Classification rationale
1
2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is below the MSH2 PM2 threshold of <0.00002 (<1 in 50,000 alleles).
gnomad_v2 ↗ gnomad_v4 ↗3
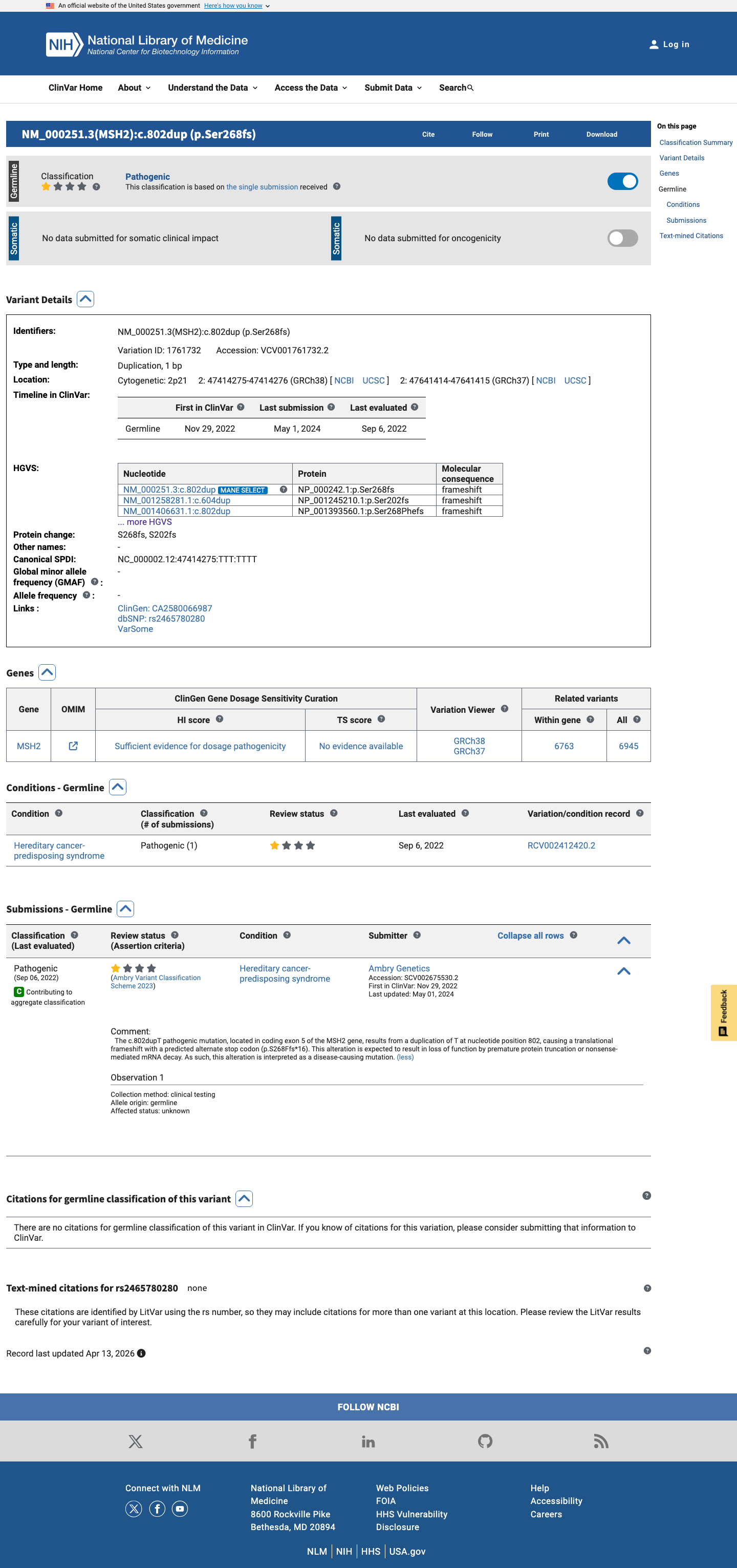
The duplication causes a frameshift with a premature stop codon at p.(Ser268PhefsTer16), and this truncation is upstream of the MSH2 codon 891 cutoff for PVS1 Very Strong in the InSiGHT specification.
cspec ↗4
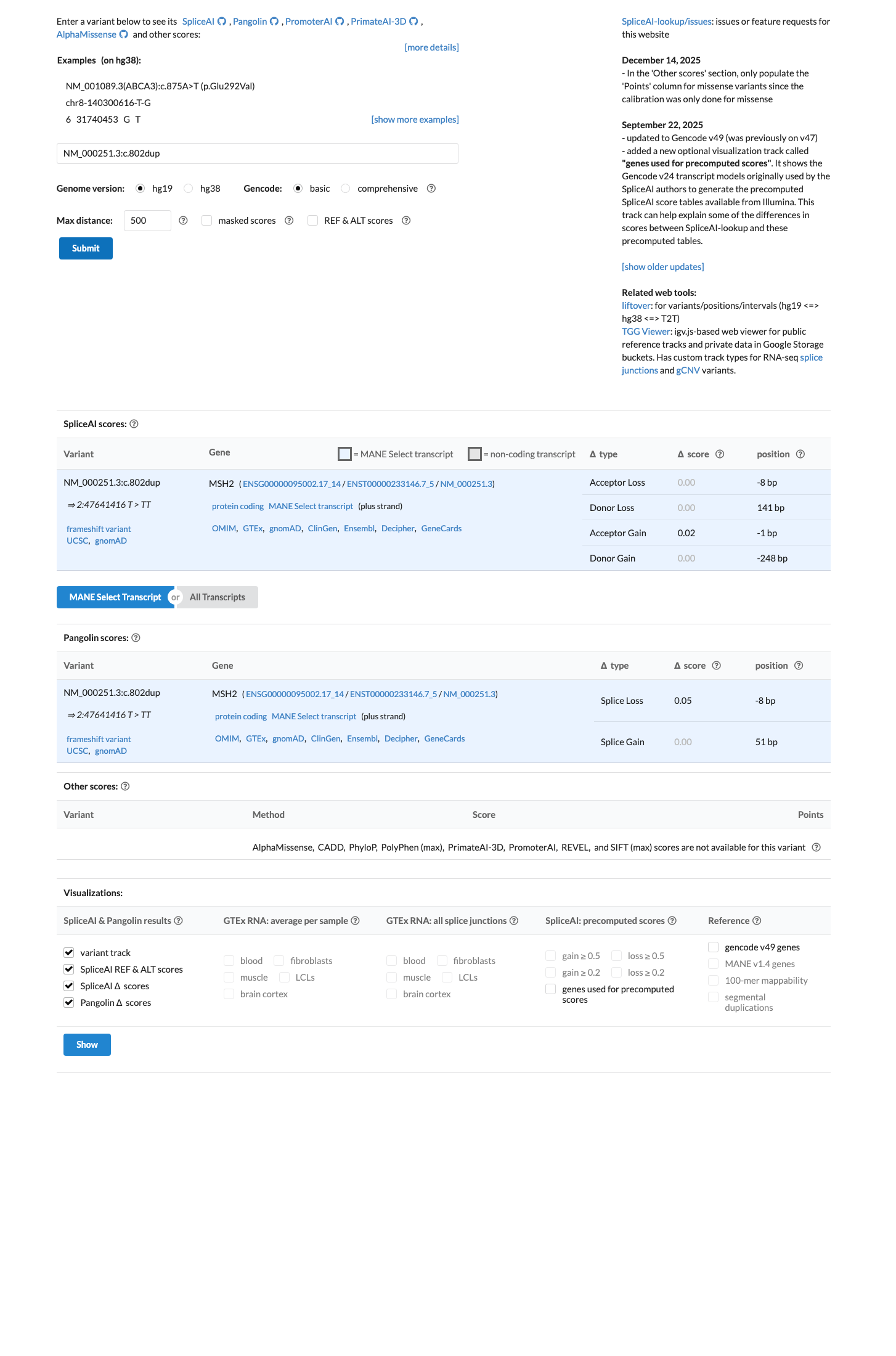
SpliceAI predicts no significant splice impact with a maximum delta score of 0.06, indicating no additional splice effect is predicted beyond the truncating consequence of the duplication.
spliceai ↗