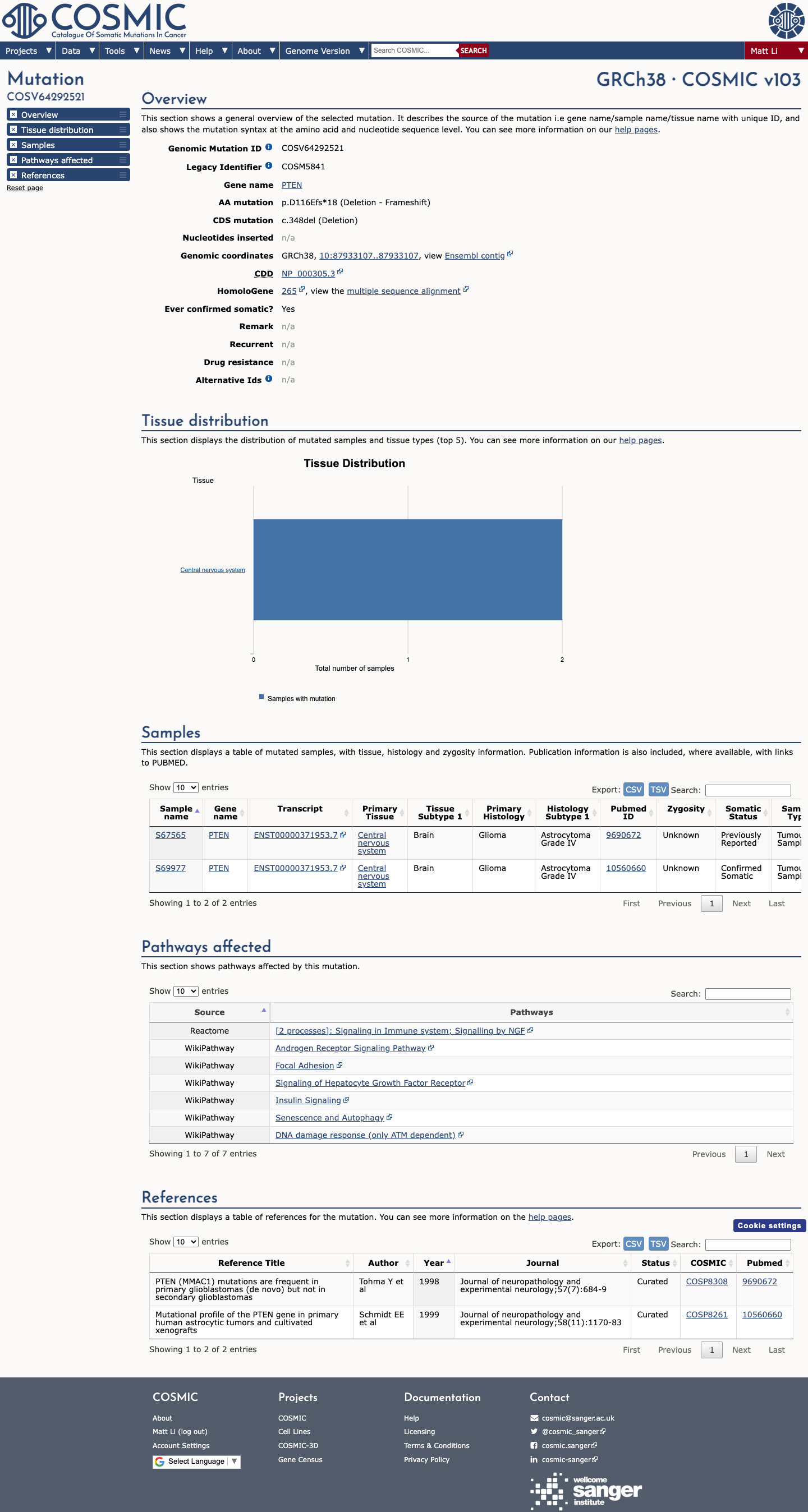
NM_000314.8:c.348del predicts PTEN p.(Asp116GlufsTer18), a frameshift expected to truncate the protein early in exon 5.
The PTEN PVS1 decision tree assigns PVS1 to frameshift variants in NM_000314.8 when the stop/disruption occurs at or 5' to p.D375 (c.1121); this variant truncates at p.Asp116fs, which is well upstream of that threshold and is consistent with loss of function.
The variant is absent from gnomAD v2.1 and gnomAD v4.1, and the PTEN specification applies PM2 at Supporting strength for variants with allele frequency below 0.00001, with a subpopulation threshold below 0.00002 when multiple alleles are present.
cspec ↗No PTEN-specific final classification framework was retrieved, so the generic ACMG/AMP combination rules were used as fallback; with PVS1 and PM2_Supporting alone, this evidence set does not reach a defined pathogenic or likely pathogenic combination under those rules, and the variant is therefore classified as a variant of uncertain significance pending human review.