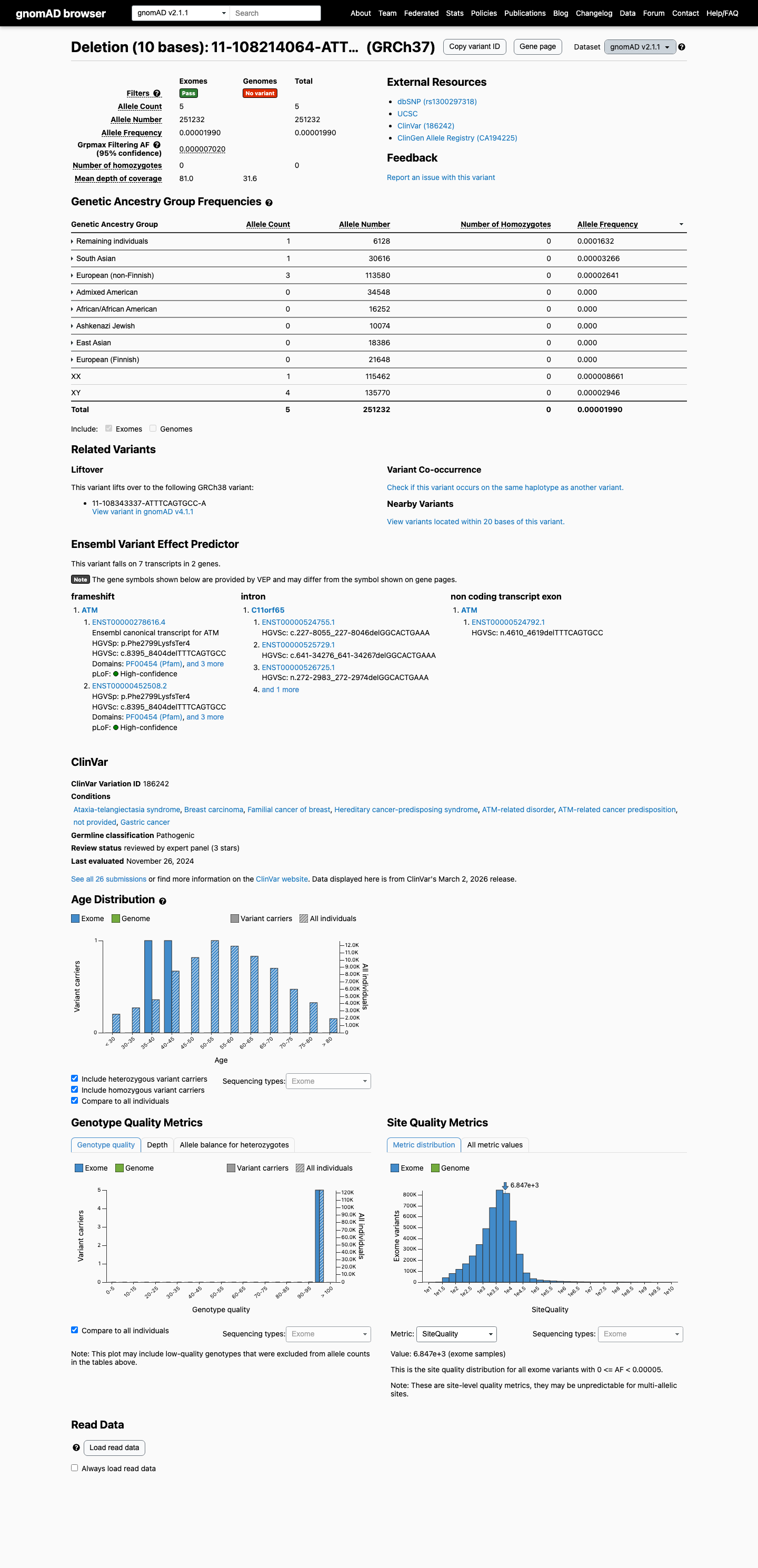
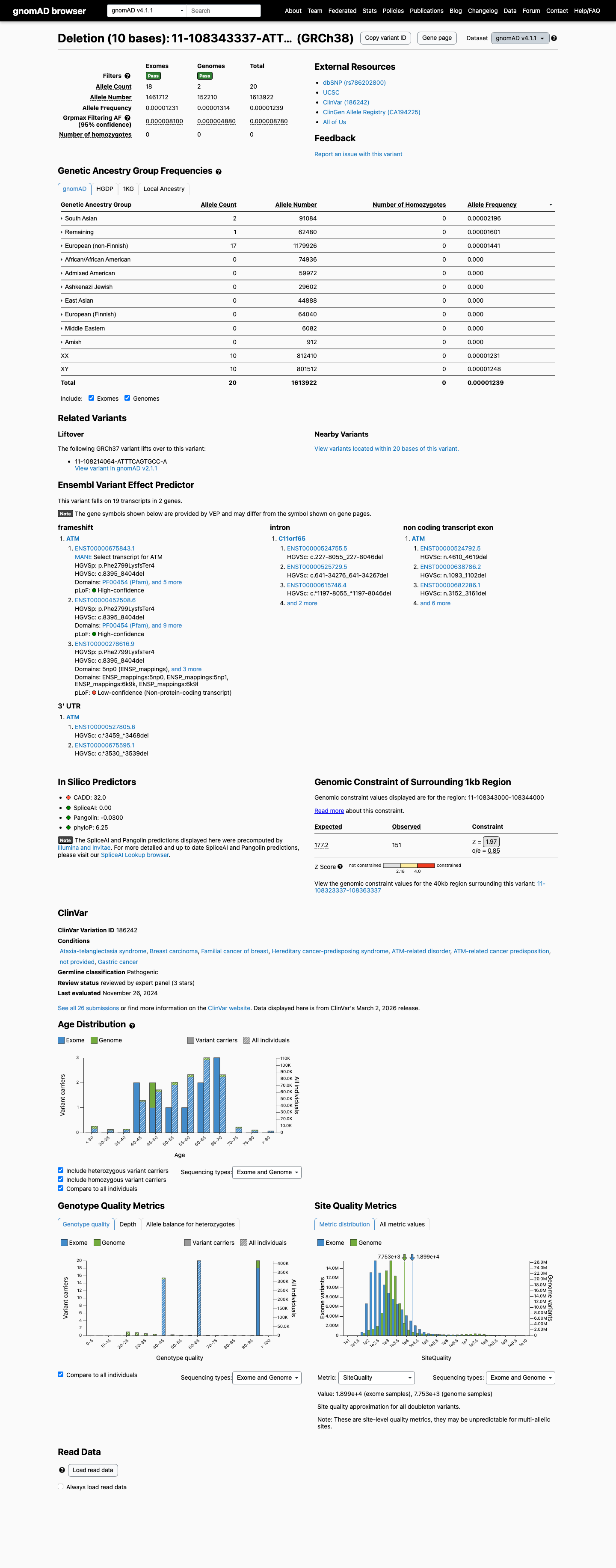
PM2 was not applied because gnomAD v4 shows 20/1613922 alleles (0.00124%), slightly above the ATM PM2_Supporting threshold of 0.001%.
gnomad_v4 ↗4
ClinVar contains a concordant ClinGen HBOP expert-panel Pathogenic classification for this variant, but that assertion was treated as contextual support rather than as a standalone ACMG criterion.
clinvar ↗