Classification rationale
1
2
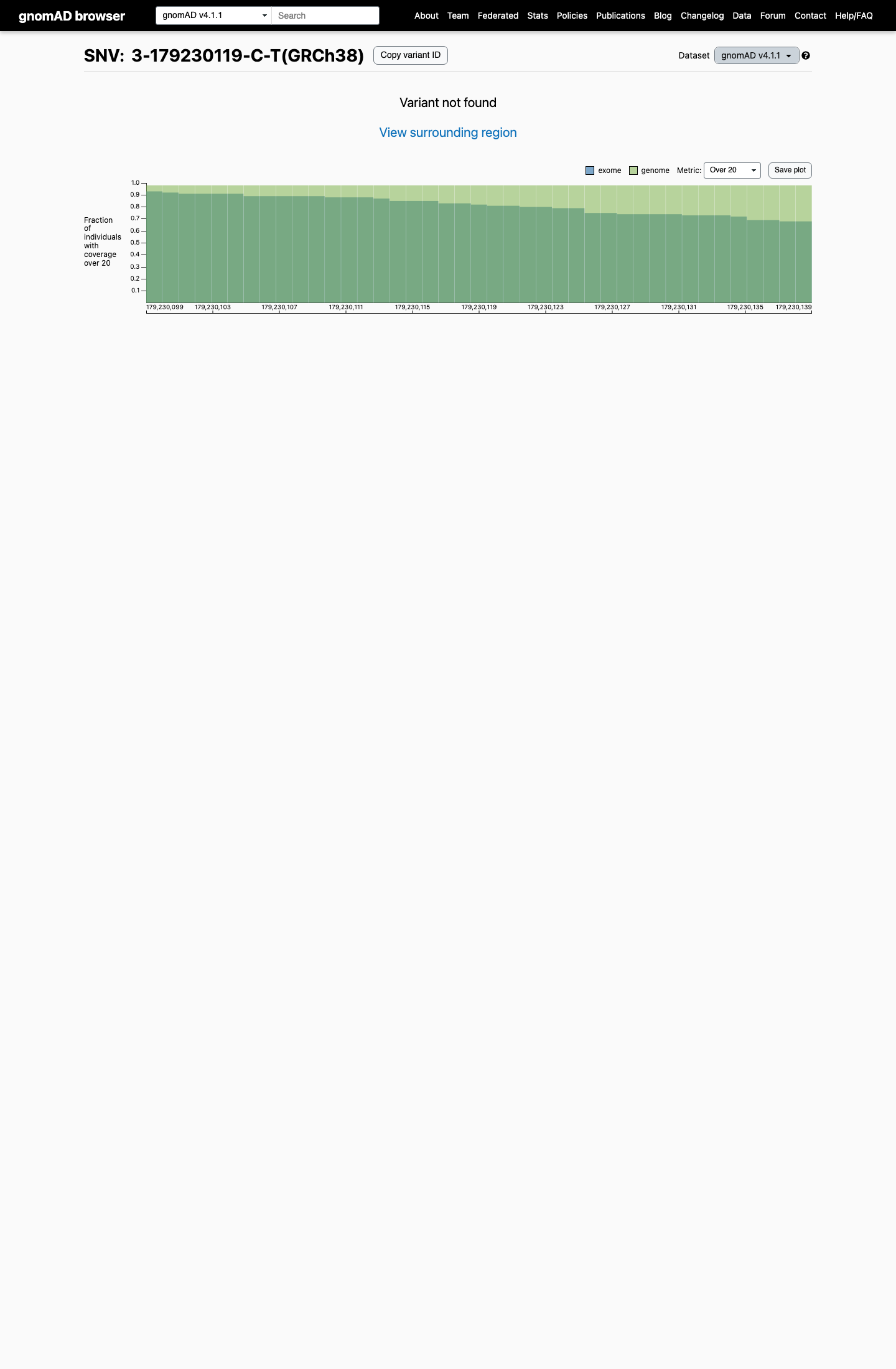
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is within the Brain Malformations VCEP PM2_Supporting threshold of at most 1 occurrence in population data and below the BS1 (>0.0185%) and BA1 (>0.0926%) thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
The variant lies within the PIK3CA kinase domain approved for PM1_Supporting by the Brain Malformations VCEP (amino acids 797-1068), supporting location in a critical functional region.
cspec ↗4
SpliceAI predicts no significant splice impact for this variant with a maximum delta score of 0.10, although PP3 and BP4 are not applicable to this nonsense variant under the Brain Malformations VCEP specifications.
spliceai ↗ cspec ↗