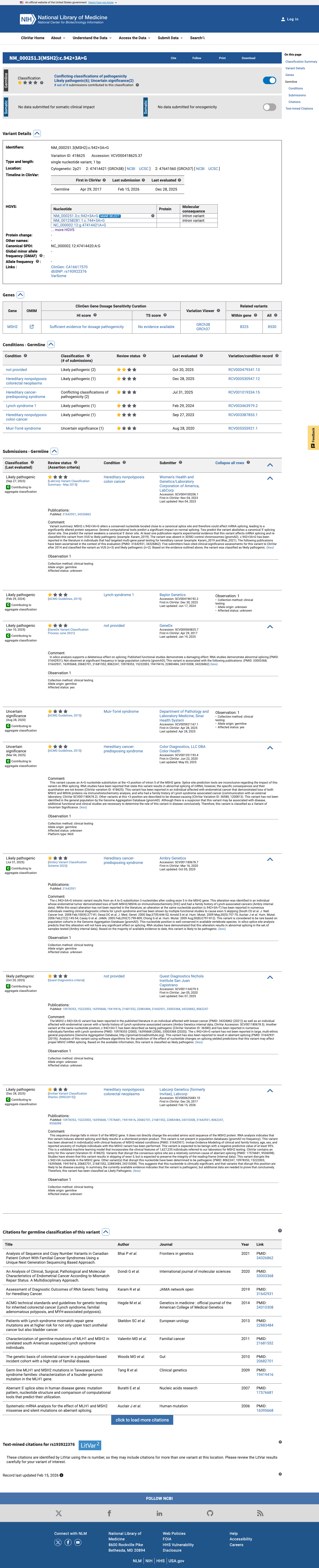
The MSH2 c.942+3A>G (p.?) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar with predominantly likely pathogenic submissions, although some submissions classify it as uncertain significance.
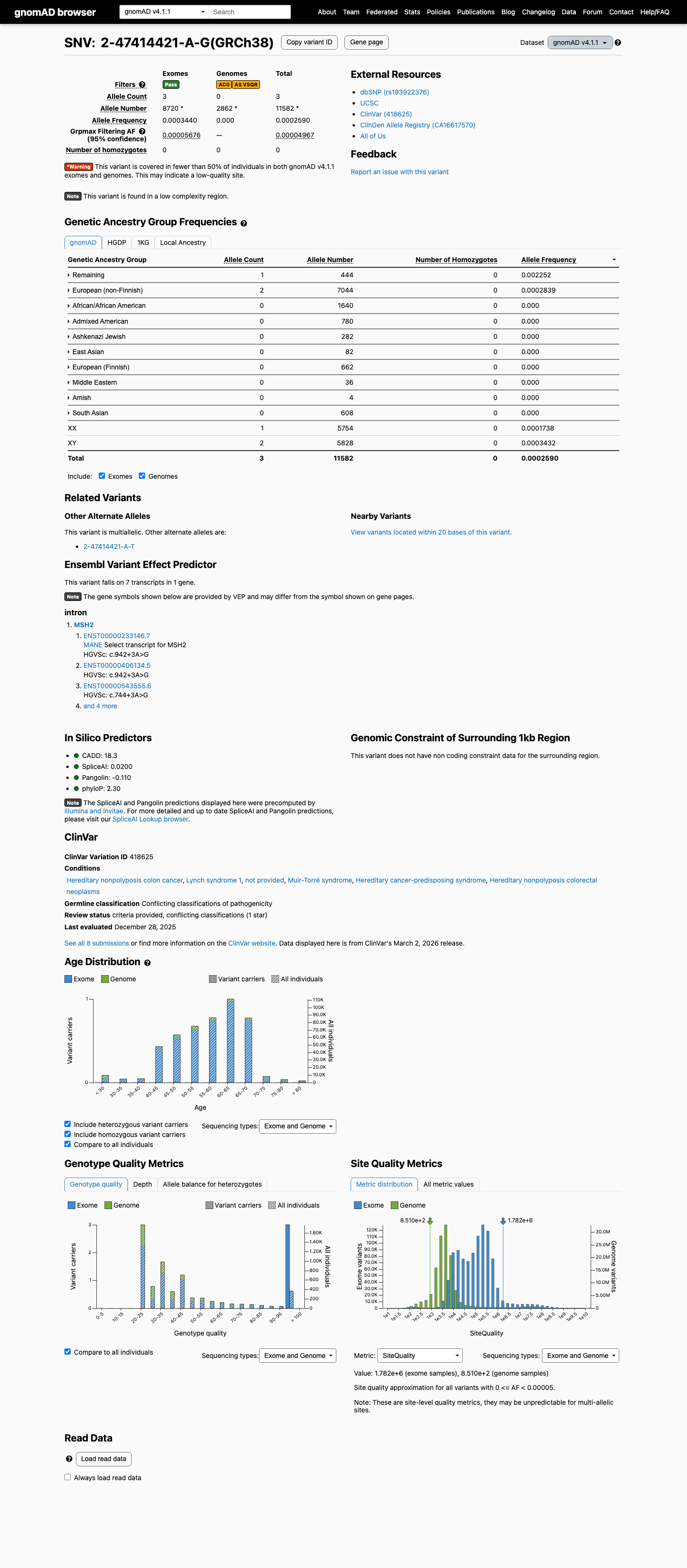
clinvar ↗This variant is absent from gnomAD v2.1 but is present in gnomAD v4.1 at 3/11,582 alleles (total AF 0.00025902) with grpmax FAF 0.00004967, which is above the MSH2 VCEP PM2 threshold of less than 0.00002 and below the BS1 threshold of at least 0.0001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Published RNA studies reported that this variant disrupts MSH2 donor splicing and generates an exon 5-skipped transcript, supporting a true splice defect despite its non-canonical +3 position.
PMID:10978353 ↗ PMID:16395668 ↗SpliceAI predicts a low splice effect with a maximum delta score of 0.02, which is below the MSH2 VCEP PP3 threshold of at least 0.2 and within the BP4 threshold of 0.1 or less, but this in silico result is inconsistent with the published RNA evidence.
spliceai ↗ cspec ↗ PMID:10978353 ↗ PMID:16395668 ↗