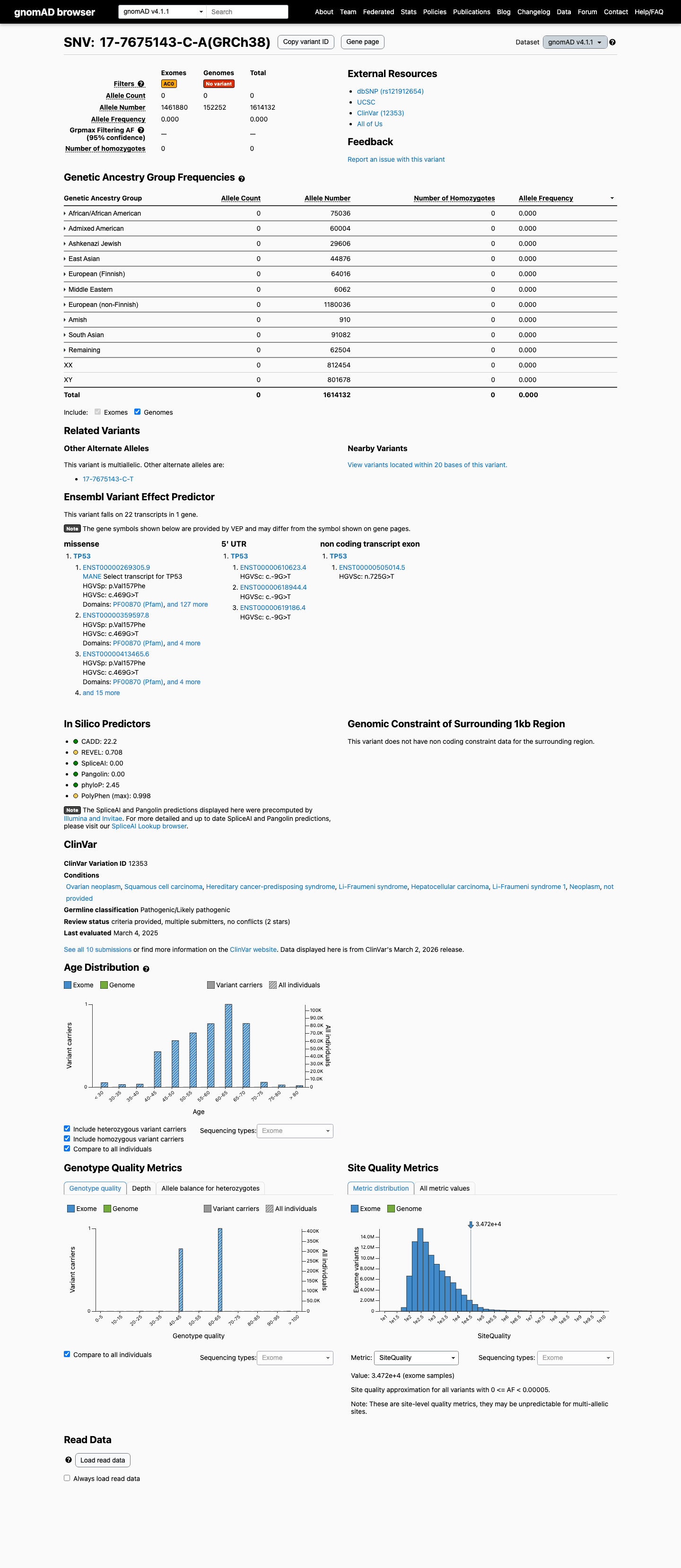
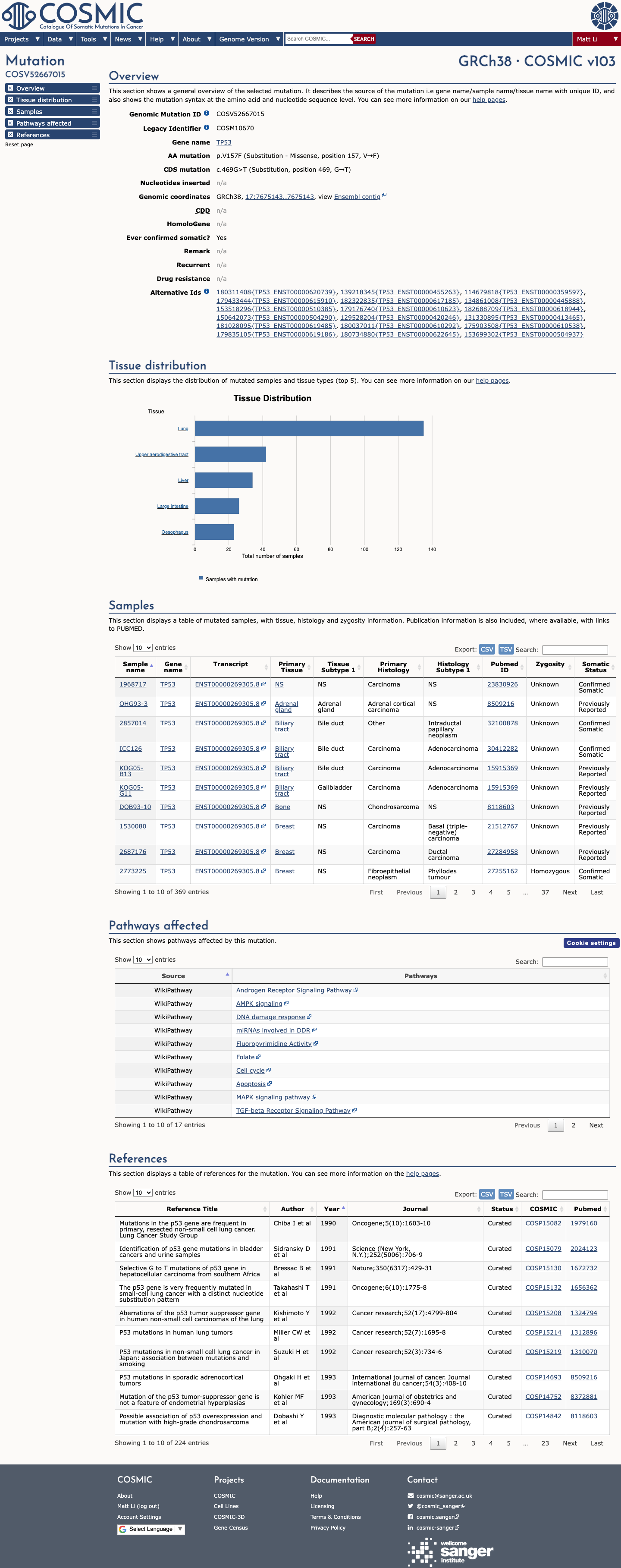
The TP53 c.469G>T (p.Val157Phe) variant has been observed in somatic cancers in COSMIC (COSV52667015; n=366) and has not been reported in ClinVar.
This variant is absent from gnomAD v2.1 and absent from gnomAD v4.1 (0/1614132 alleles; AF 0.00000%), which is below the TP53 PM2_Supporting threshold of less than 0.00003.
TP53 expert panel functional data classify p.Val157Phe as PS3 because the variant is non-functional in Kato, shows loss of function in Funk and Kotler, and is loss-of-function by the majority of eligible assays.
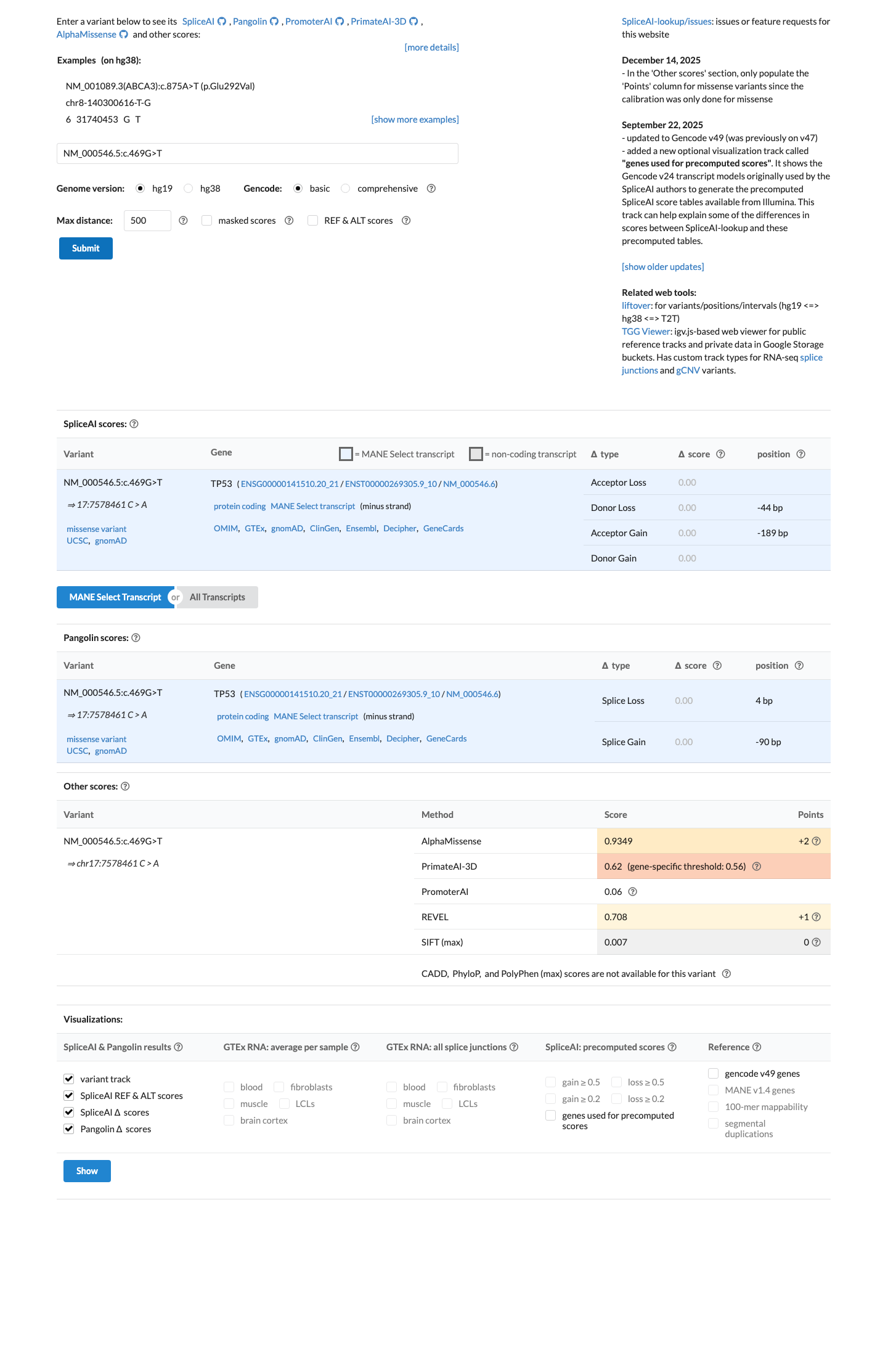
Cancer Hotspots shows p.Val157Phe in 67 tumors at the significant TP53 V157 hotspot residue (77 tumors at the residue overall), supporting PM1, while the TP53 bioinformatic worksheet assigns no PP3/BP4 evidence and SpliceAI predicts no splice impact (max delta score 0.00).