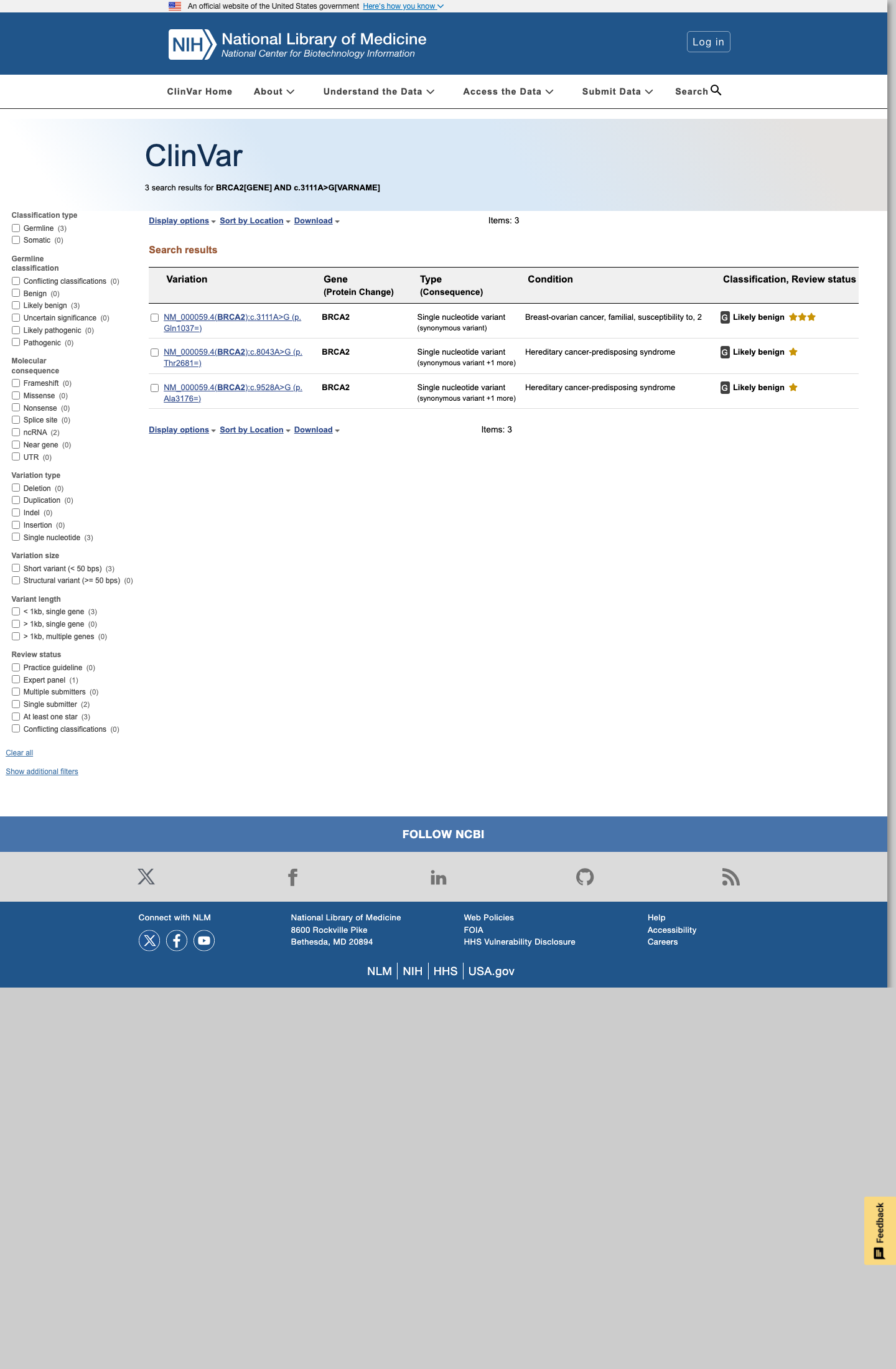
NM_000059.4:c.3111A>G normalizes to the synonymous BRCA2 protein consequence p.(Gln1037=) / p.(Q1037=).
cspec ↗Q1037 is outside the BRCA2 clinically important domains defined by the specification (PALB2-binding aa 10-40 and DNA-binding aa 2481-3186).
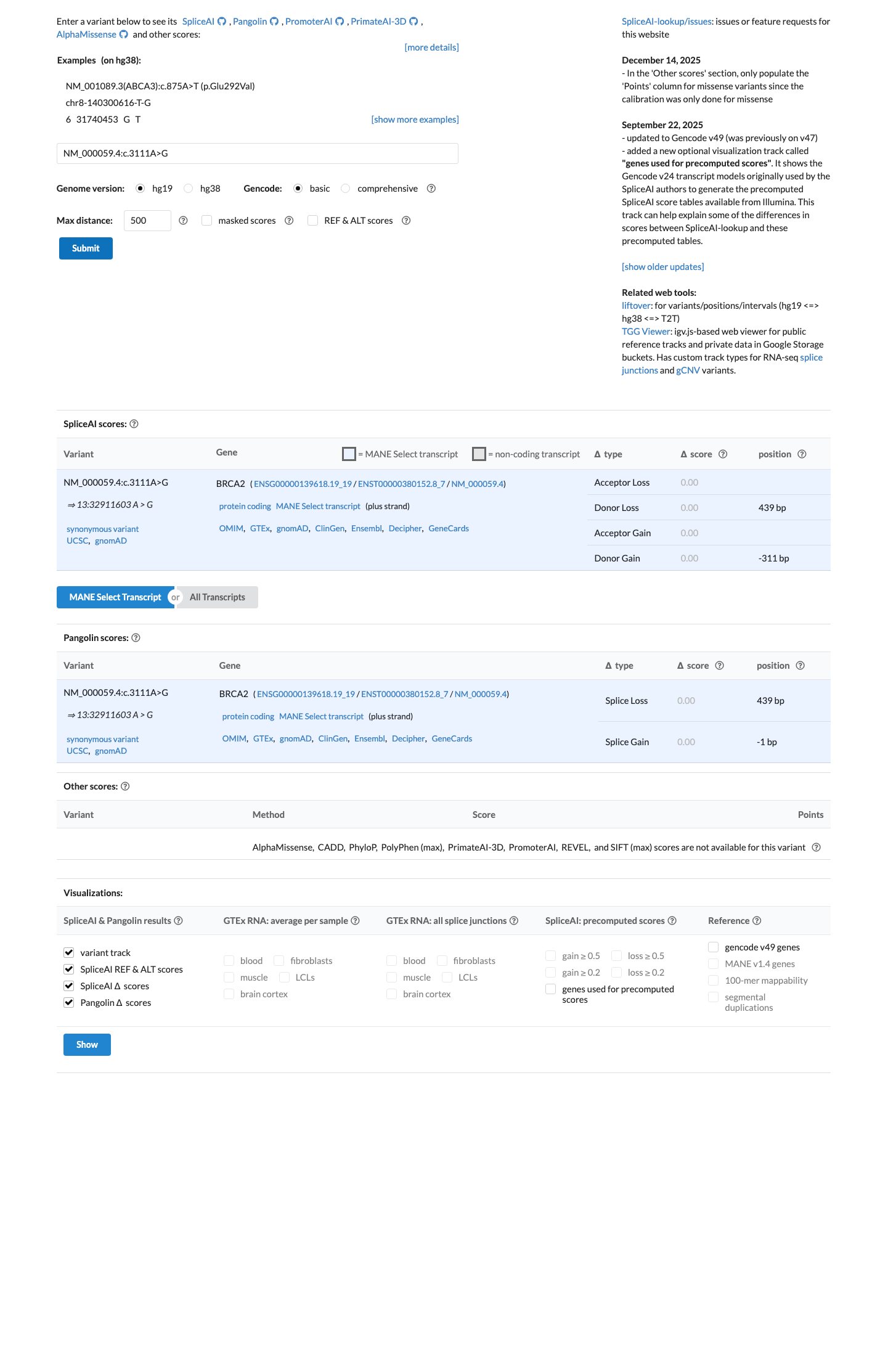
SpliceAI predicts no significant splice impact for this variant, with max delta score 0.00, satisfying the no-splicing-predicted part of BP1_Strong.
spliceai ↗The BRCA2 ENIGMA specification states to apply BP1_Strong for silent variants outside clinically important domains with SpliceAI ≤0.1, and its Table 3 states Likely Benign can be assigned from one Strong benign code when multiple evidence types contribute; BP1_Strong is given as an explicit example.
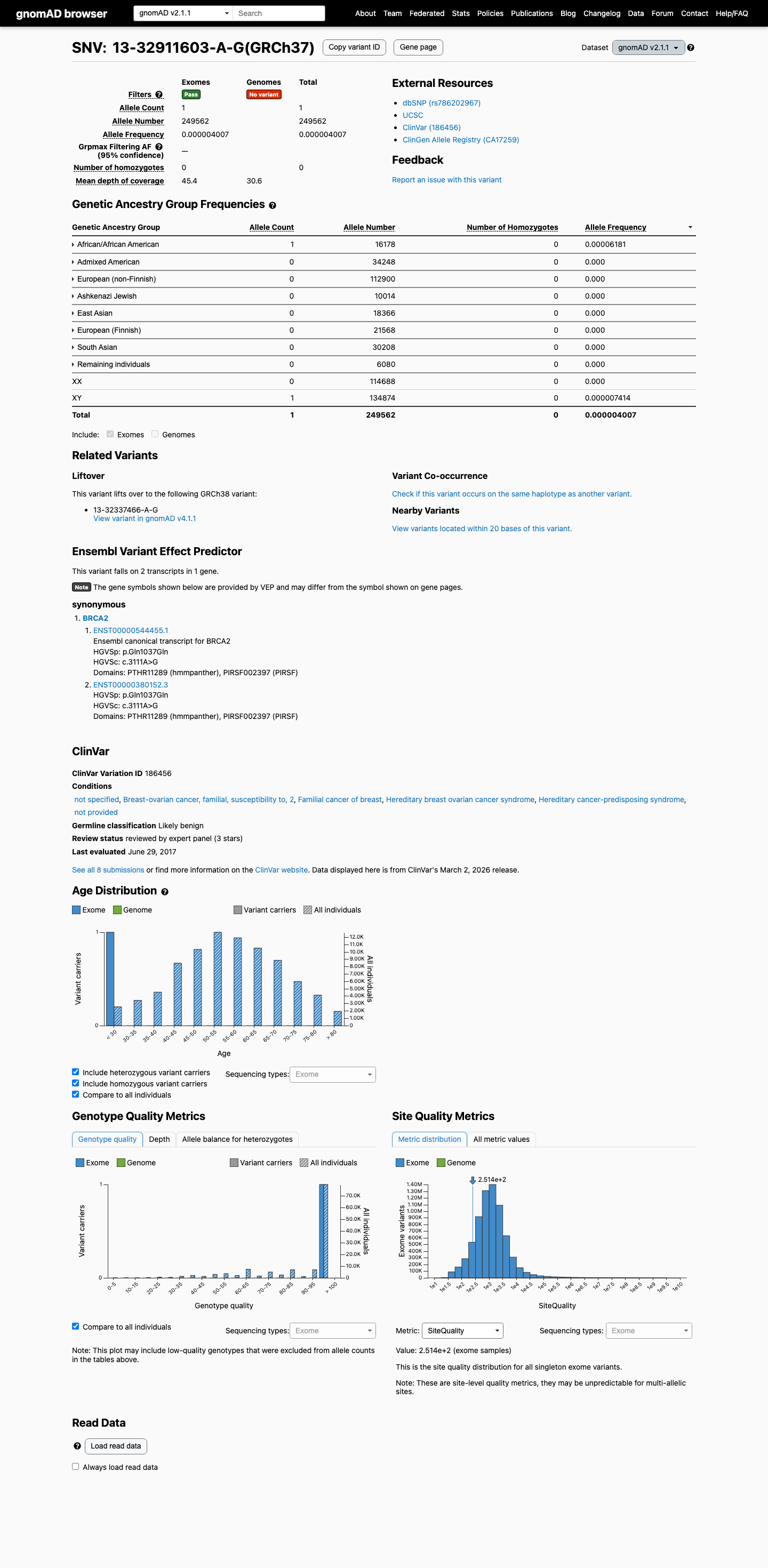
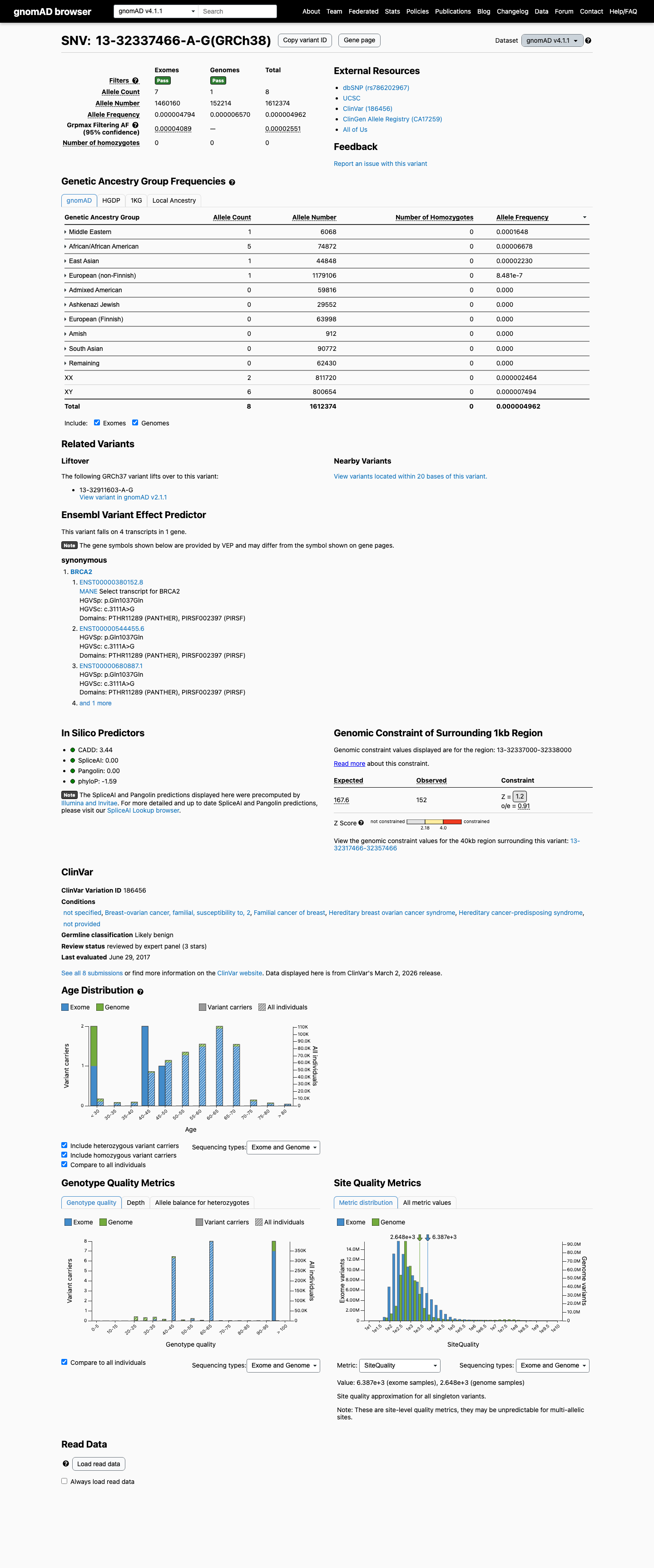
Population data do not support BA1 or BS1, and the variant is not absent from controls, so PM2 is also not met; these findings do not overturn the BP1_Strong-based Likely Benign classification.
gnomad_v4 ↗