Classification rationale
1
PTEN final classification was assigned using the explicit CSPEC/VCEP criteria-combination framework rather than generic ACMG/AMP combination rules.
2
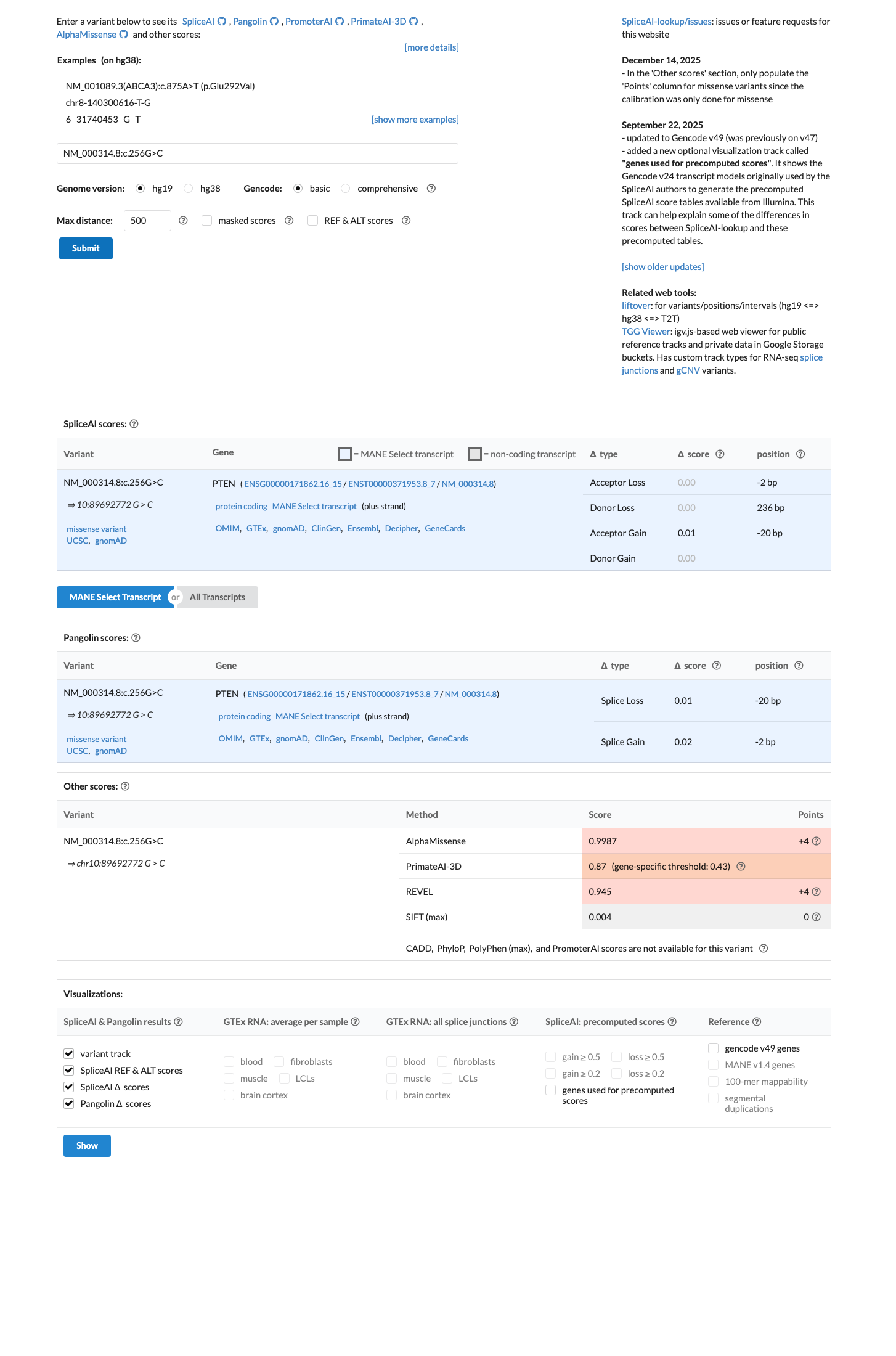
The applicable pathogenic evidence consists of PS3_Moderate, PM2_Supporting, PP2, and PP3.
3
This combination provides 1 Moderate and 3 Supporting pathogenic criteria, which does not match a defined PTEN rule for Likely Pathogenic or Pathogenic.
4
No benign standalone, strong, or supporting rule combination is satisfied, so the variant remains of uncertain significance.