Classification rationale
1
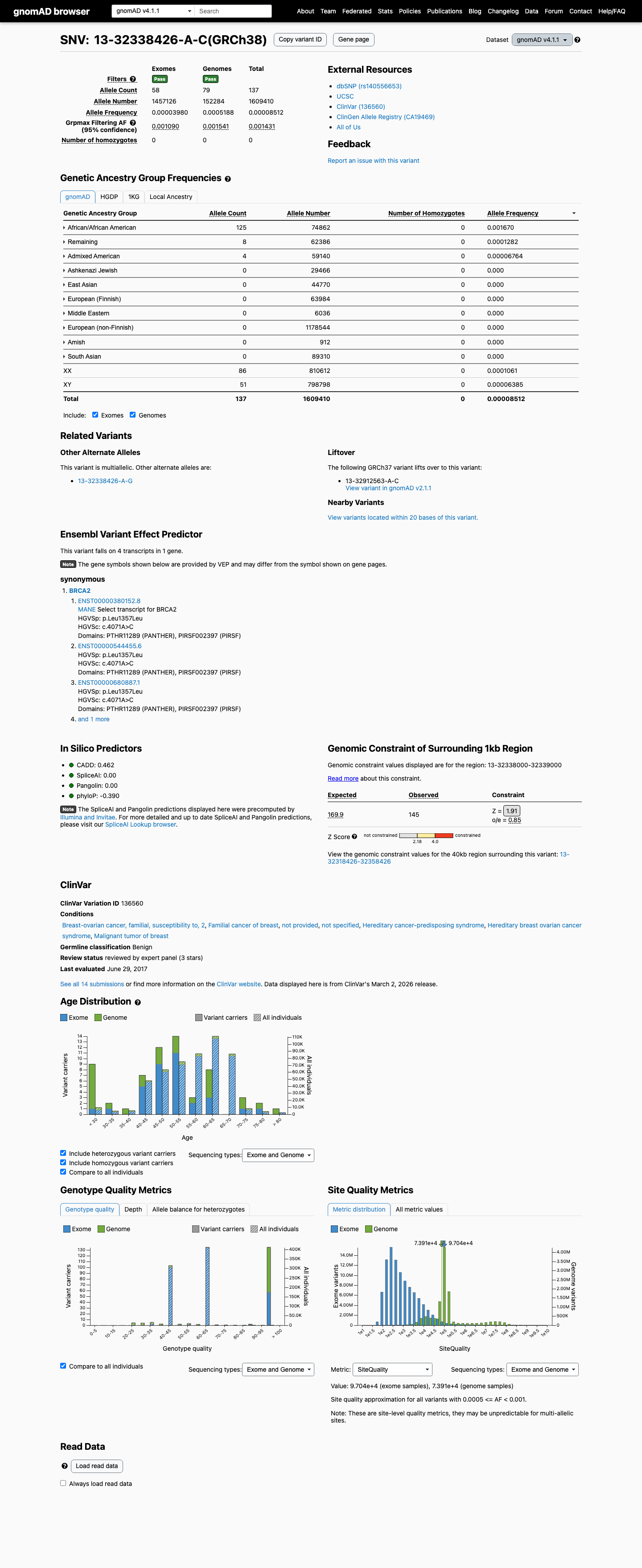
BRCA2 ENIGMA BA1 is met because the variant exceeds the stand-alone benign frequency threshold in gnomAD: v2.1 grpmax FAF 0.00161392 and v4.1 grpmax FAF 0.00154091, both >0.001.
gnomad_v2 ↗2
The variant is synonymous, p.(Leu1357=), and lies outside the BRCA2 clinically important domains defined by ENIGMA; with SpliceAI max delta score 0.00, this satisfies BP1_Strong.
cspec ↗3
ClinVar shows a concordant benign expert-panel assertion from ENIGMA, which supports but does not replace the VCEP rule-based classification.
clinvar ↗