Classification rationale
1
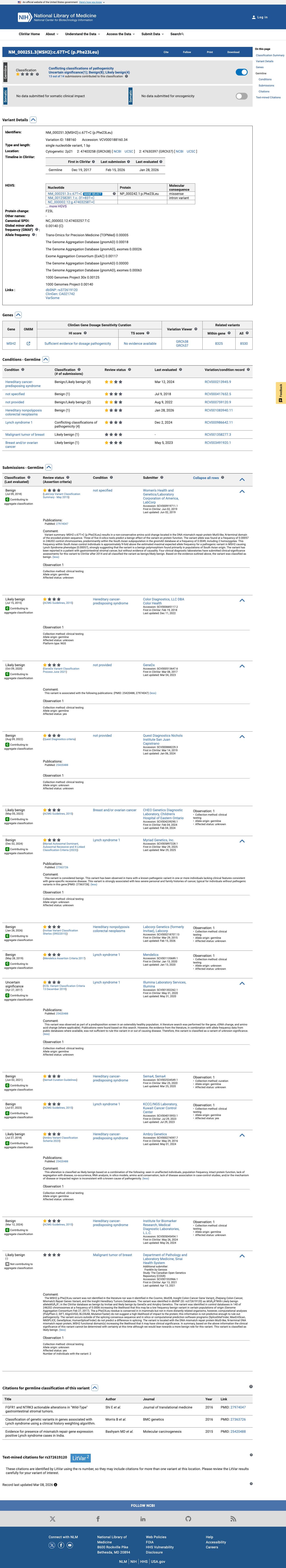
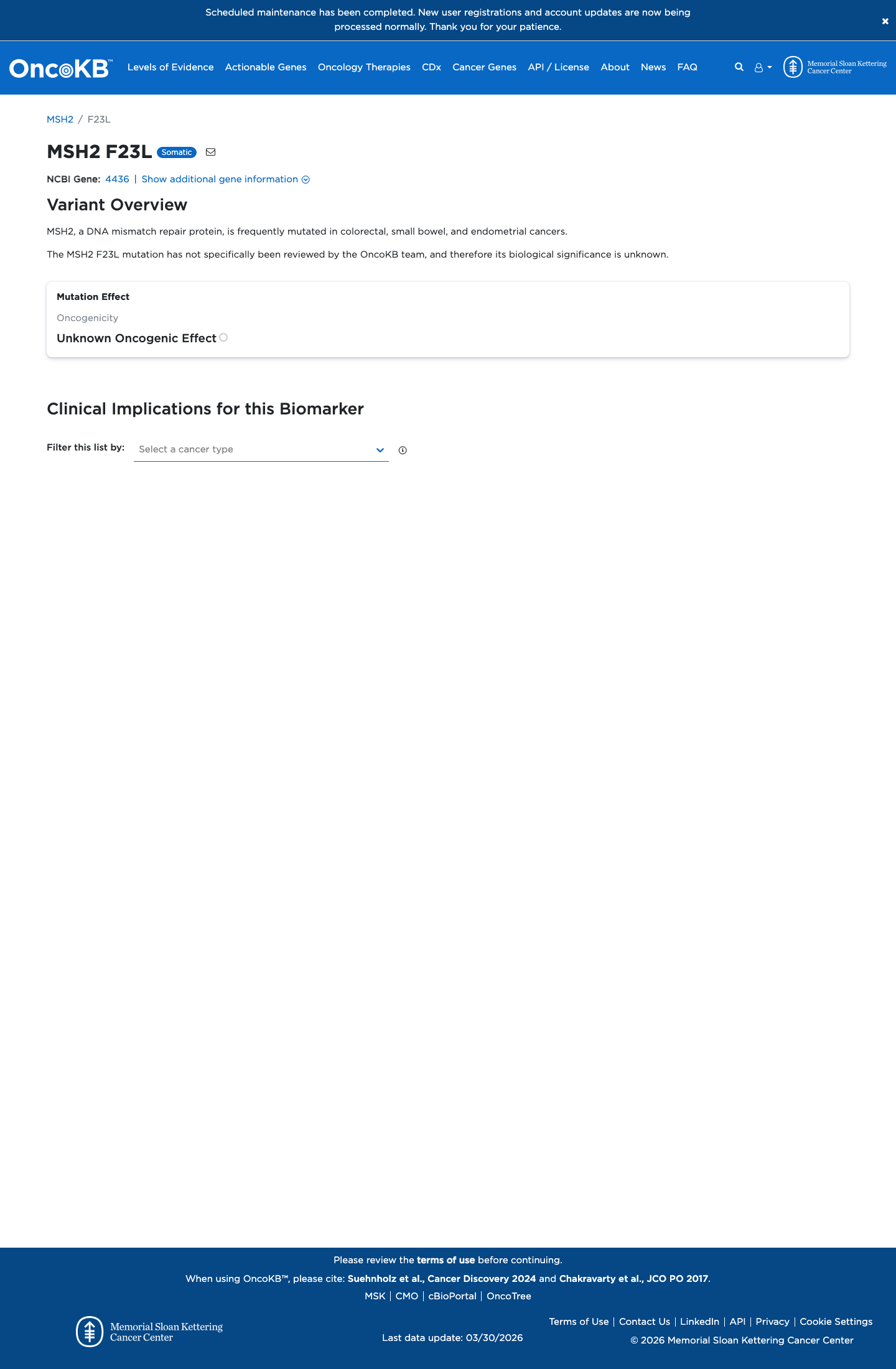
The MSH2 c.67T>C (p.Phe23Leu) variant has not been observed in COSMIC somatic cancer records and has been reported in ClinVar predominantly as benign or likely benign, with an aggregate ClinVar classification of Benign.
clinvar ↗2
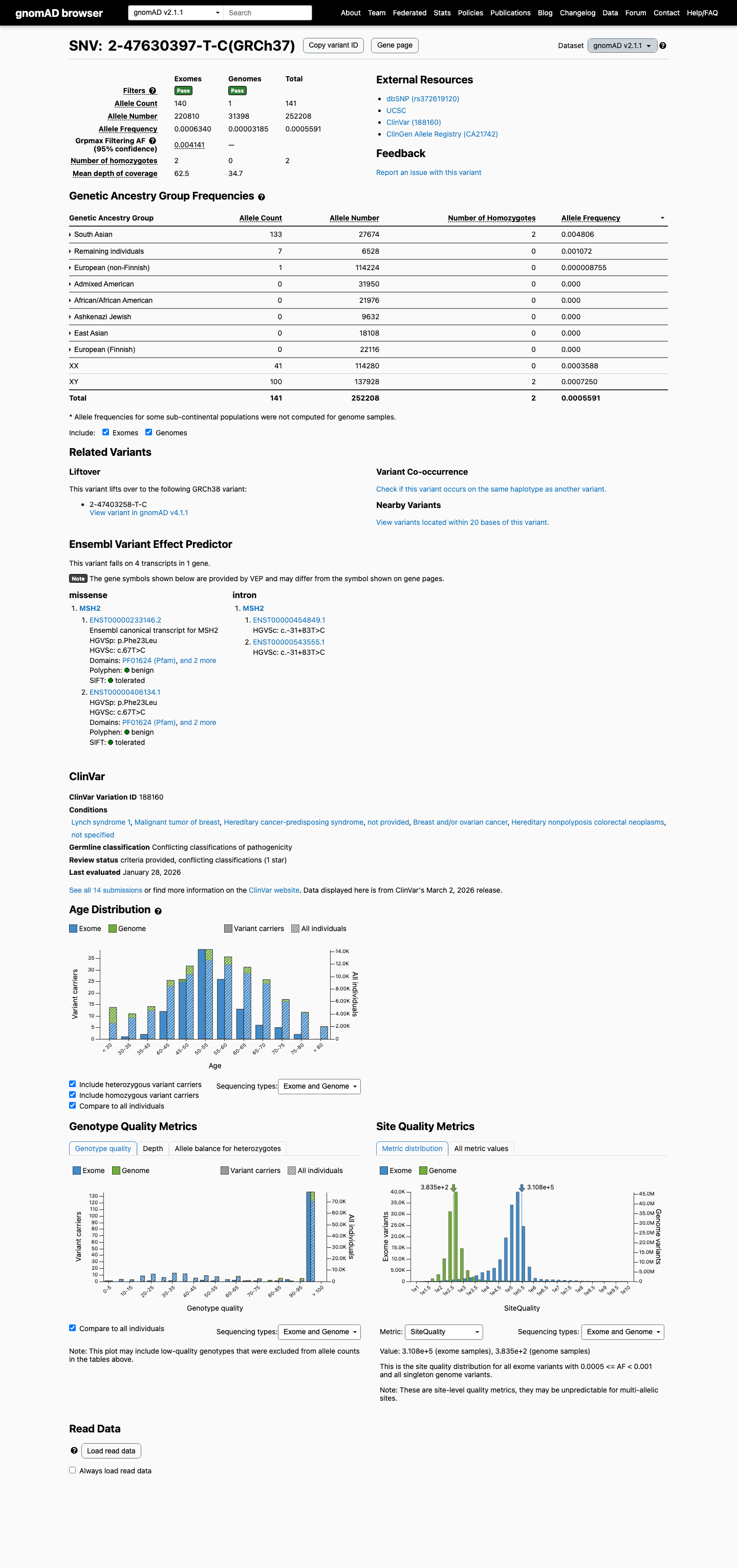
This variant is present at high frequency in population databases, including gnomAD v4.1 at 410/1600398 alleles with 8 homozygotes and grpmax filtering allele frequency 0.00394201, which exceeds the MSH2 VCEP BA1 threshold.
gnomad_v4 ↗ gnomad_v2 ↗3
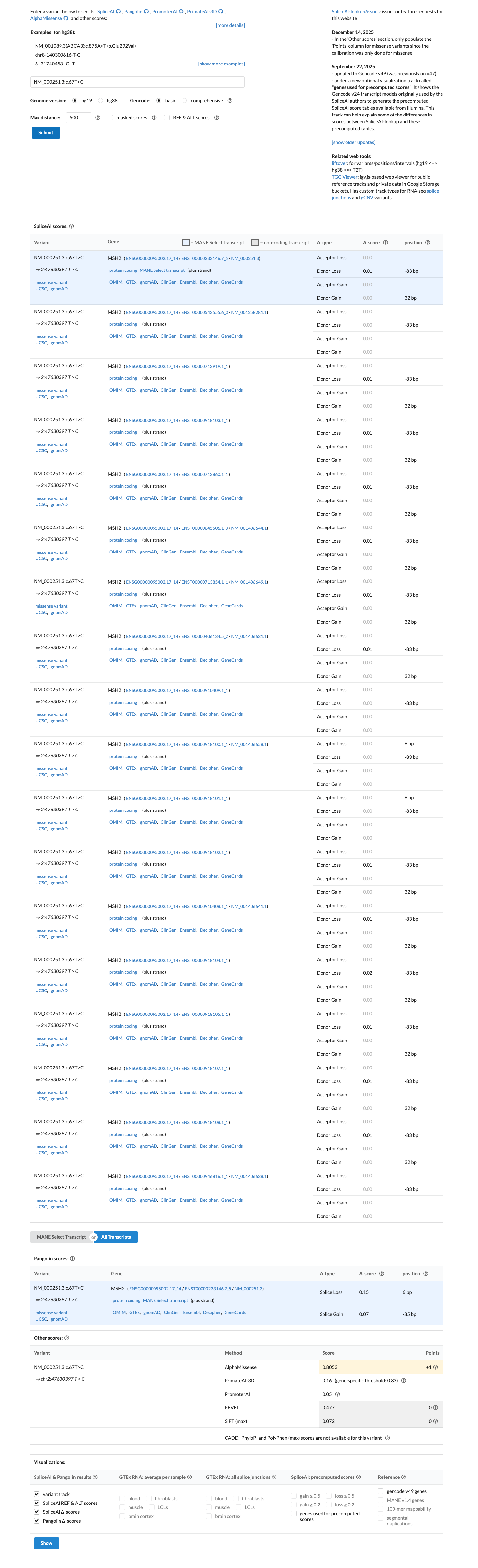
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01; no missense-specific HCI prior value was available to support PP3 or BP4 assessment.
spliceai ↗