Classification rationale
1
The ATM c.3137T>C (p.Leu1046Pro) variant has not been observed in somatic cancers in COSMIC and has not been reported in ClinVar.
2
This variant is absent from gnomAD v2.1 and is present at very low frequency in gnomAD v4.1 overall (AF 1.85893e-06; 3/1613832 alleles; 0 homozygotes), with the highest observed frequency in the South Asian population at 2.19587e-05 (2/91080 alleles; 0 homozygotes).
3
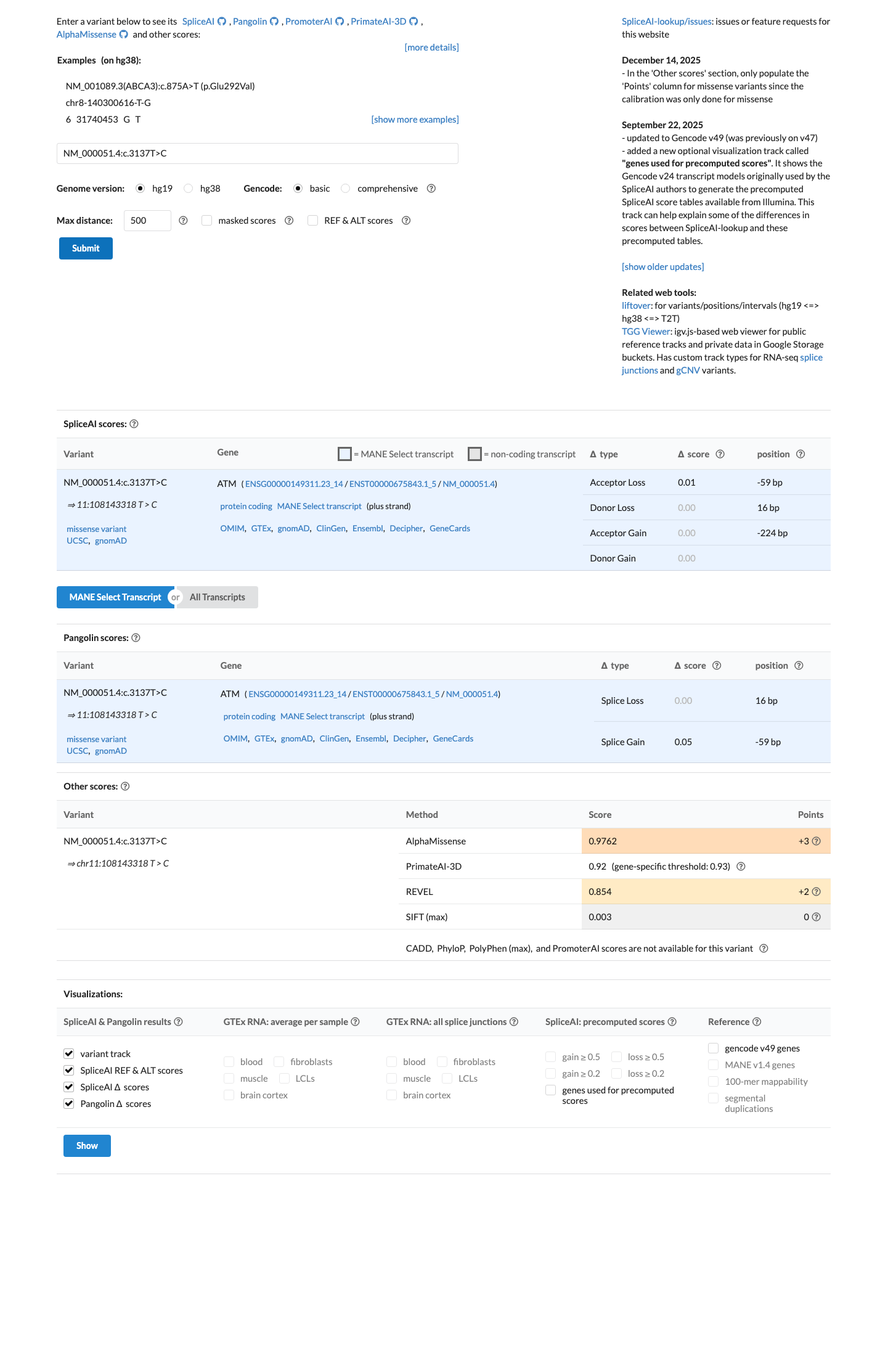
In silico evidence supports a deleterious missense effect, with REVEL 0.854 exceeding the ATM PP3 threshold of 0.7333, while SpliceAI predicts no significant splice impact (maximum delta score 0.01).