Classification rationale
1
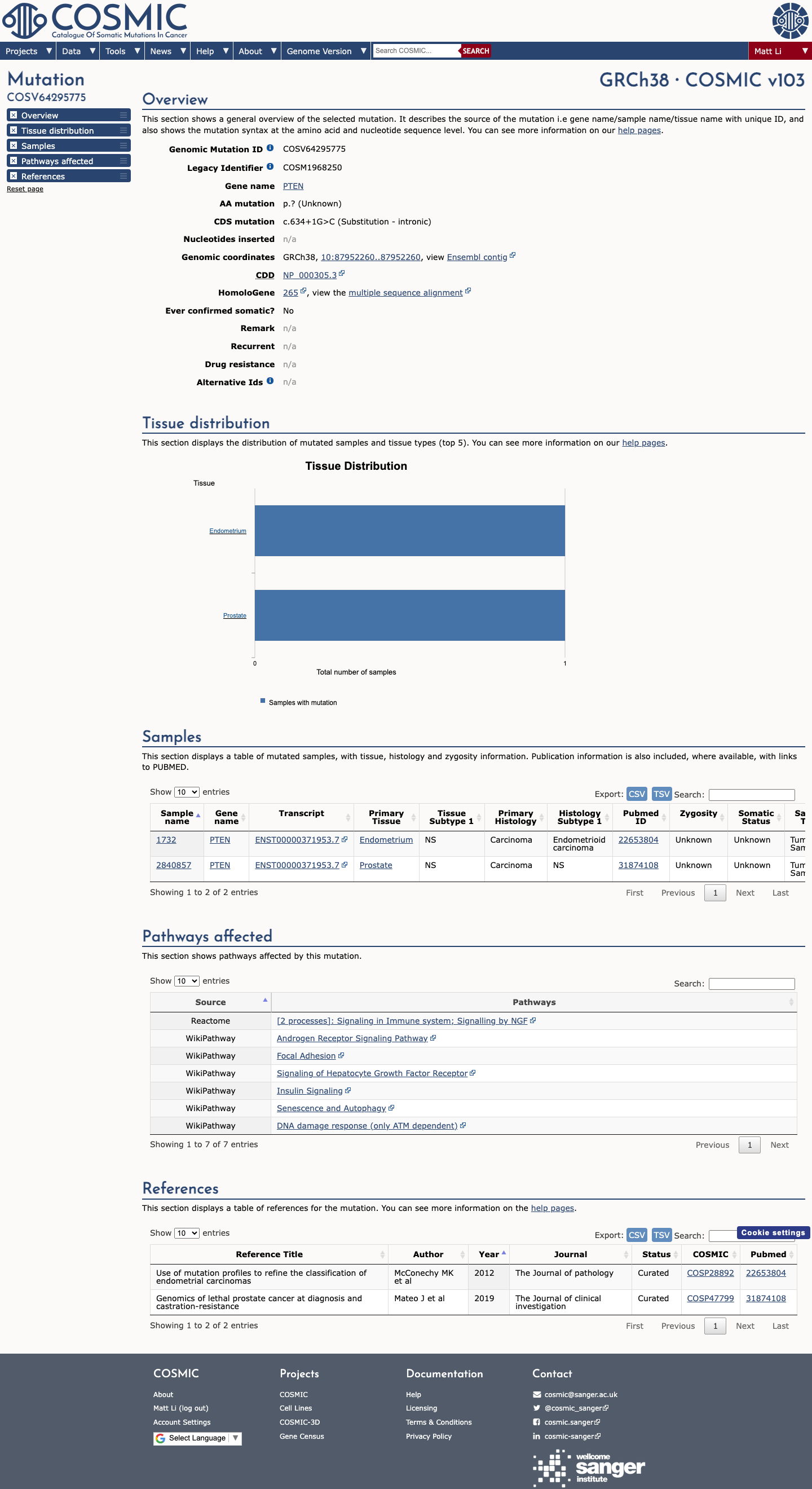
The PTEN c.634+1G>C (p.?) variant has been observed in somatic cancers in COSMIC (COSV64295775; 2 occurrences) and has been reported in ClinVar as pathogenic by three clinical laboratories.
clinvar ↗2
3
In silico splicing analysis predicts a marked effect on RNA processing, with a SpliceAI maximum delta score of 0.99, consistent with disruption of the canonical donor site.
spliceai ↗