Classification rationale
1
2
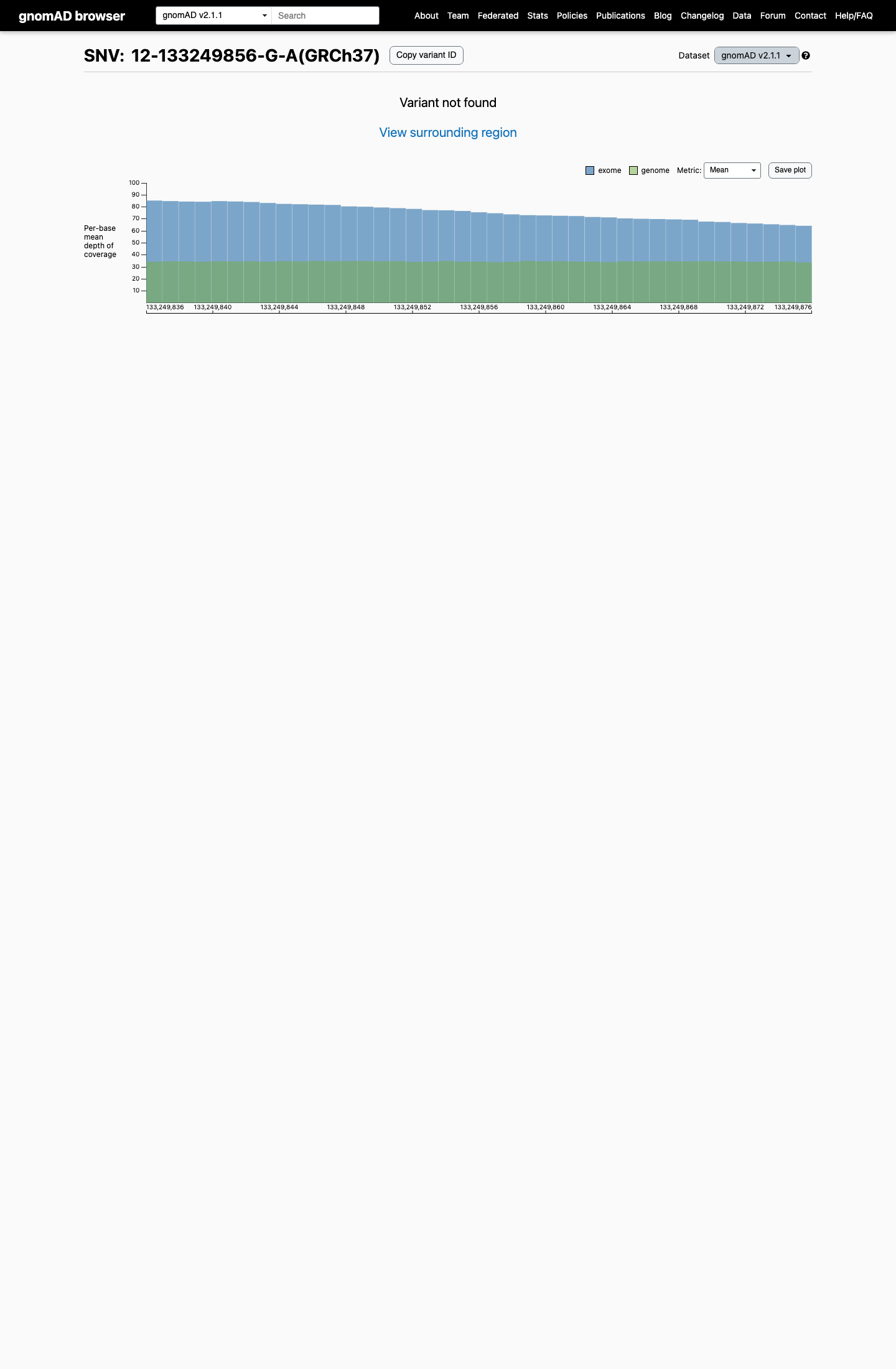
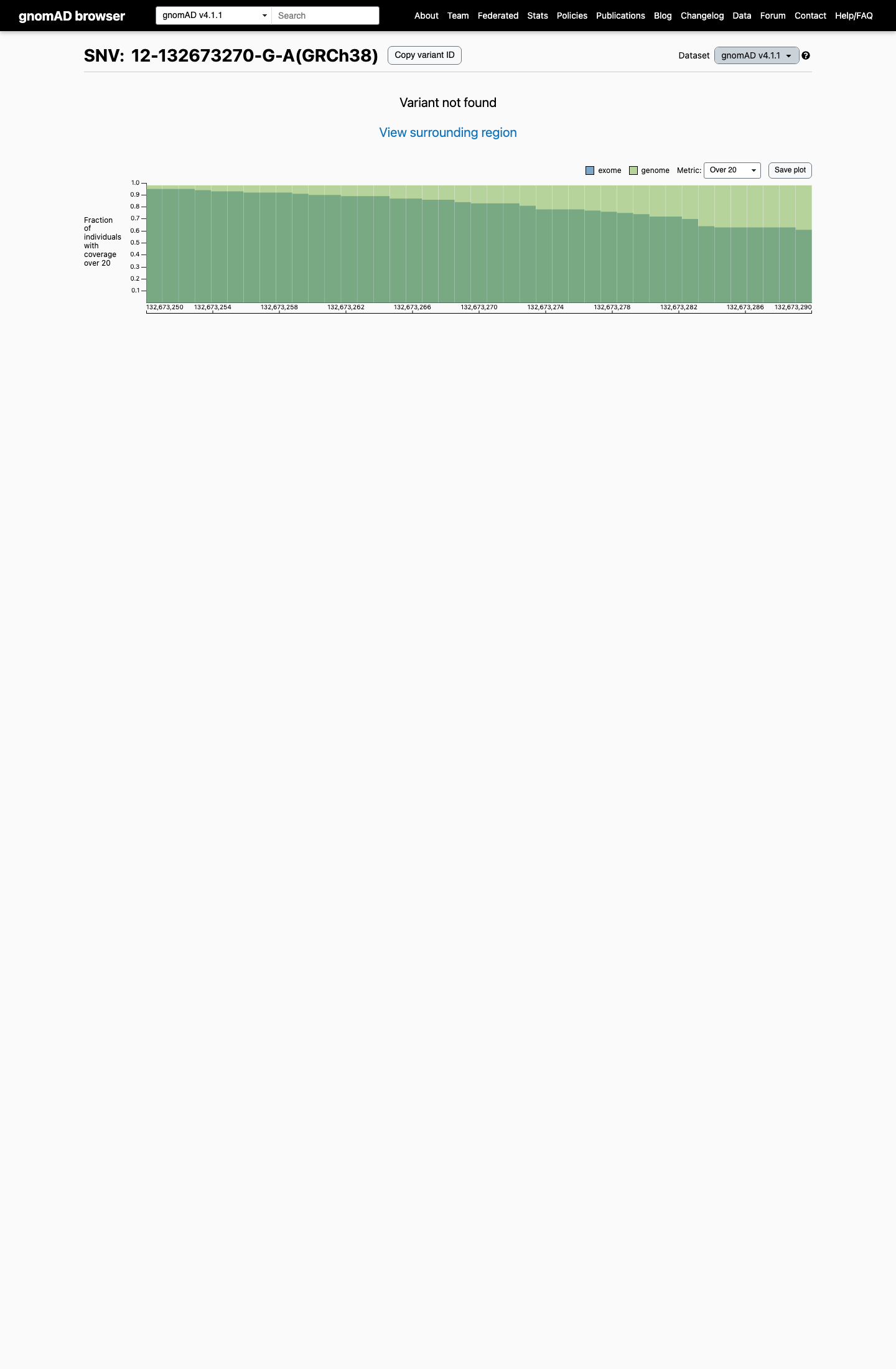
The variant is absent from gnomAD v2.1 and gnomAD v4.1, with an observed population frequency of 0 that is below the project's PM2 threshold of 0.1%.
gnomad_v2 ↗ gnomad_v4 ↗3
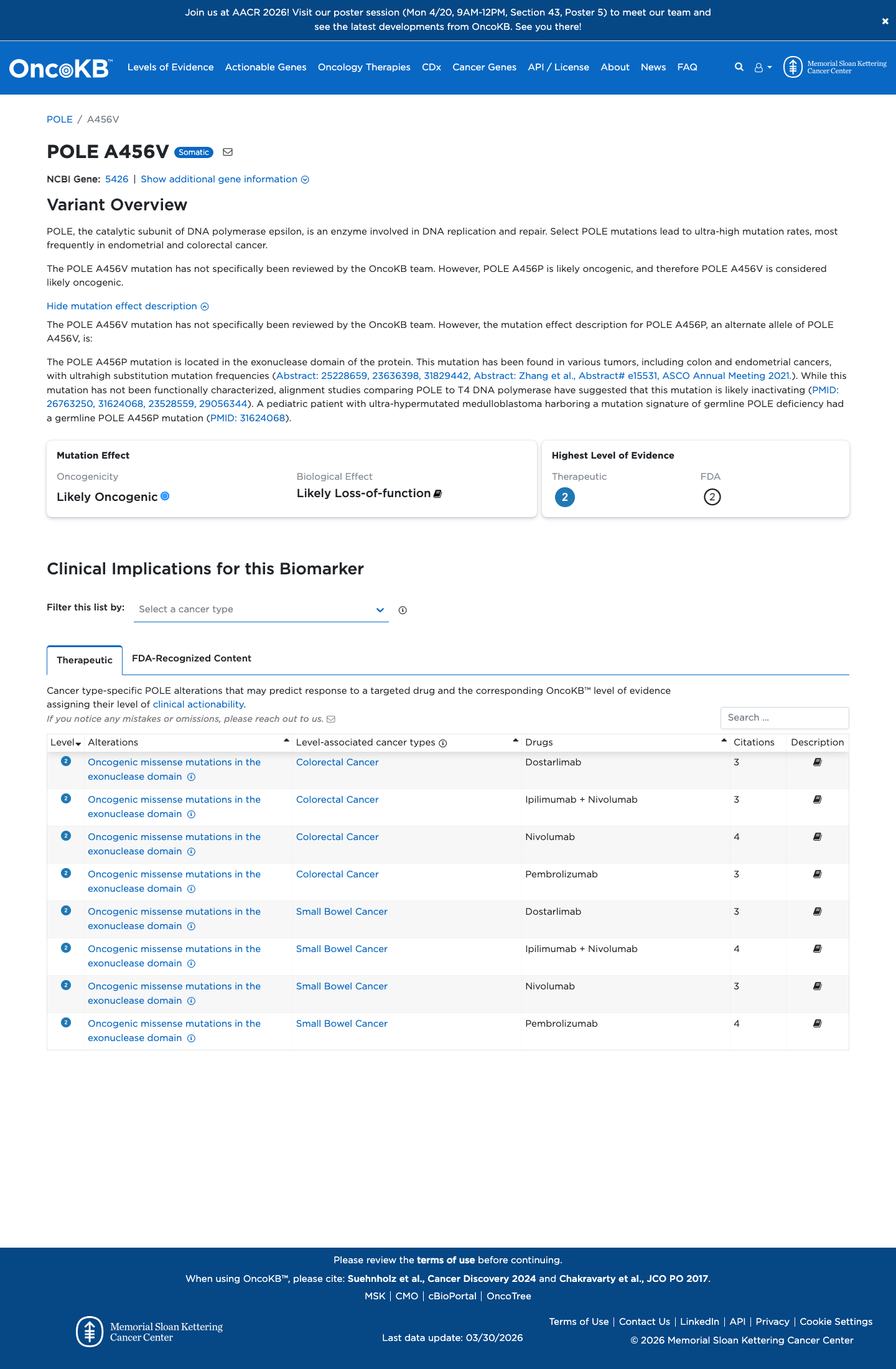
In León-Castillo Supplementary Table S3, p.Ala456Val is listed as likely disease causing with 0 benign in silico results, meeting the local POLE PP3 rule; SpliceAI predicts no significant splice impact with a maximum delta score of 0.10.
spliceai ↗