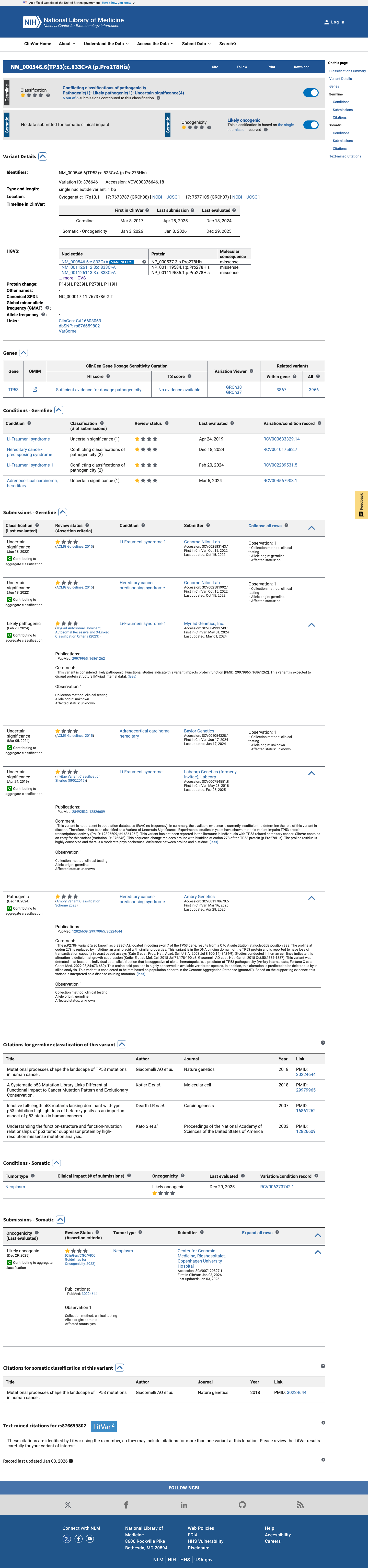
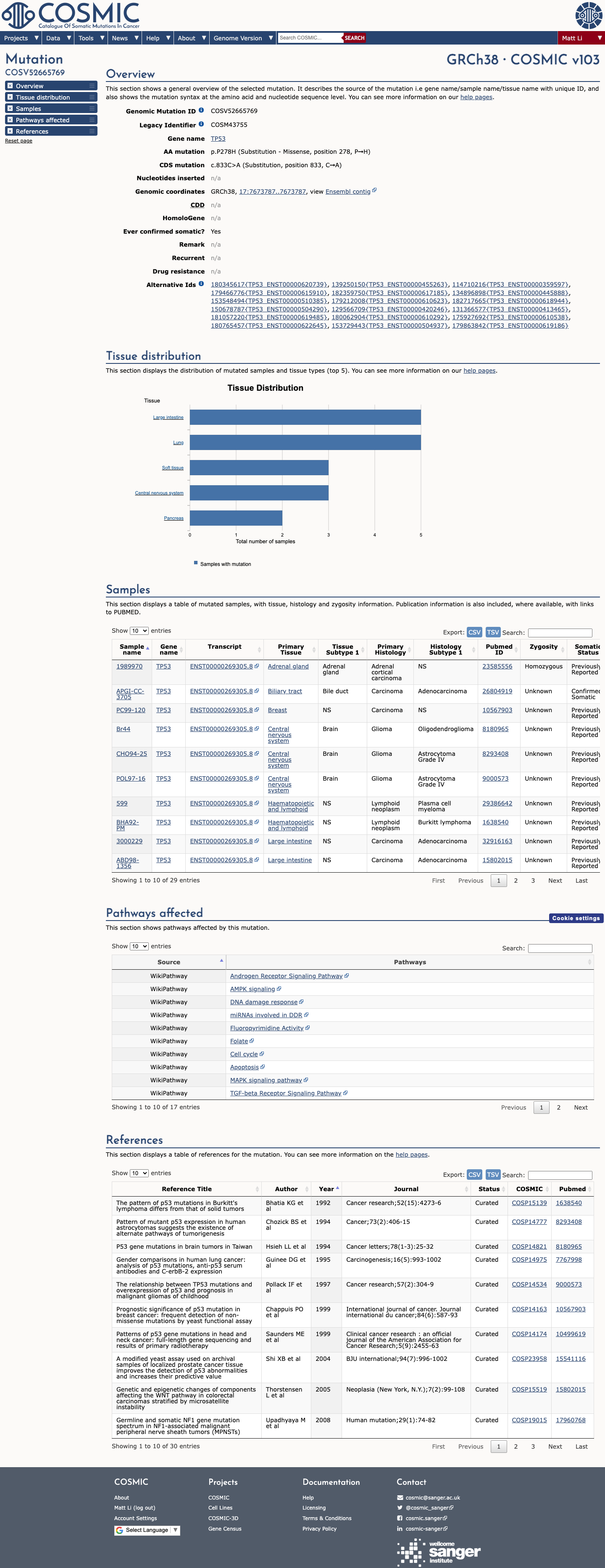
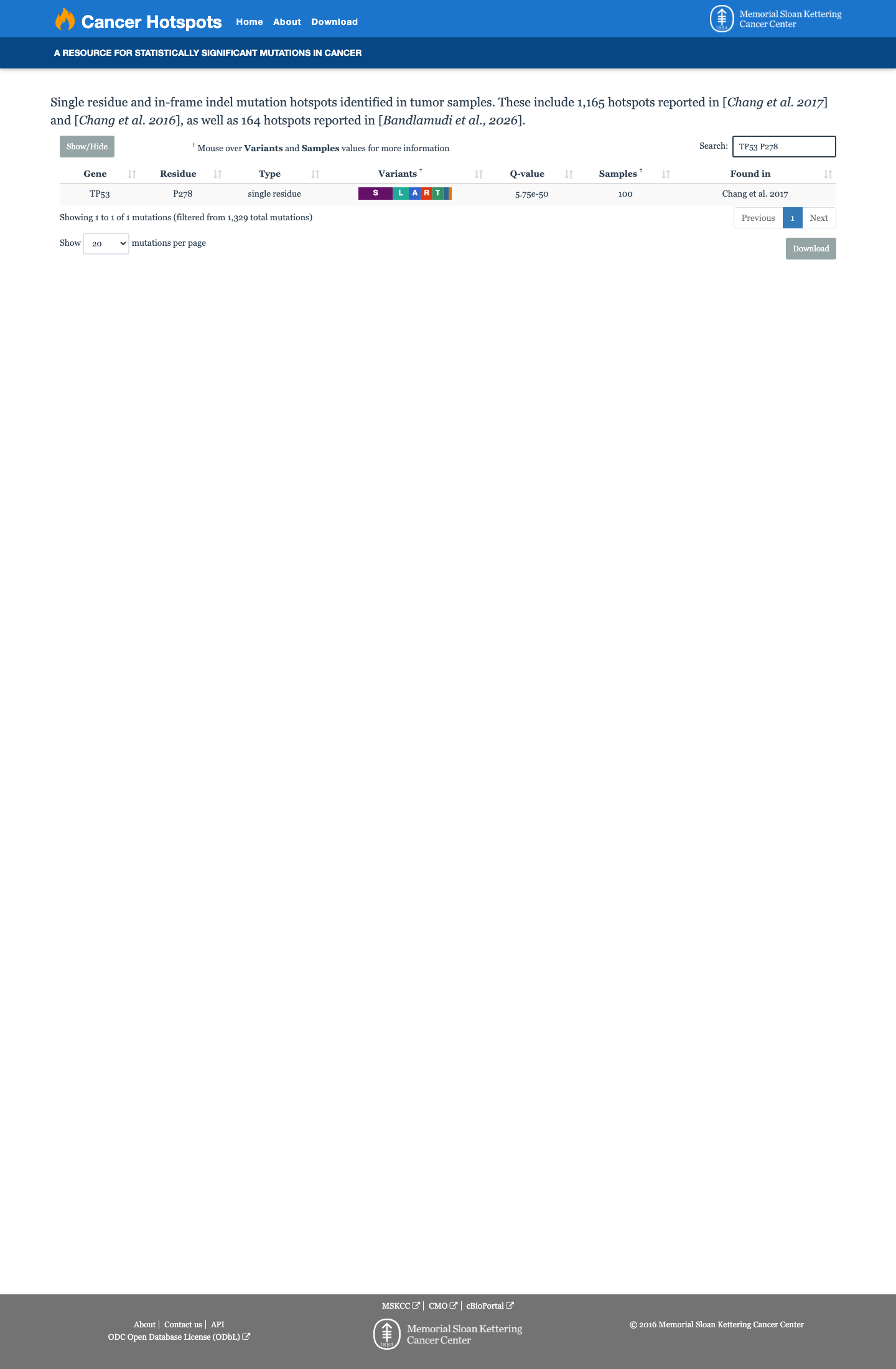
TP53 p.Pro278His has abnormal functional evidence, and the TP53 VCEP functional worksheet assigns PS3 for this amino acid substitution.
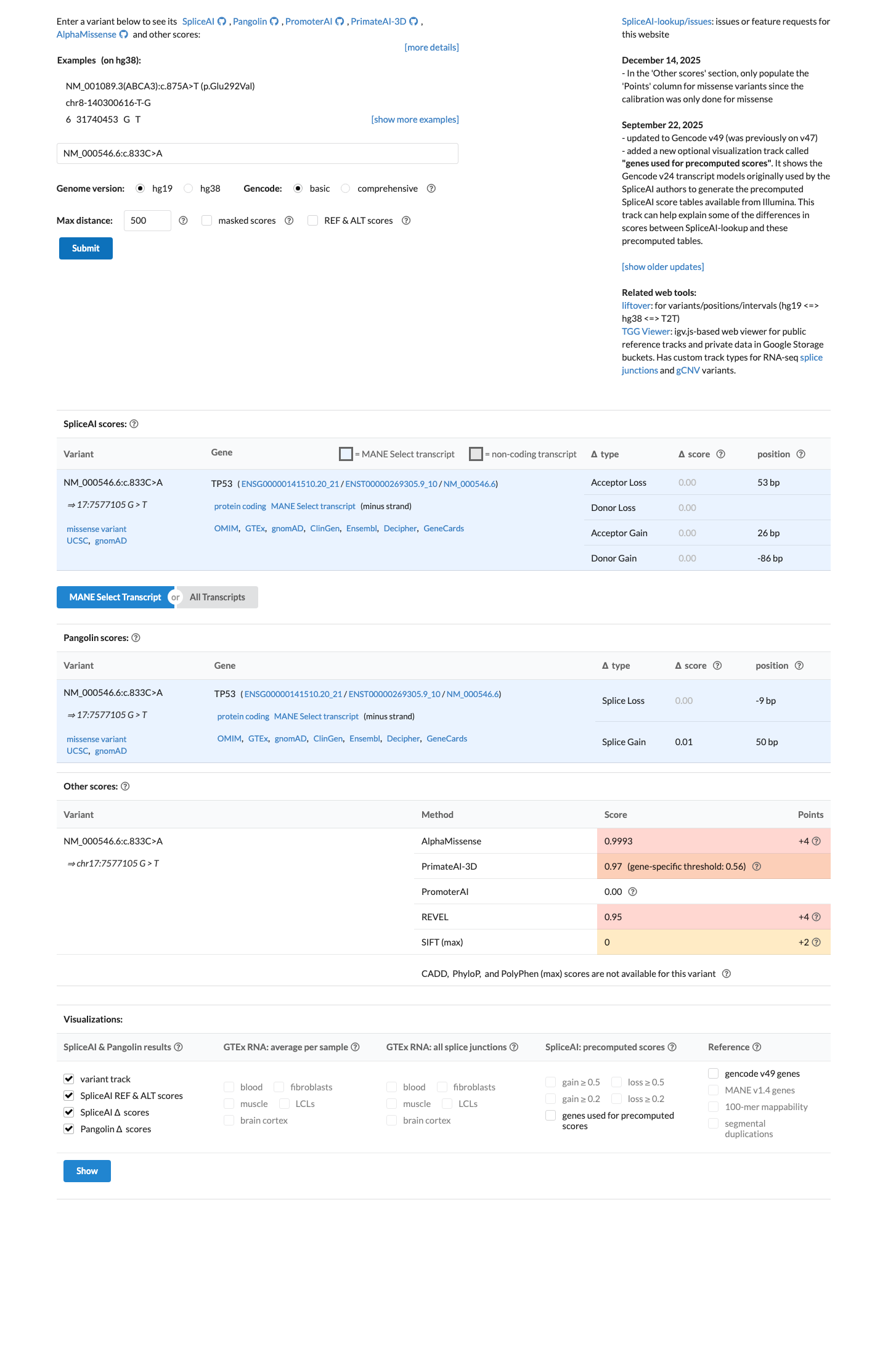
TP53 in silico assessment assigns c.833C>A as PP3_Moderate, with BayesDel 0.607821 and Class C65 supporting a deleterious missense effect.
SpliceAI predicts a max delta score of 0.00, which is below the TP53 splice-impact threshold of 0.2 and does not indicate a predicted splice effect.
spliceai ↗The variant is absent from gnomAD v4.1, and absence from population databases is consistent with the TP53 PM2_Supporting threshold of less than 0.00003.
gnomad_v4 ↗Using the generic ACMG/AMP combination rules as a fallback because explicit TP53 VCEP final Tavtigian score thresholds were not explicitly retrieved, PS3 with PP3_Moderate and PM2_Supporting supports a Likely Pathogenic classification.