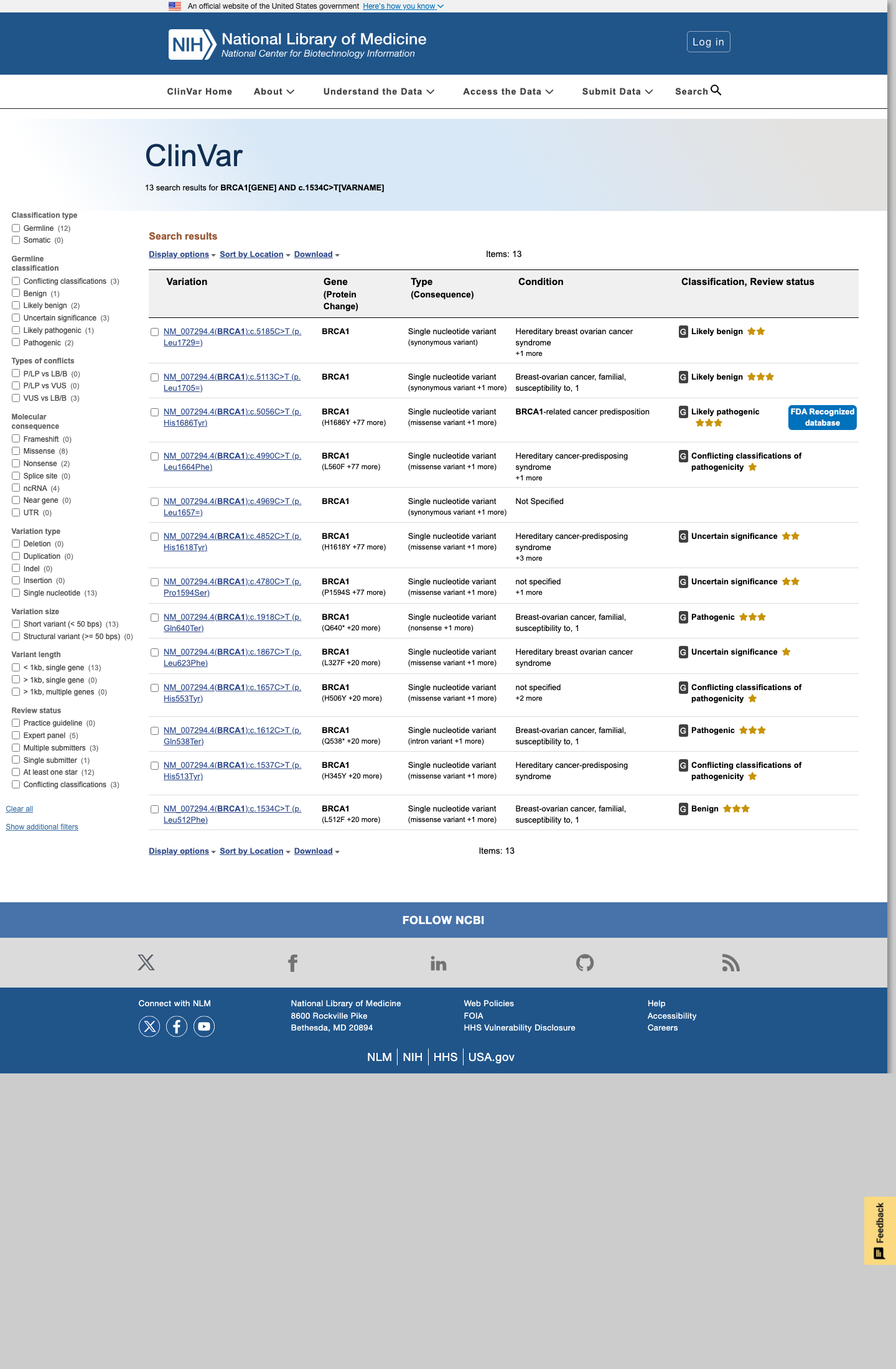
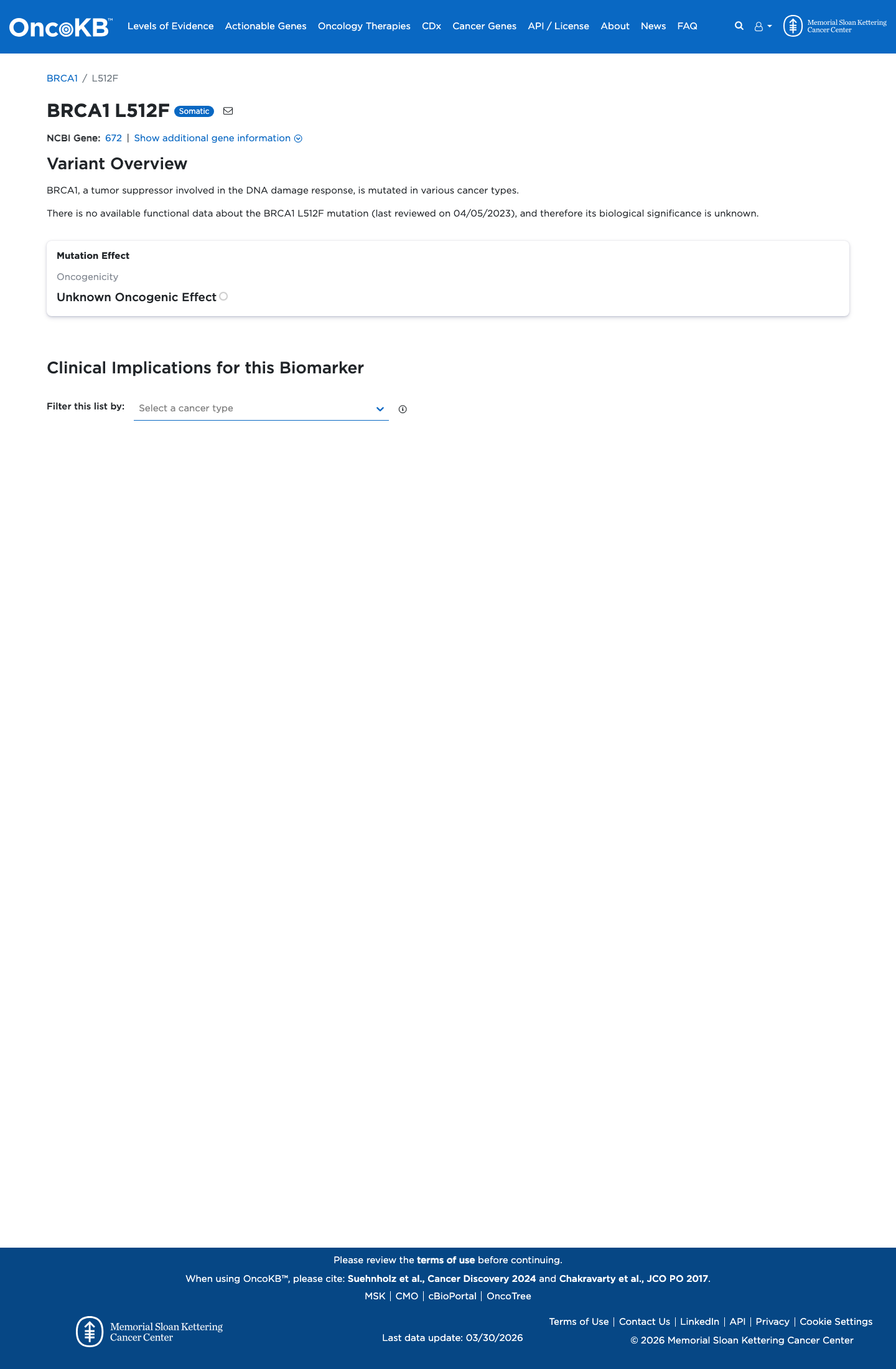
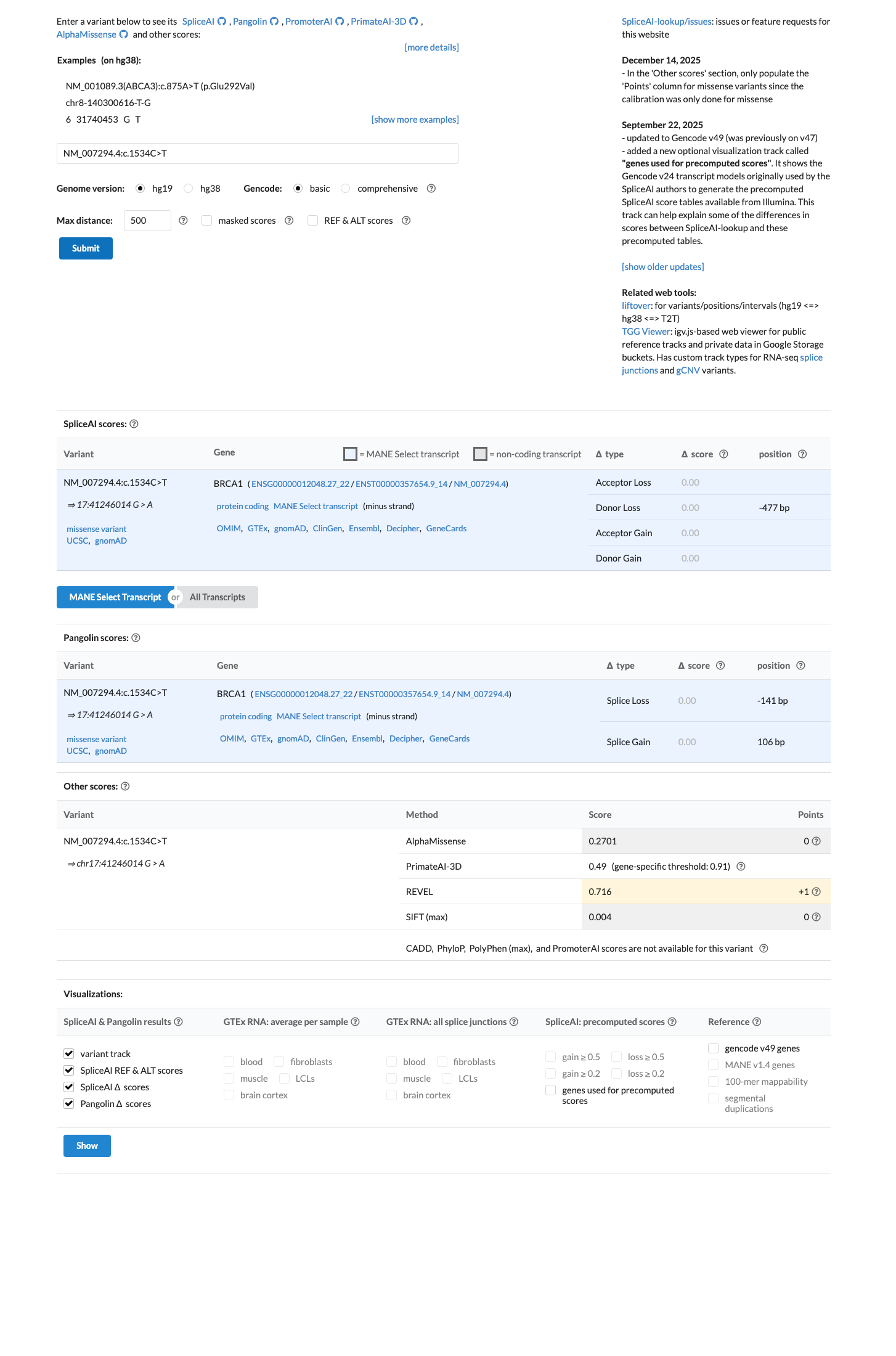
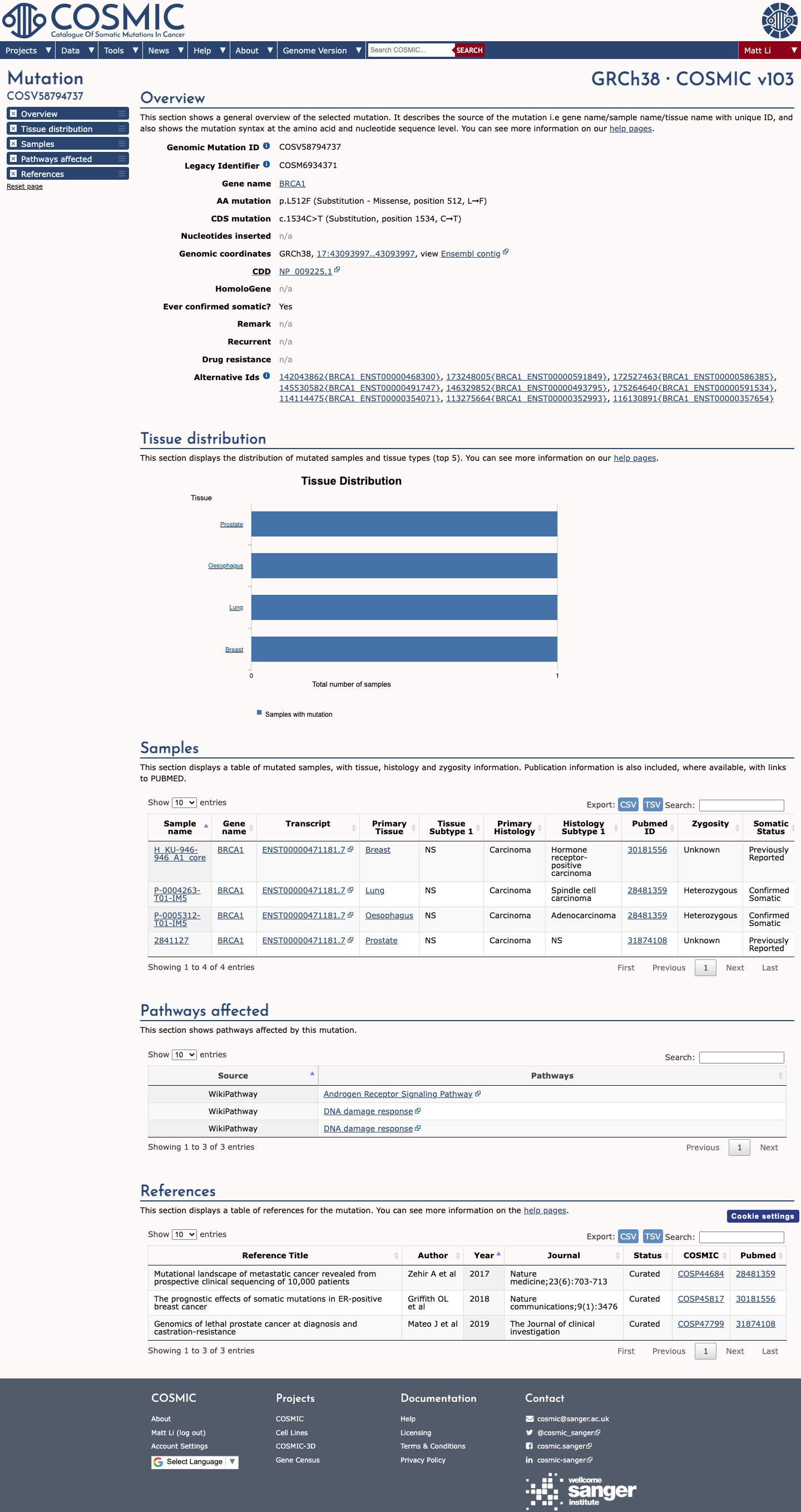
BRCA1 c.1534C>T (p.Leu512Phe, p.L512F) has been shown in a calibrated functional assay to have protein function similar to benign control variants, supporting BS3_Strong.
Multifactorial likelihood analysis yielded a combined LR for causality of 0.007006151007218386, which is below the ENIGMA BP5_Strong threshold of 0.05 and supports strong evidence against pathogenicity.
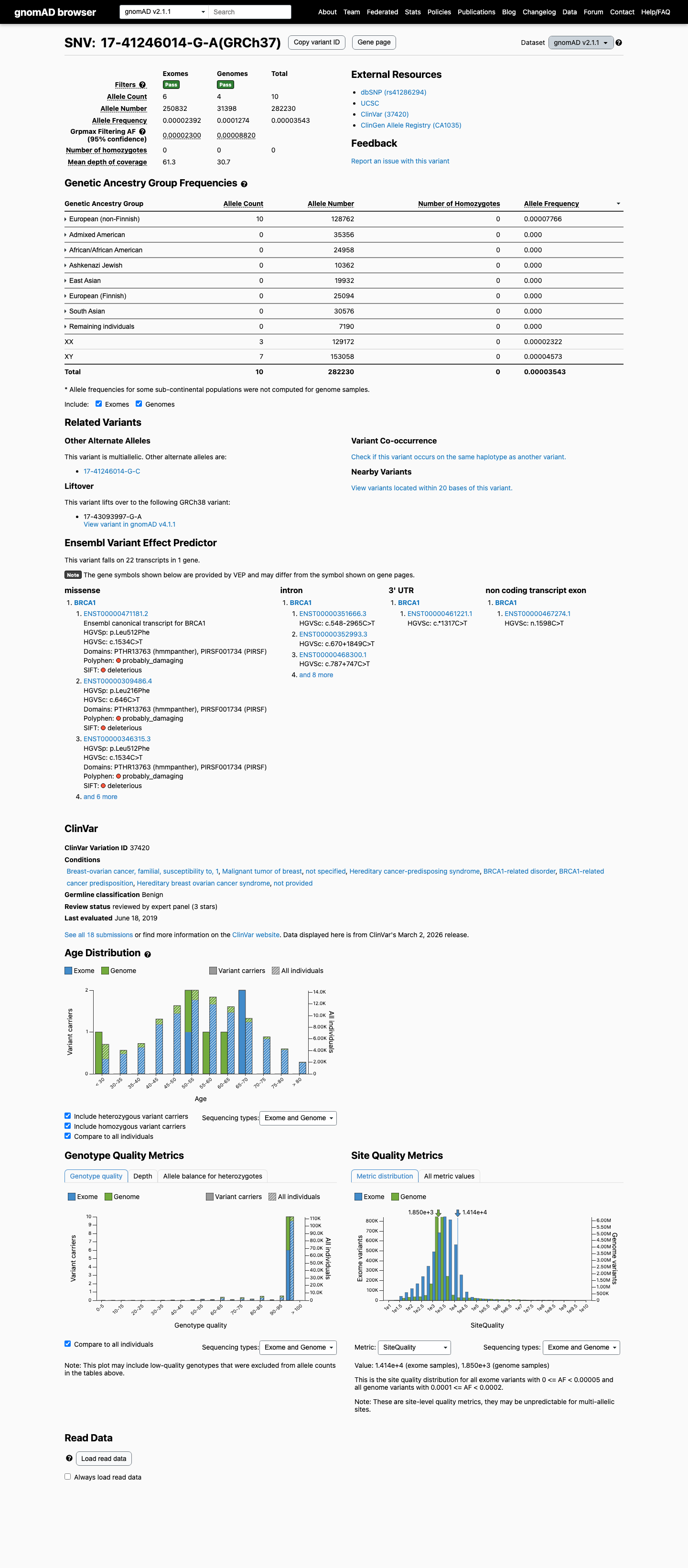
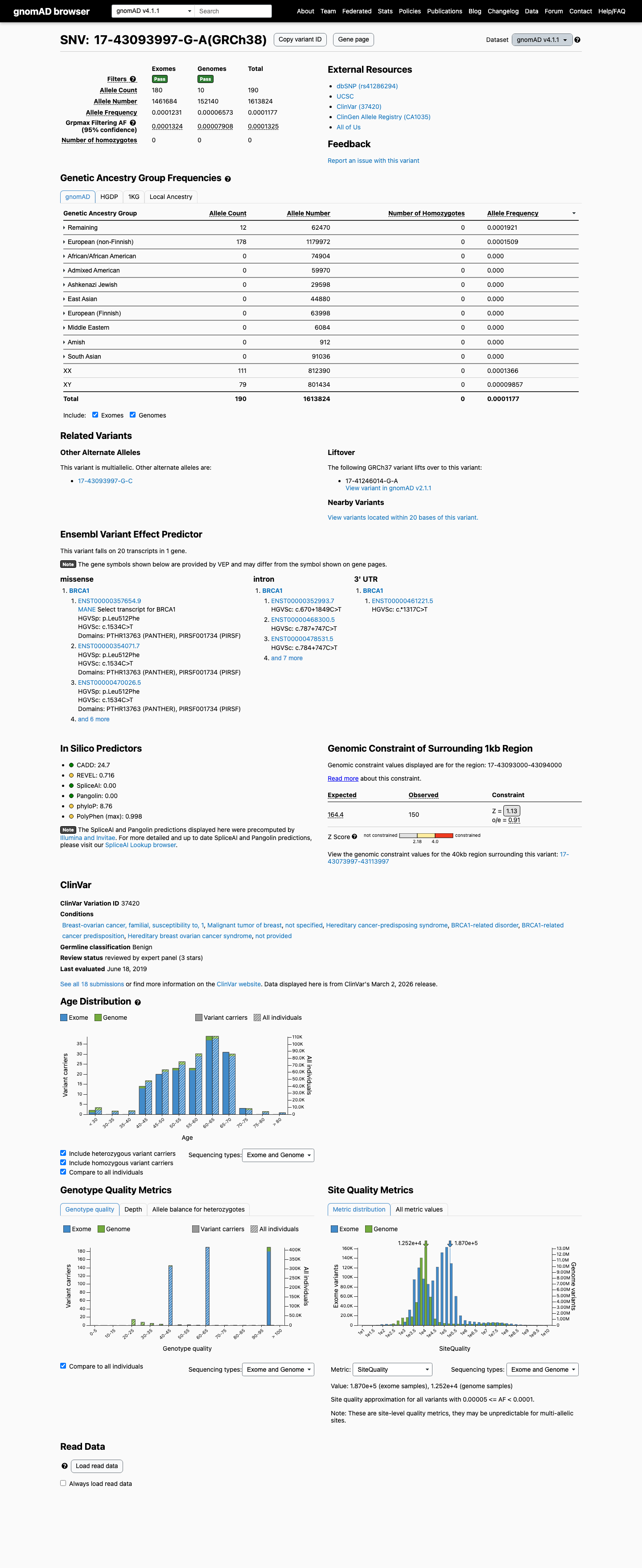
The gnomAD v2.1 non-founder grpmax FAF is 8.82e-05, which is within the BS1_Supporting range of >2e-05 to <=1e-04, and the missense change lies outside the BRCA1 clinically important domains with SpliceAI 0.00, supporting BS1_Supporting and BP1_Strong benign evidence.
gnomad_v2 ↗Using the retrieved generic ACMG/AMP combination rules as fallback final-classification framework, the presence of at least two Strong benign criteria supports classification of this variant as Benign.