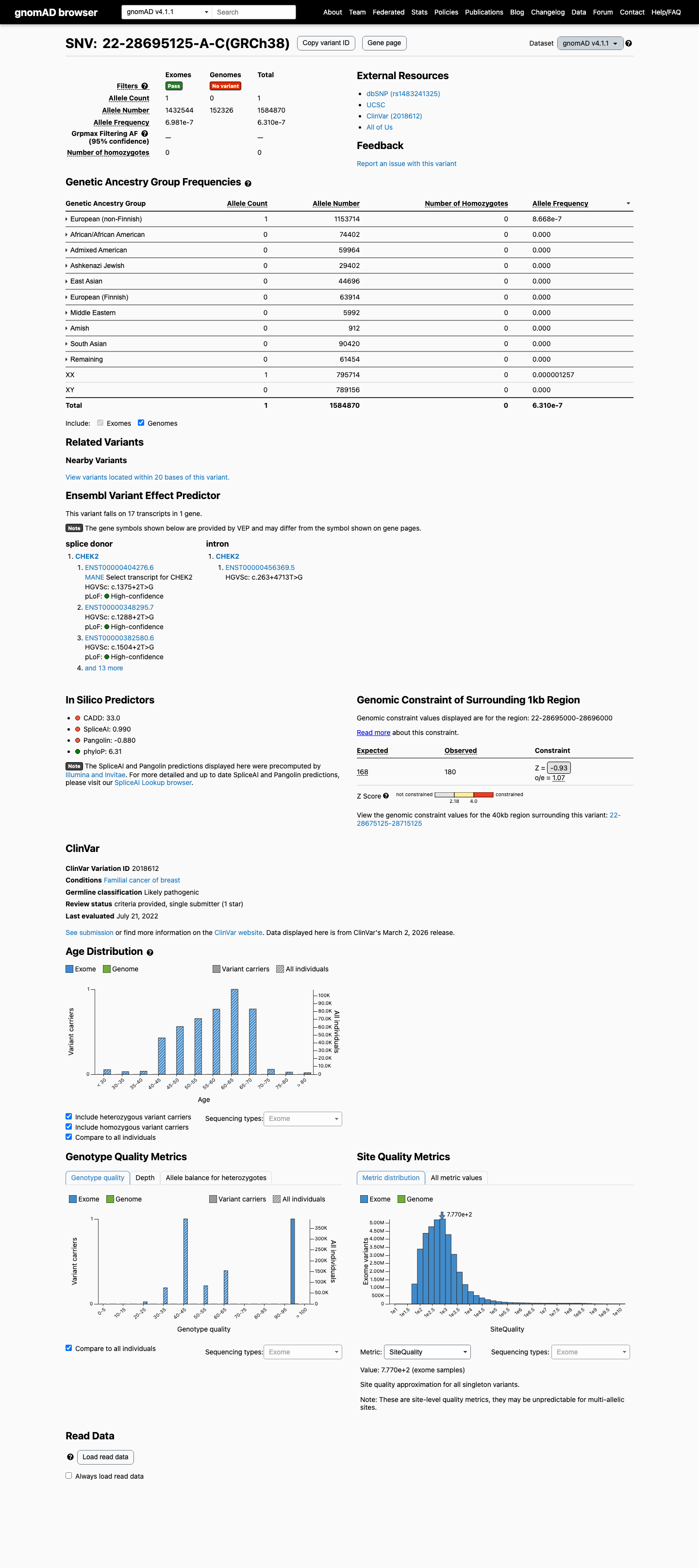
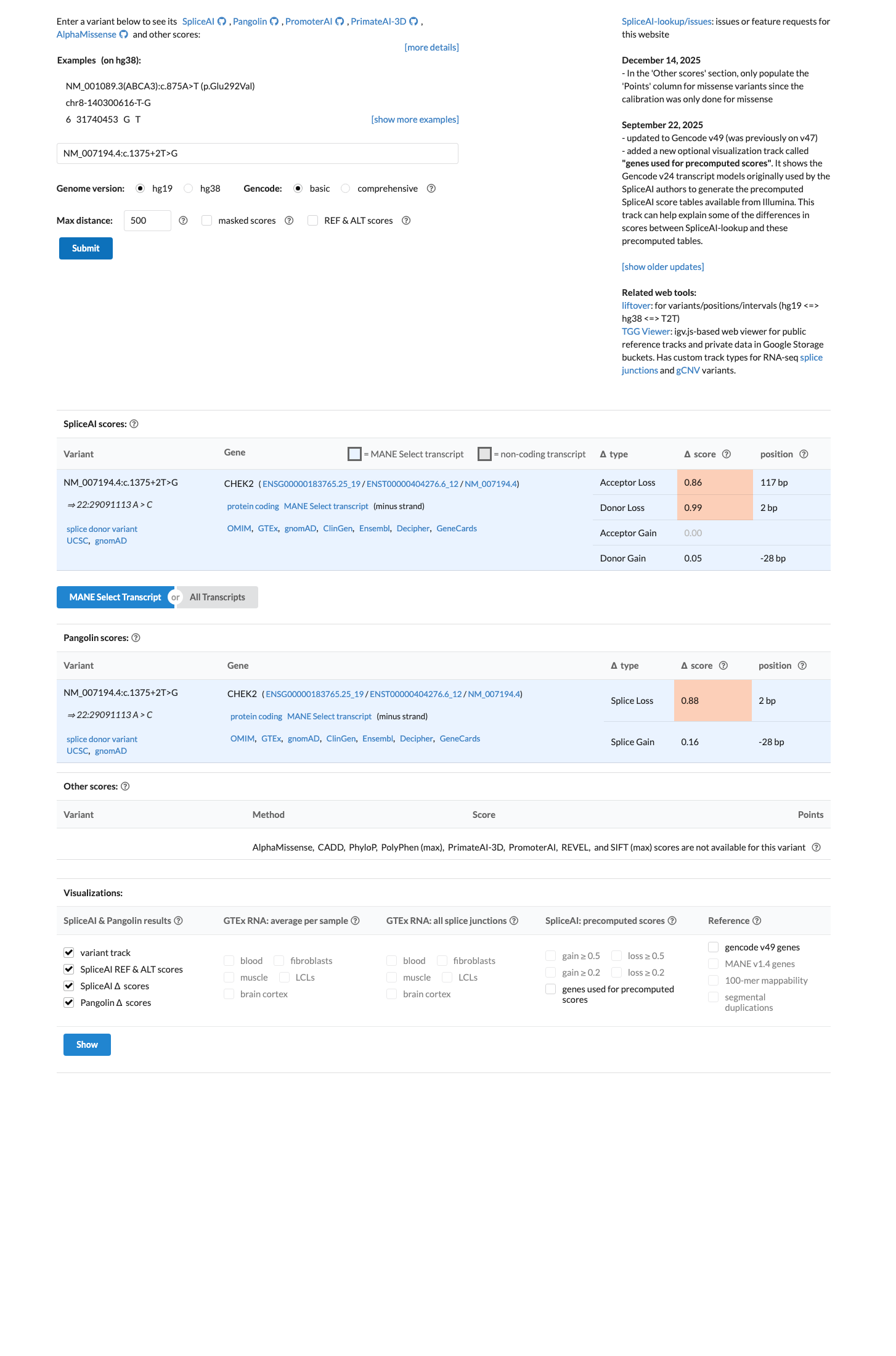
The variant affects the canonical +2 donor position of CHEK2 and is expected to disrupt normal splicing, which supports PVS1 in a loss-of-function disease mechanism context.
spliceai ↗The reviewed CHEK2 framework mode is VCEP and the case workspace points to the CHEK2 ClinGen specification as the primary interpretive authority, although the local criterion table was not populated.
cspec ↗Population data support rarity: the variant is absent from gnomAD v2.1 and observed only once in gnomAD v4.1 with no homozygotes, supporting PM2 at supporting strength.
gnomad_v4 ↗No convincing variant-specific case-control, segregation, de novo, or functional RNA evidence for this exact allele was identified in the reviewed workspace, so no additional criteria were added.
Using ACMG/AMP combining rules, PVS1 plus PM2_Supporting supports a Likely Pathogenic classification.
cspec ↗