The MLH1 c.1990-23G>T (p.?) variant has not been observed in somatic cancers in COSMIC and has not been reported in ClinVar.
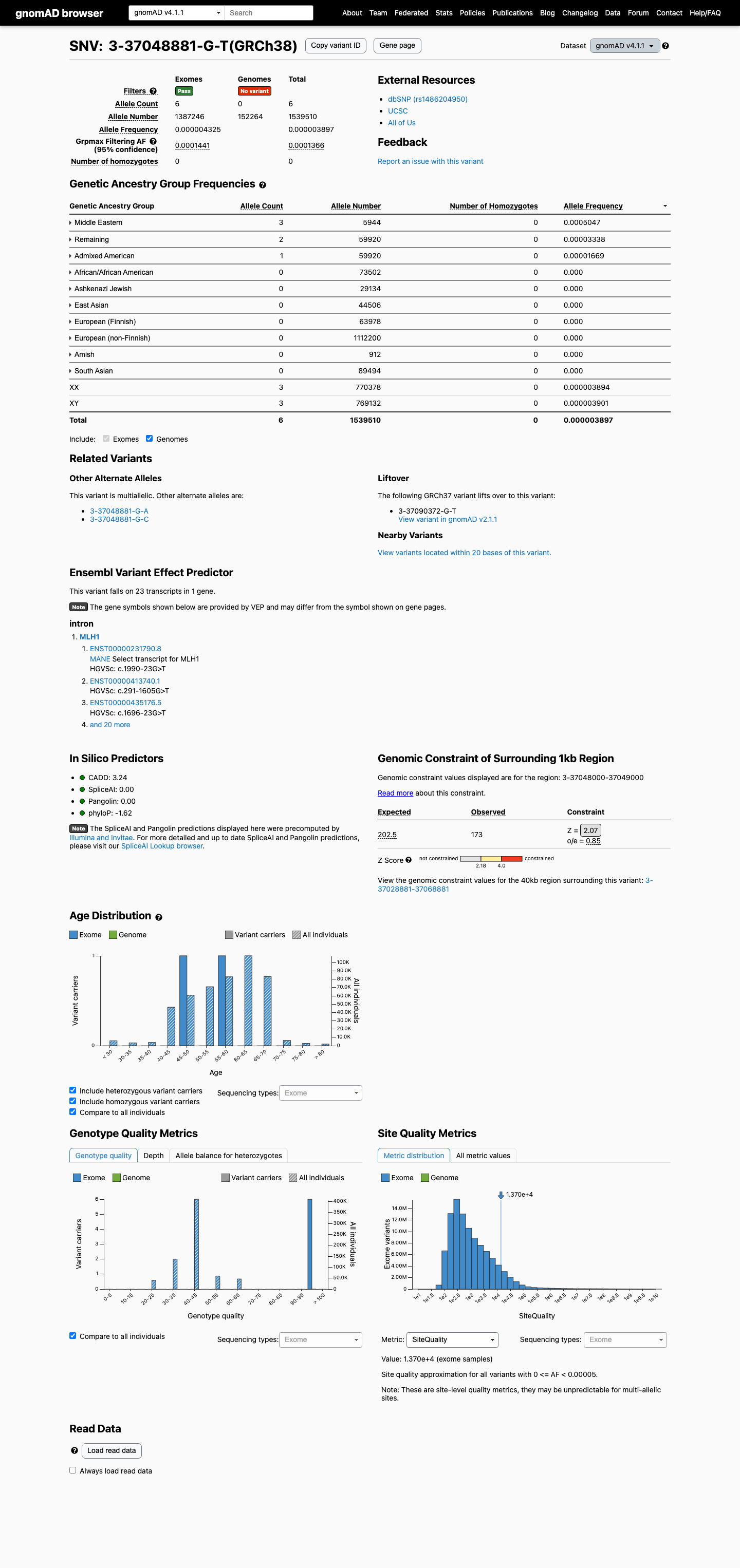
In gnomAD v4.1, this variant is very rare overall at 6/1539510 alleles (AF 0.00000389734), which is below the MLH1 PM2_Supporting threshold of 0.00002, but the gnomAD v4.1 grpmax filtering allele frequency is 0.00013658 in the Middle Eastern population, which falls within the MLH1 BS1 range of 0.0001 to less than 0.001; gnomAD v2.1 also shows only 1/250612 alleles (AF 0.00000399023).
No variant-specific functional or constitutional RNA assay evidence was identified for this variant, so PS3 and BS3 were not assessed.
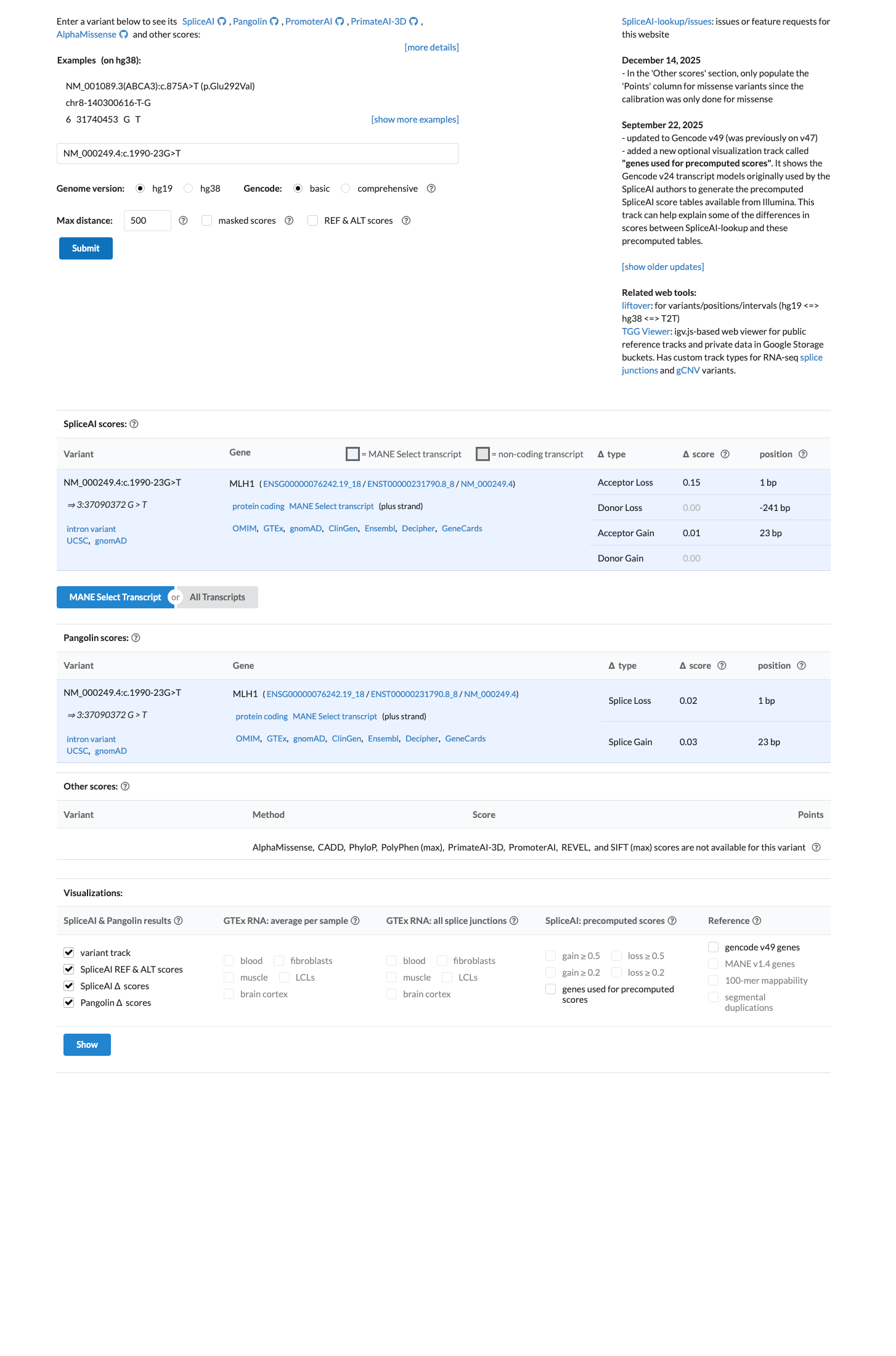
SpliceAI predicts no significant splice impact with a maximum delta score of 0.00, which is below the BP4 threshold of 0.1 and below the PP3 threshold of 0.2, and the intronic position c.1990-23 is beyond the BP7 distance threshold of -21.