Classification rationale
1
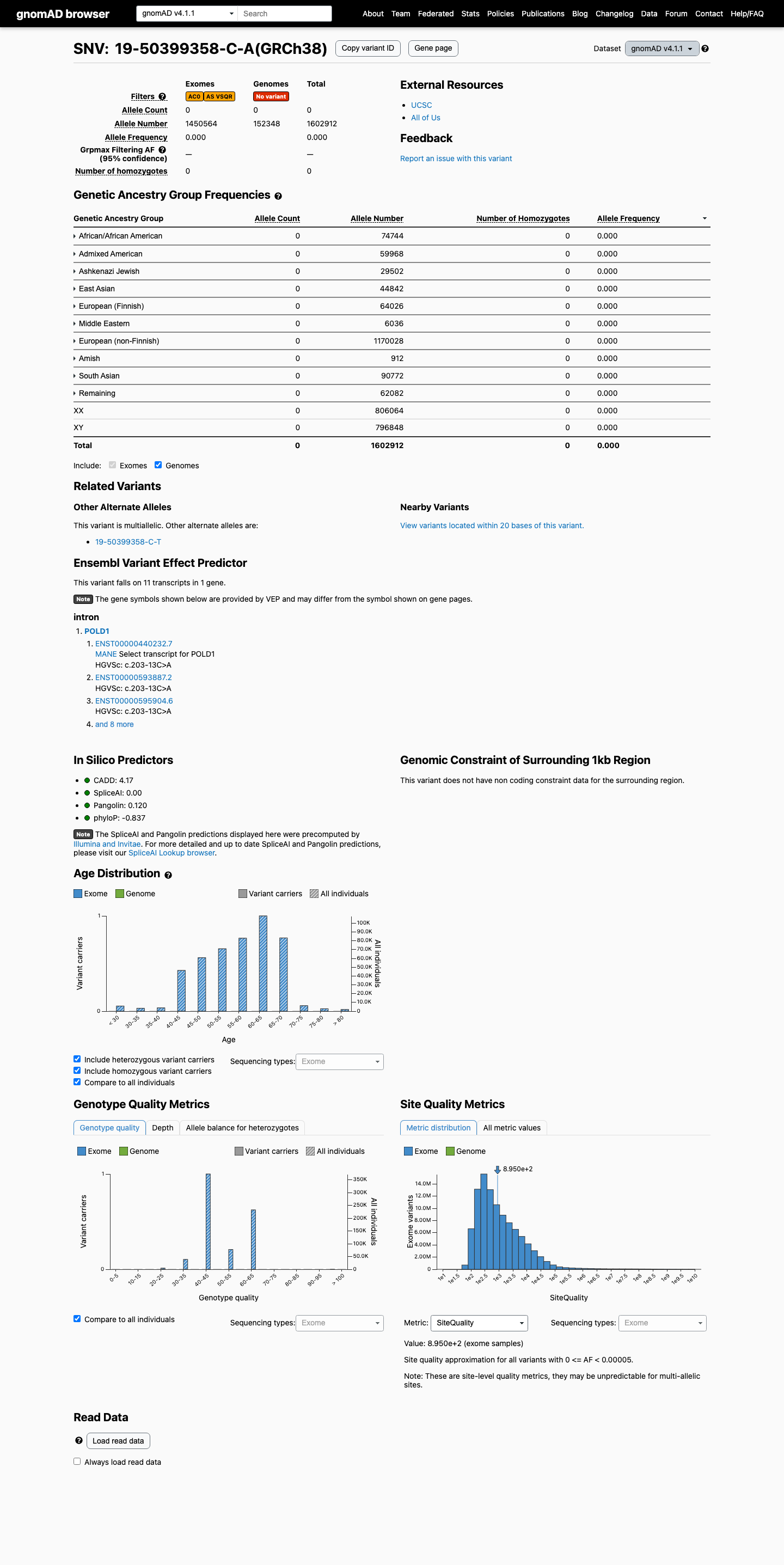
The POLD1 c.203-13C>A (p.?) variant has not been observed in somatic cancers in COSMIC and has not been reported in ClinVar.
2
This variant is absent from gnomAD v2.1 and shows 0/1,602,912 alleles in gnomAD v4.1 (AF 0.0%), which is below the 0.1% rarity threshold and supports PM2 at supporting strength.
3
SpliceAI predicts no significant splice impact for this intronic change, with a maximum delta score of 0.15, which supports BP4 and does not support PP3.