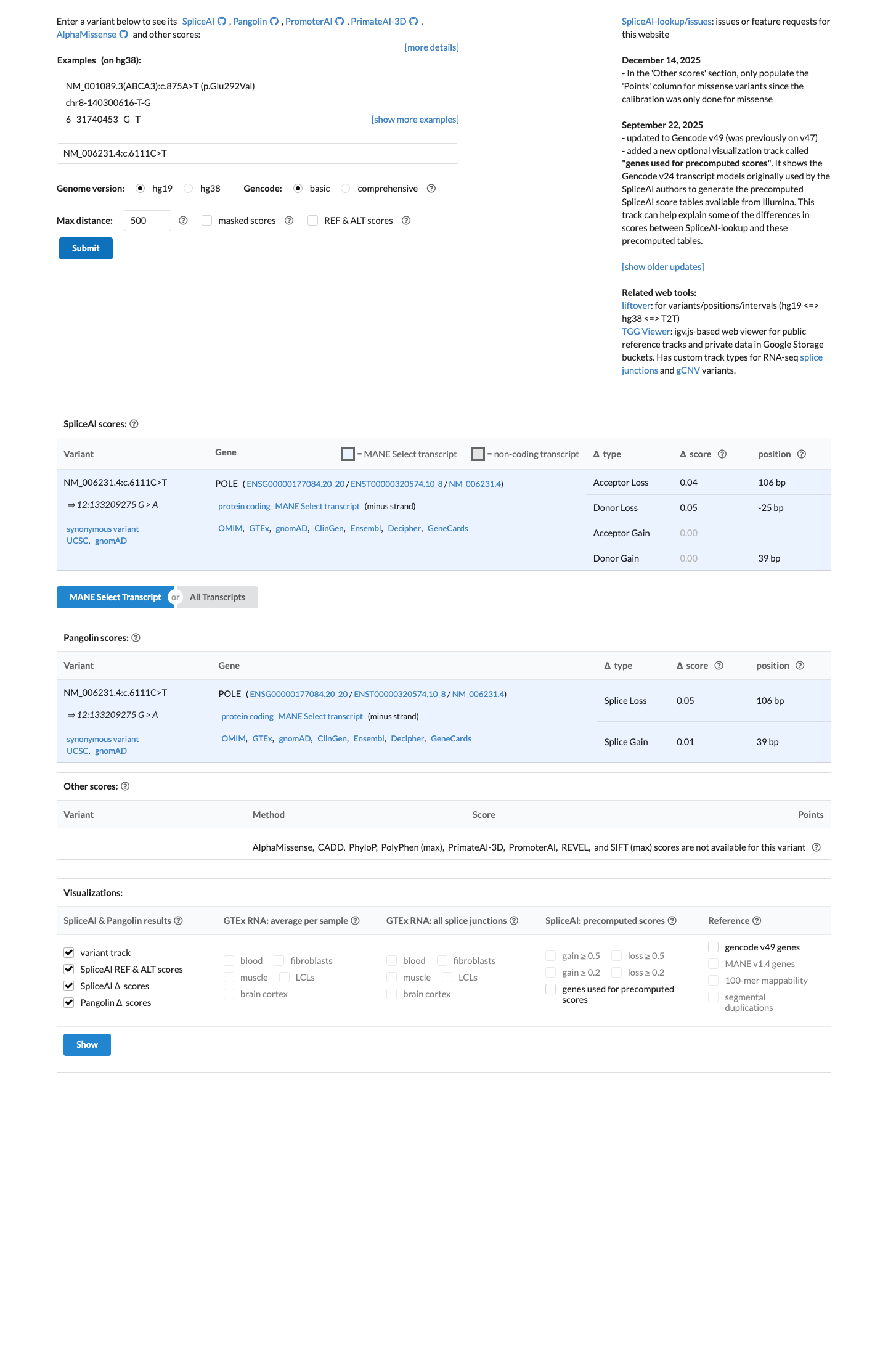
NM_006231.4:c.6111C>T is a synonymous POLE variant, NP_006222.2:p.(Ala2037=), and SpliceAI predicts no significant splice impact with a maximum delta score of 0.05, supporting BP7.
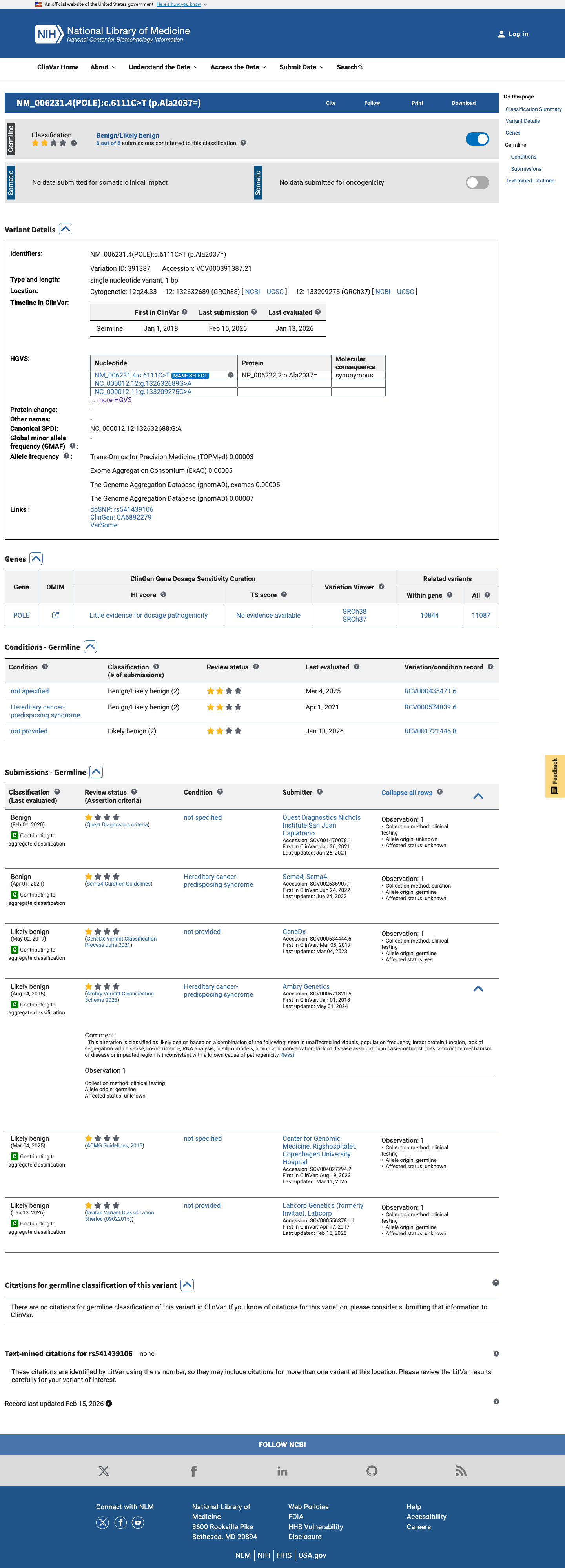
spliceai ↗The variant has been submitted to ClinVar as Likely benign by 4 clinical laboratories and as Benign by 1 clinical laboratory, supporting BP6 under the generic ACMG/AMP framework.
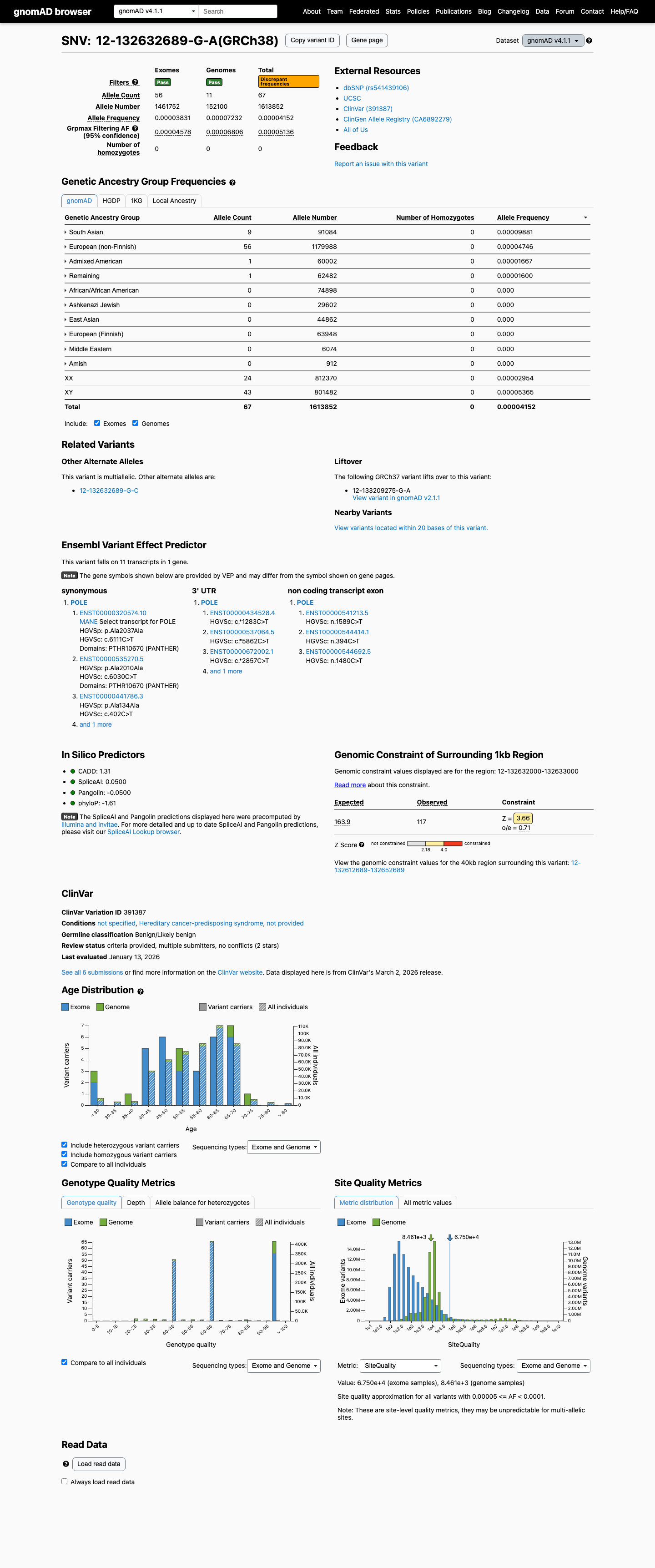
clinvar ↗Population frequency is low in gnomAD v4.1 at 4.15156e-05 (0.00415%), which is below the BS1 threshold of 0.3% and the BA1 threshold of 1.0%, so benign frequency criteria stronger than supporting are not met.
gnomad_v4 ↗Using the retrieved generic ACMG/AMP final classification combination rules, BP6 plus BP7 constitutes two supporting benign criteria and is consistent with a Likely benign classification.