Classification rationale
1
2
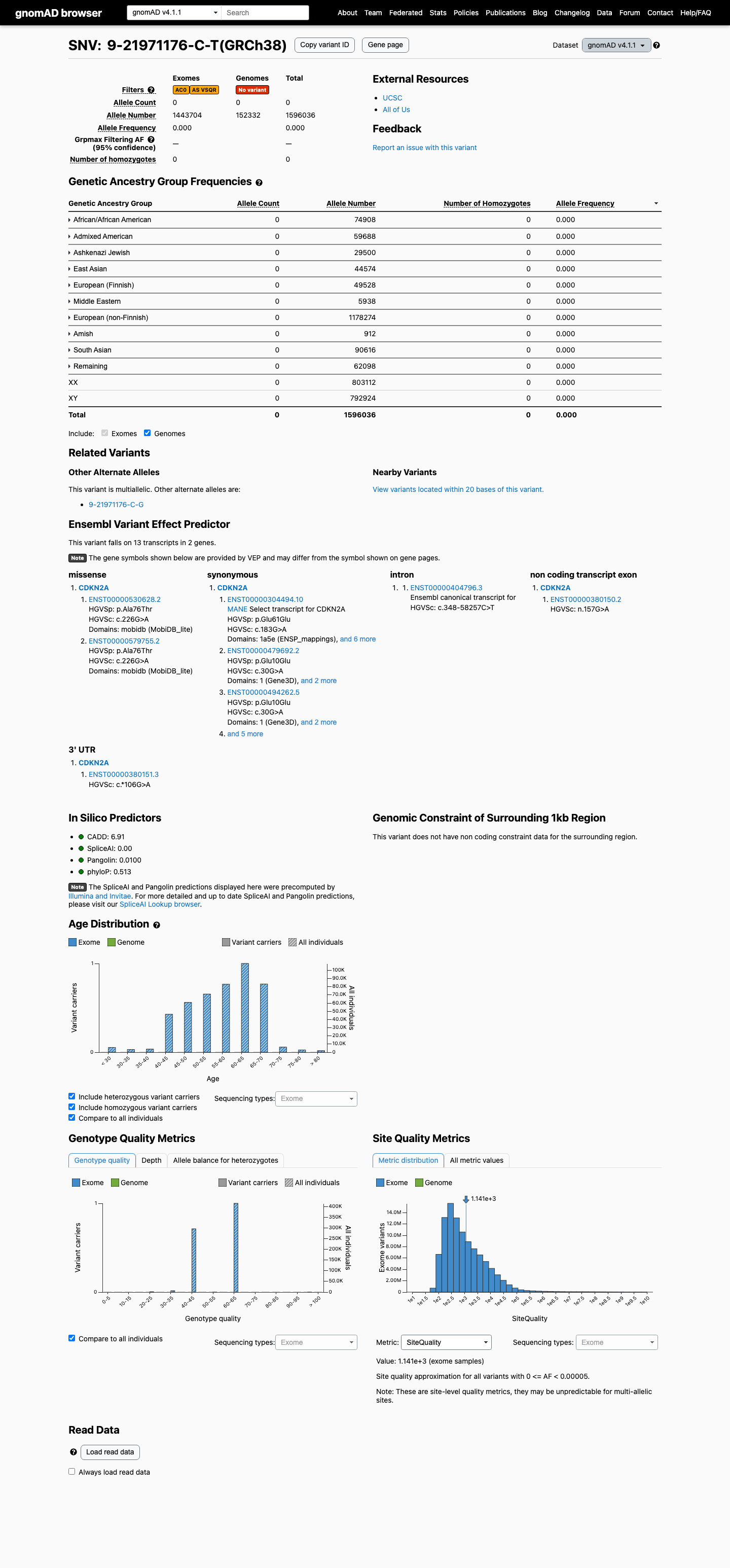
This variant is absent from gnomAD v2.1 and observed at 0/1,596,036 alleles in gnomAD v4.1 (AF 0.00000%), which is below the 0.1% PM2 rarity threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
In silico data support a silent variant without splice effect: the protein consequence is p.(Glu61=), SpliceAI predicts no significant splice impact with a maximum delta score of 0.00, and REVEL is 0.083.
spliceai ↗