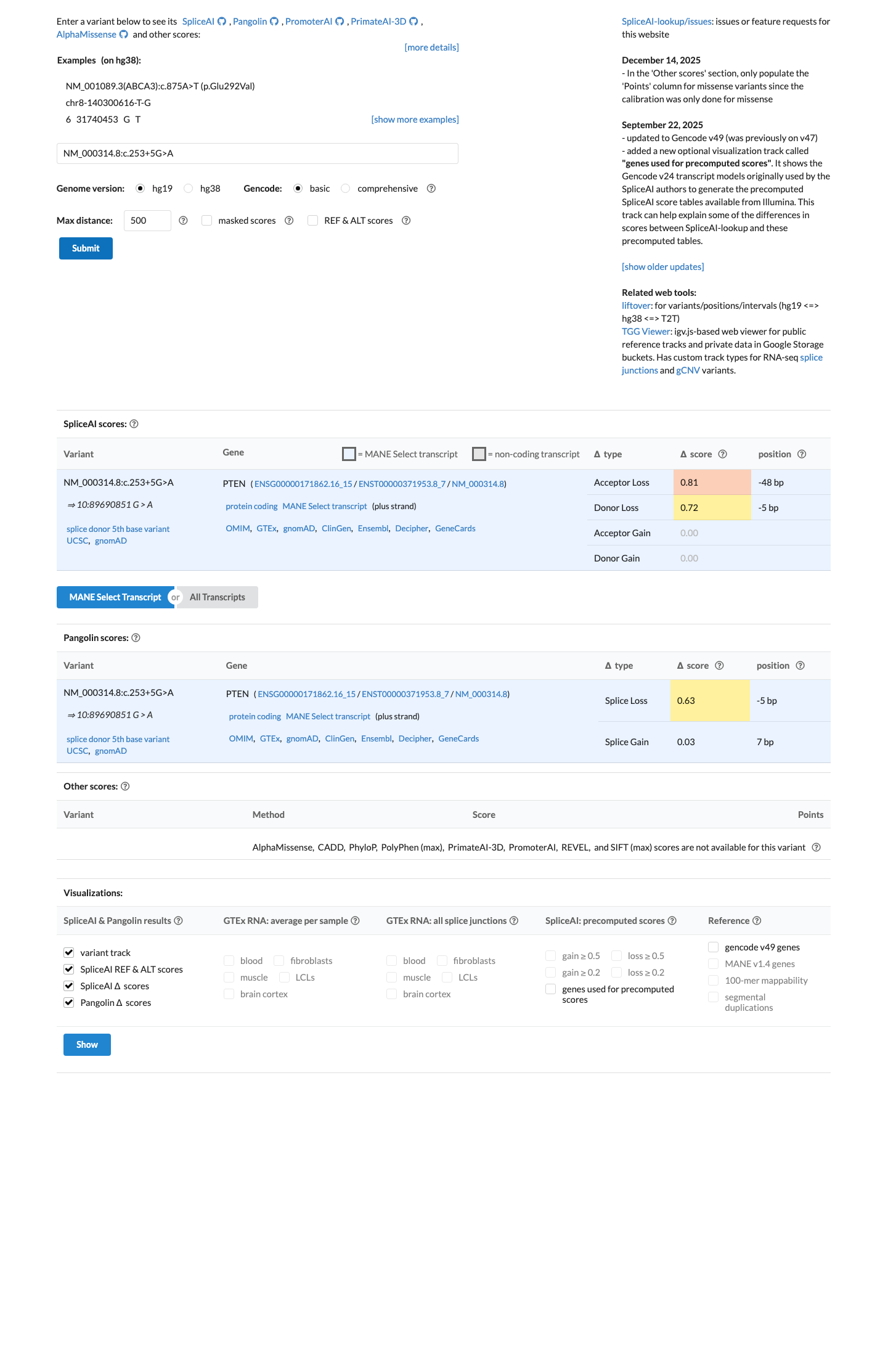
An RNA study has shown that NM_000314.8:c.253+5G>A causes exon 4 skipping, resulting in the predicted frameshift transcript consequence p.(Ala72Thrfs*5).
PMID:28677221 ↗Under the PTEN-specific PVS1 decision tree, a splice alteration that disrupts the reading frame in biologically relevant transcript NM_000314.8 and lies 5′ to p.D375 (c.1121) meets PVS1 at very strong strength; exon 4 skipping to p.(Ala72Thrfs*5) is consistent with this rule.
The variant is absent from gnomAD v2.1 and is observed at 0/1,448,024 alleles in gnomAD v4.1, with 0/33,128 alleles in the highest observed subpopulation, which is below the PTEN PM2 thresholds of 0.001% overall and 0.002% within any subpopulation and supports PM2_Supporting.
gnomad_v4 ↗With PVS1 and PM2_Supporting under the PTEN VCEP framework, NM_000314.8:c.253+5G>A is classified as likely pathogenic.
cspec ↗