Classification rationale
1
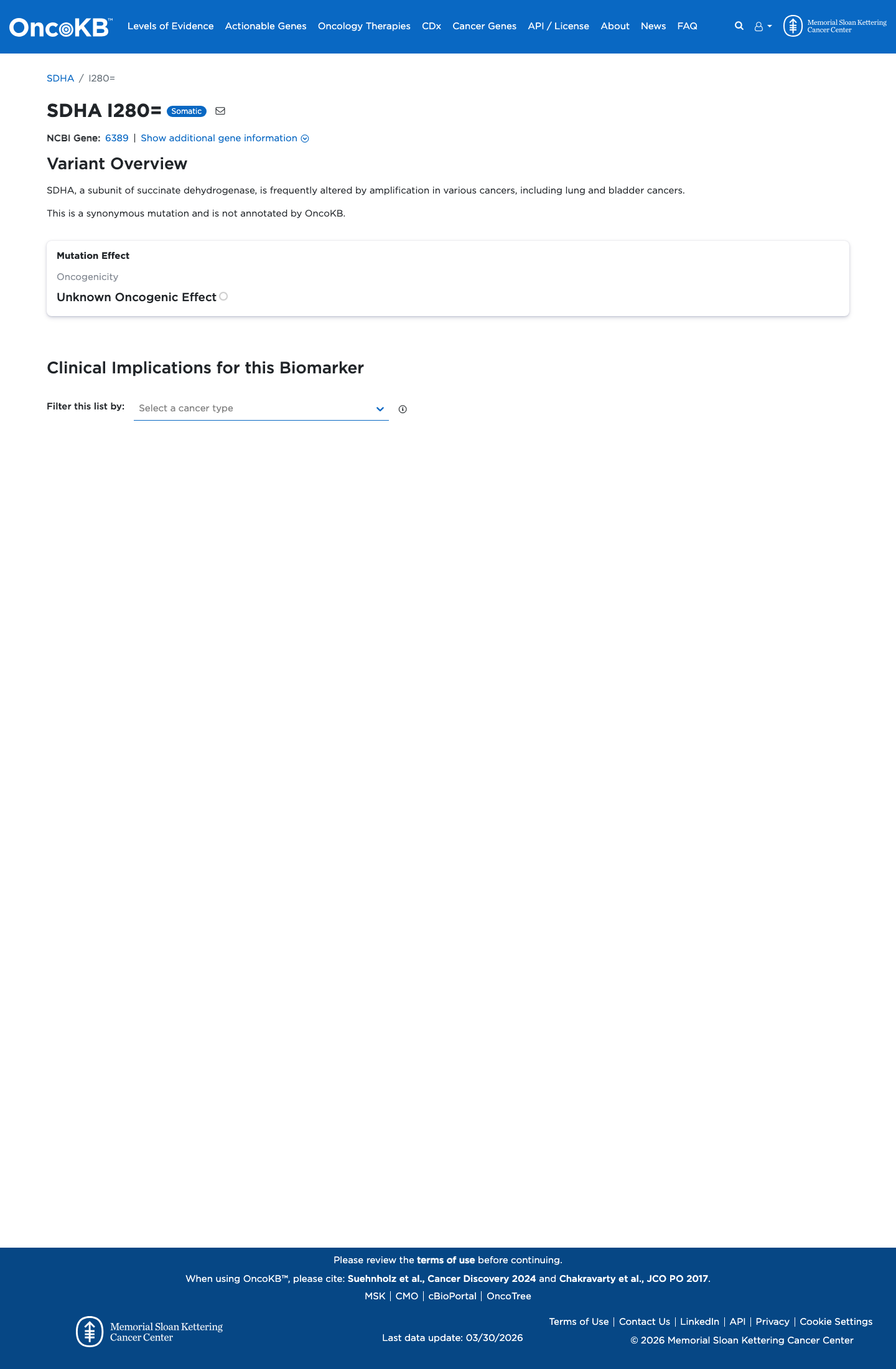
The SDHA c.840C>T (p.Ile280=) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as Likely benign with criteria provided from a single submitter.
clinvar ↗2
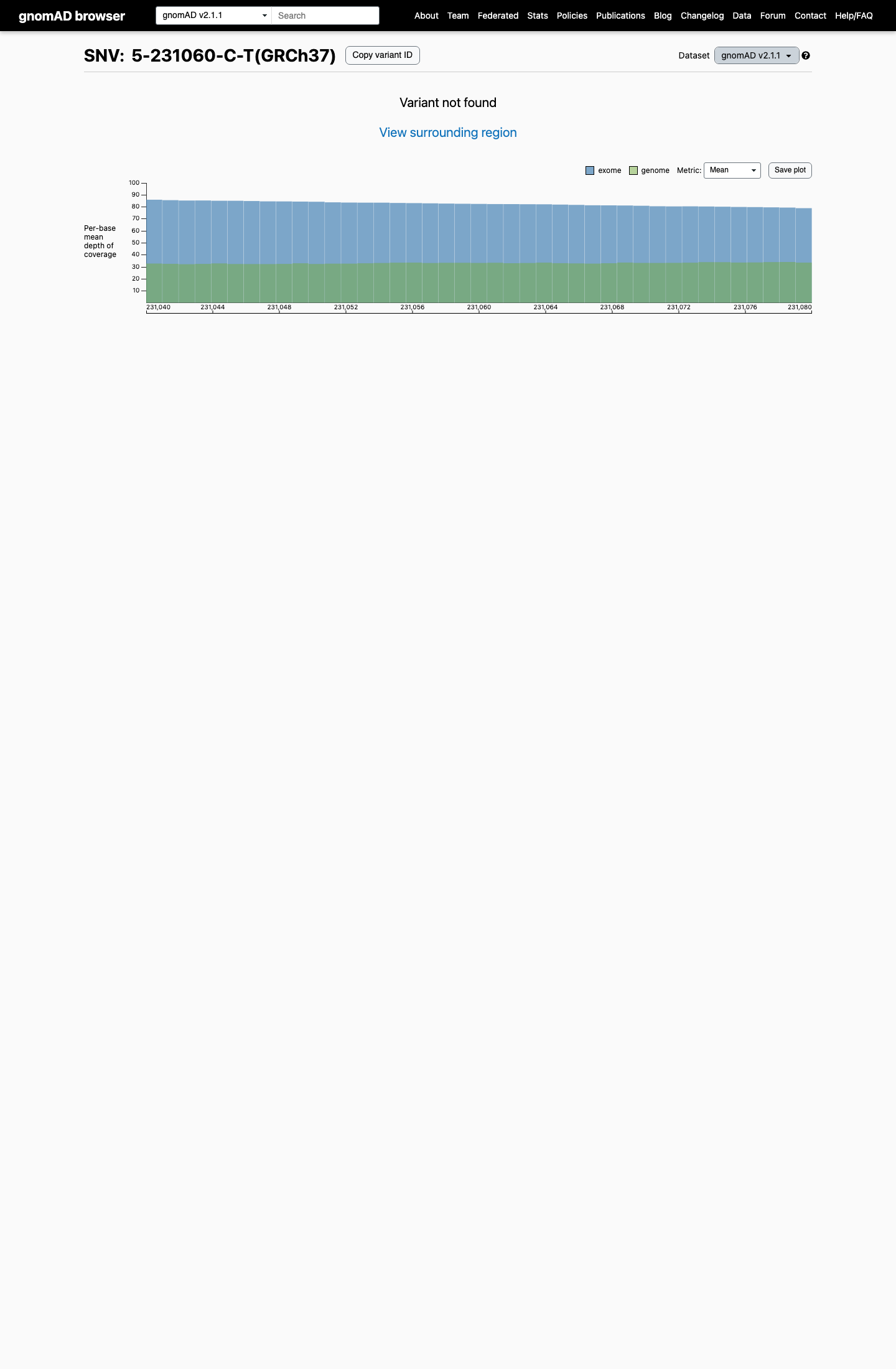
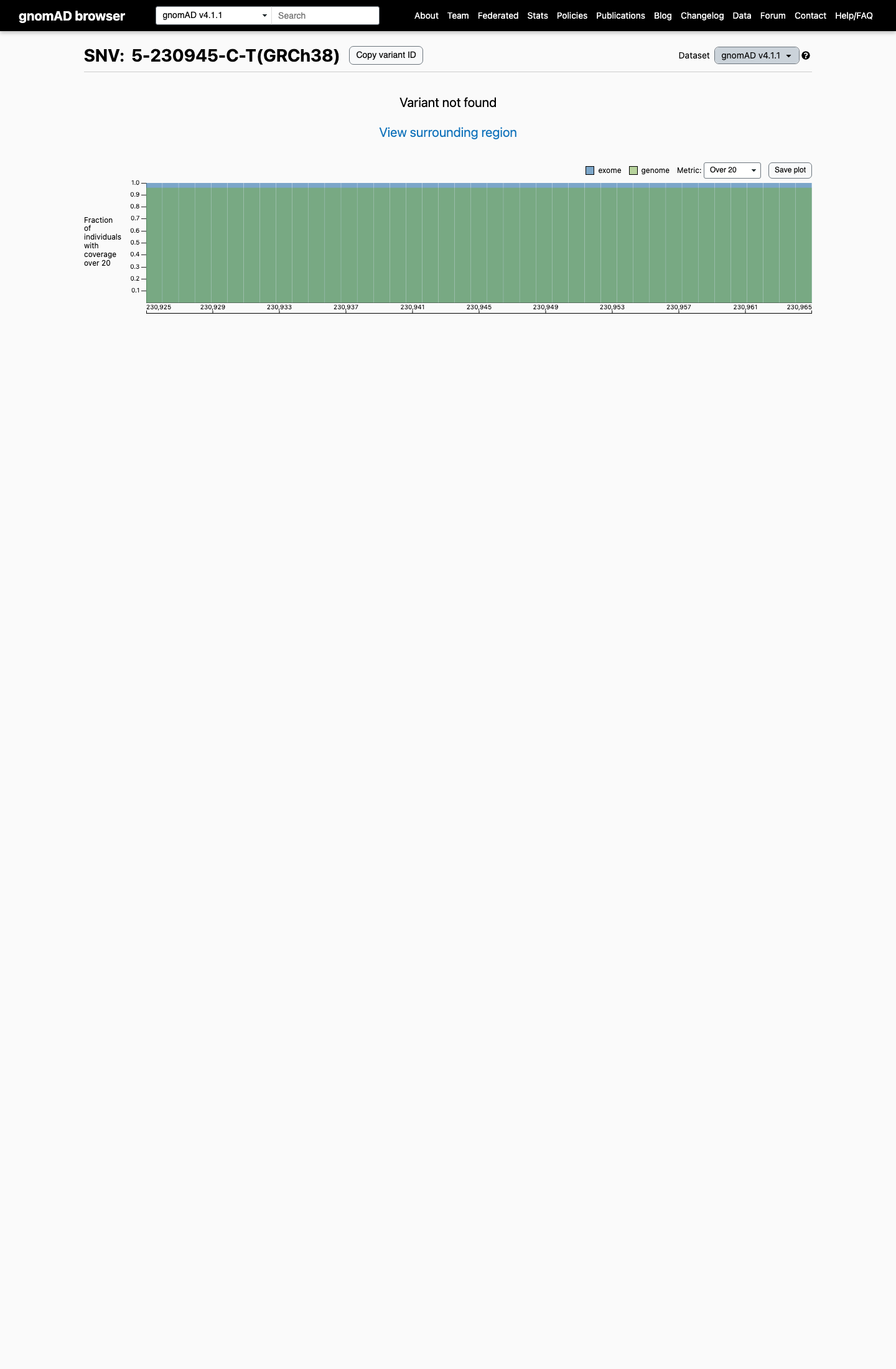
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is below the default PM2 rarity threshold of 0.1% and does not reach the BS1 (>0.3%) or BA1 (>1%) benign frequency thresholds.
gnomad_v4 ↗3
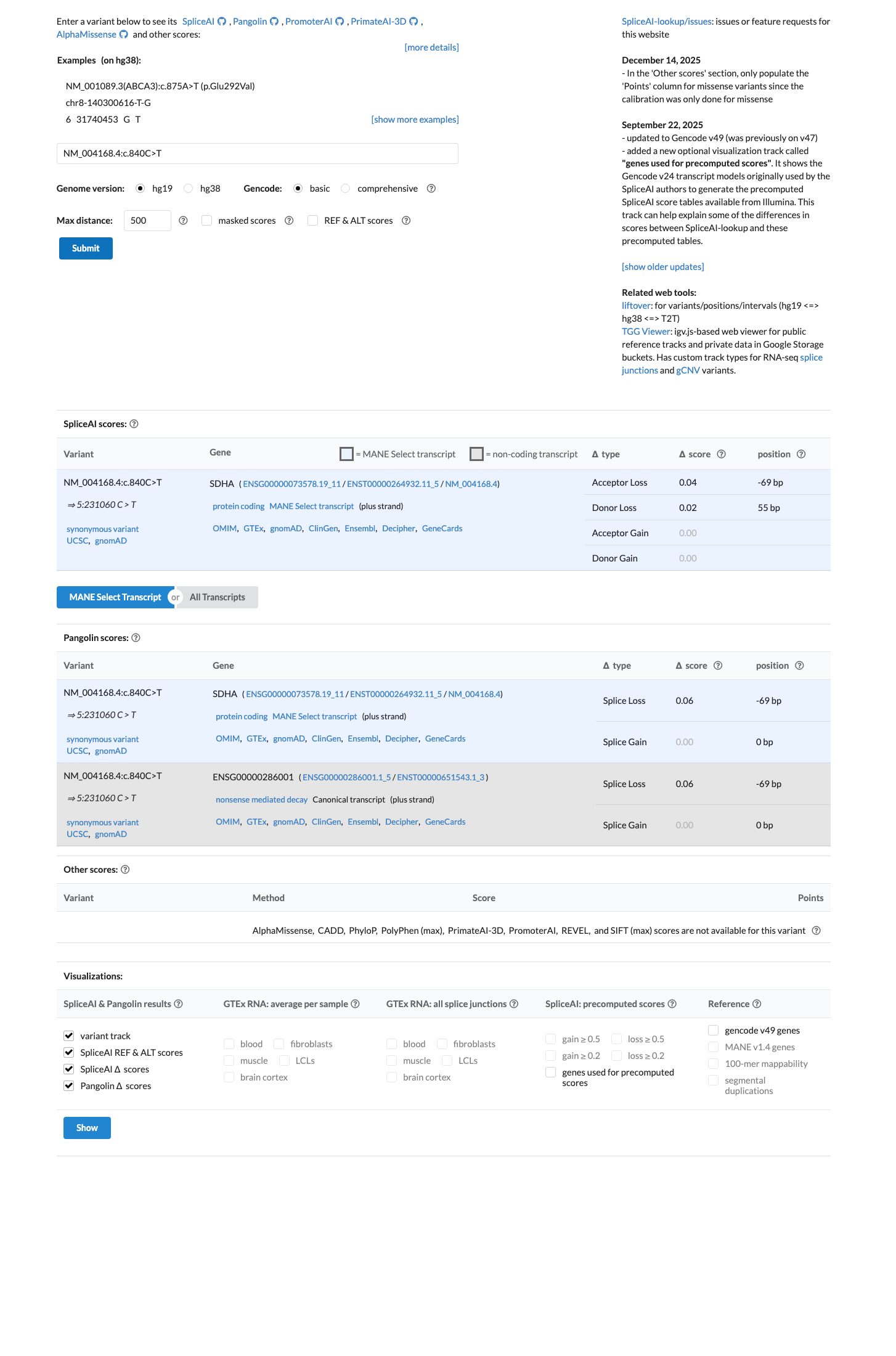
In silico assessment is consistent with a benign synonymous effect, as the variant does not change the amino acid sequence and SpliceAI predicts no significant splice impact with a maximum delta score of 0.04.
spliceai ↗