Classification rationale
1
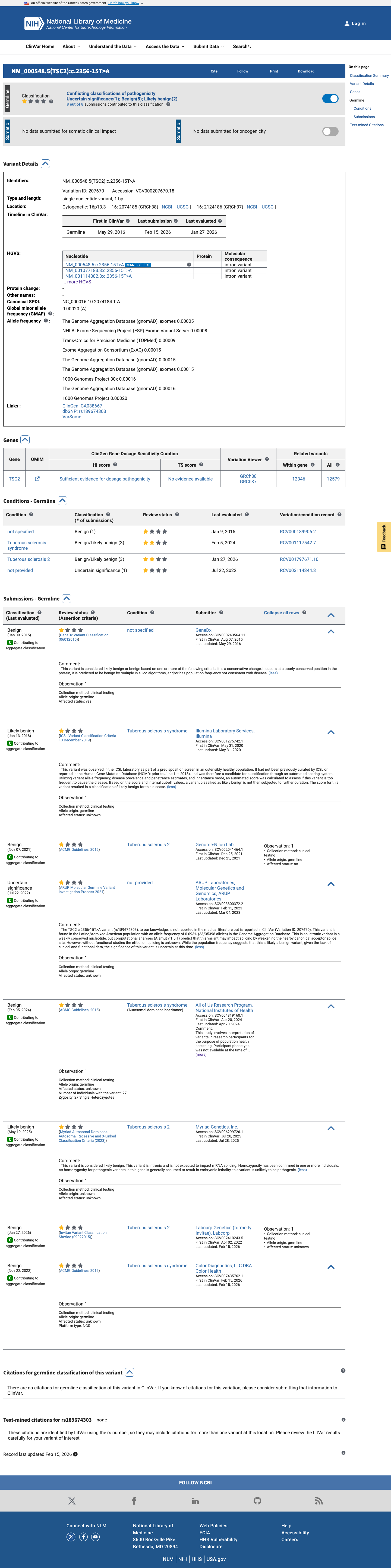
The TSC2 c.2356-15T>A (p.?) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar, where aggregate submissions include Benign, Likely benign, and Uncertain significance classifications.
clinvar ↗2
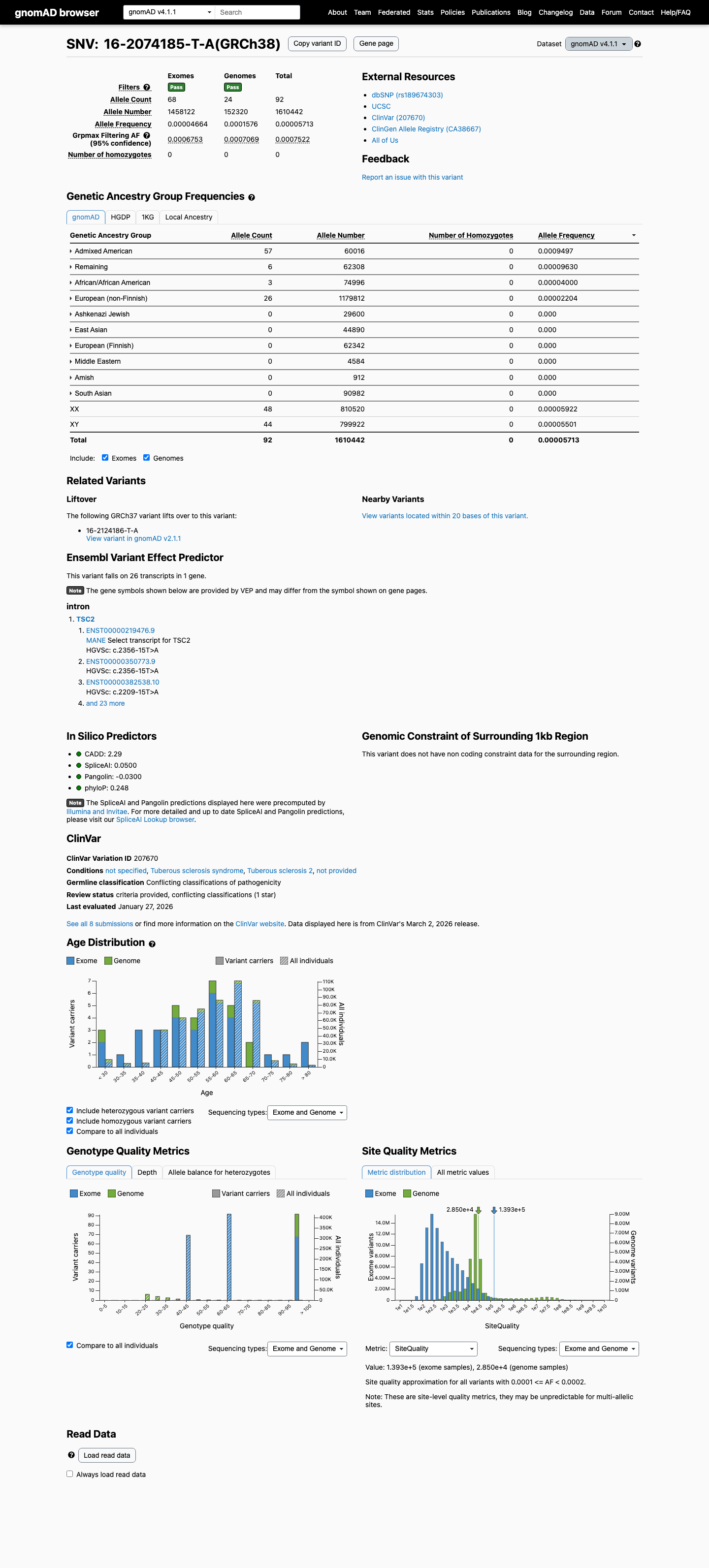
The variant is present at low frequency in population databases, with gnomAD v2.1 total AF 0.01435% (40/278790 alleles) and gnomAD v4.1 total AF 0.00571% (92/1610442 alleles), with no homozygotes reported.
gnomad_v2 ↗ gnomad_v4 ↗3
In silico splicing prediction is consistent with no significant splice effect, with SpliceAI maximal delta score 0.14.
spliceai ↗