Classification rationale
1
2
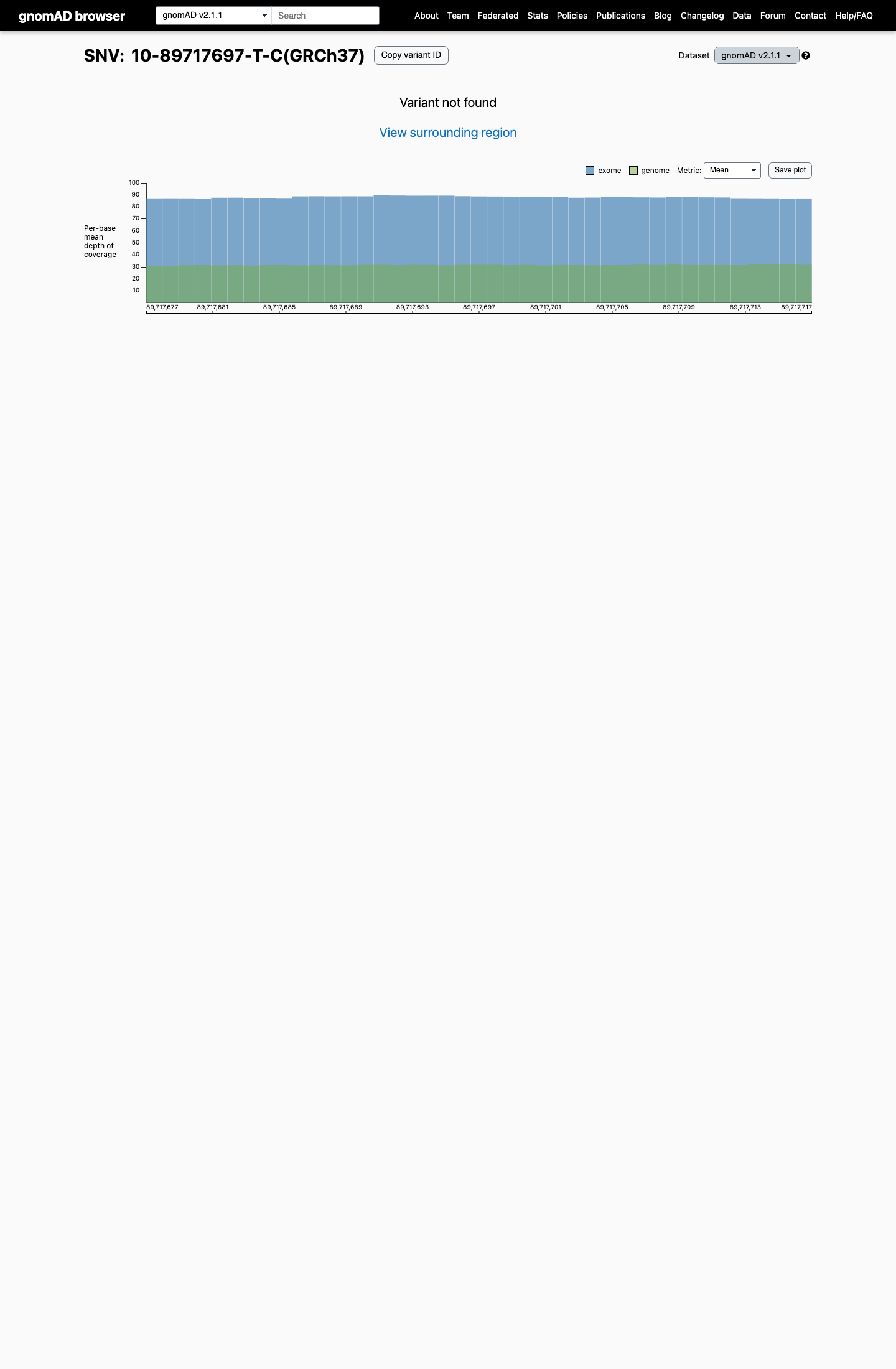
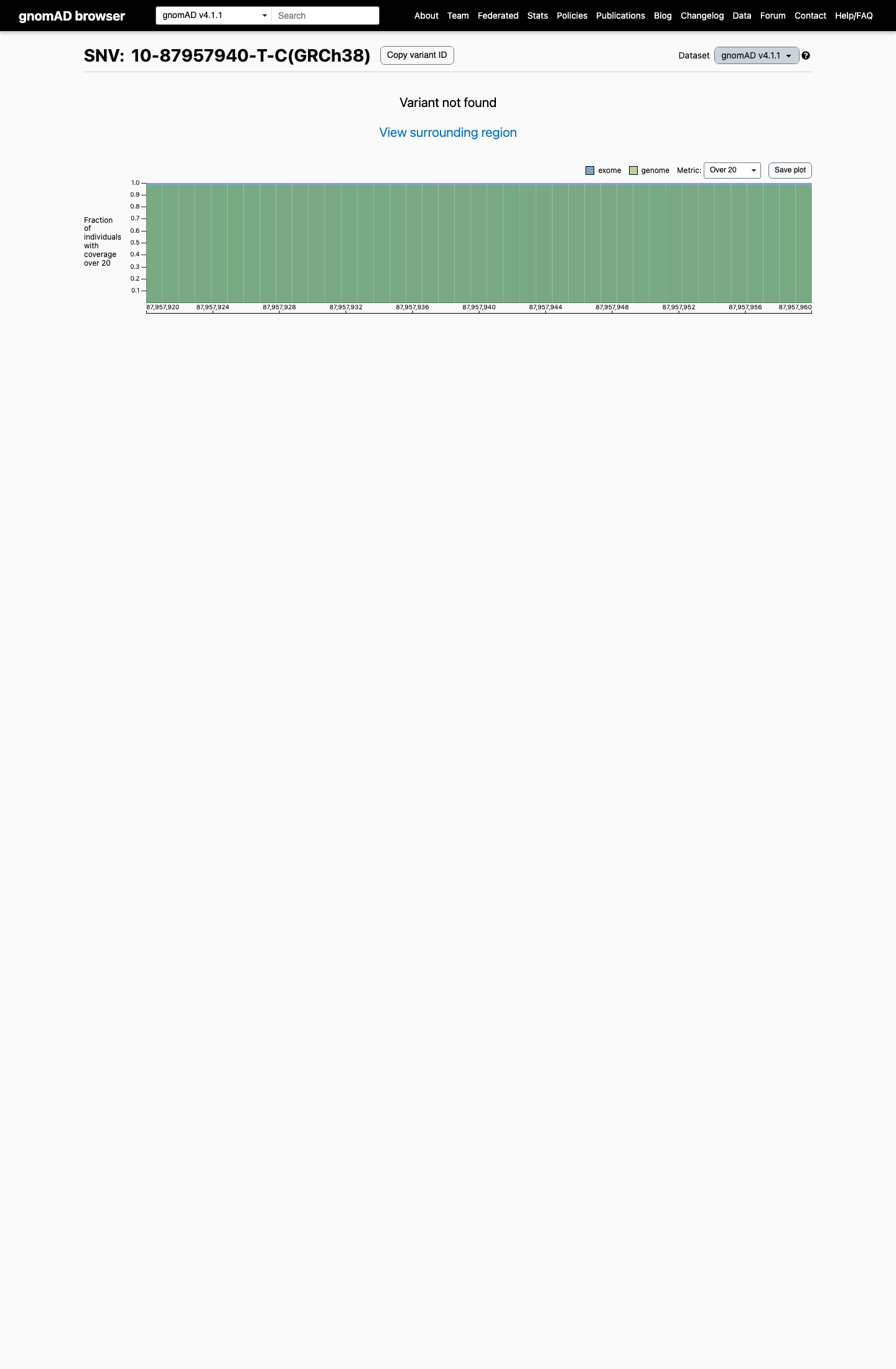
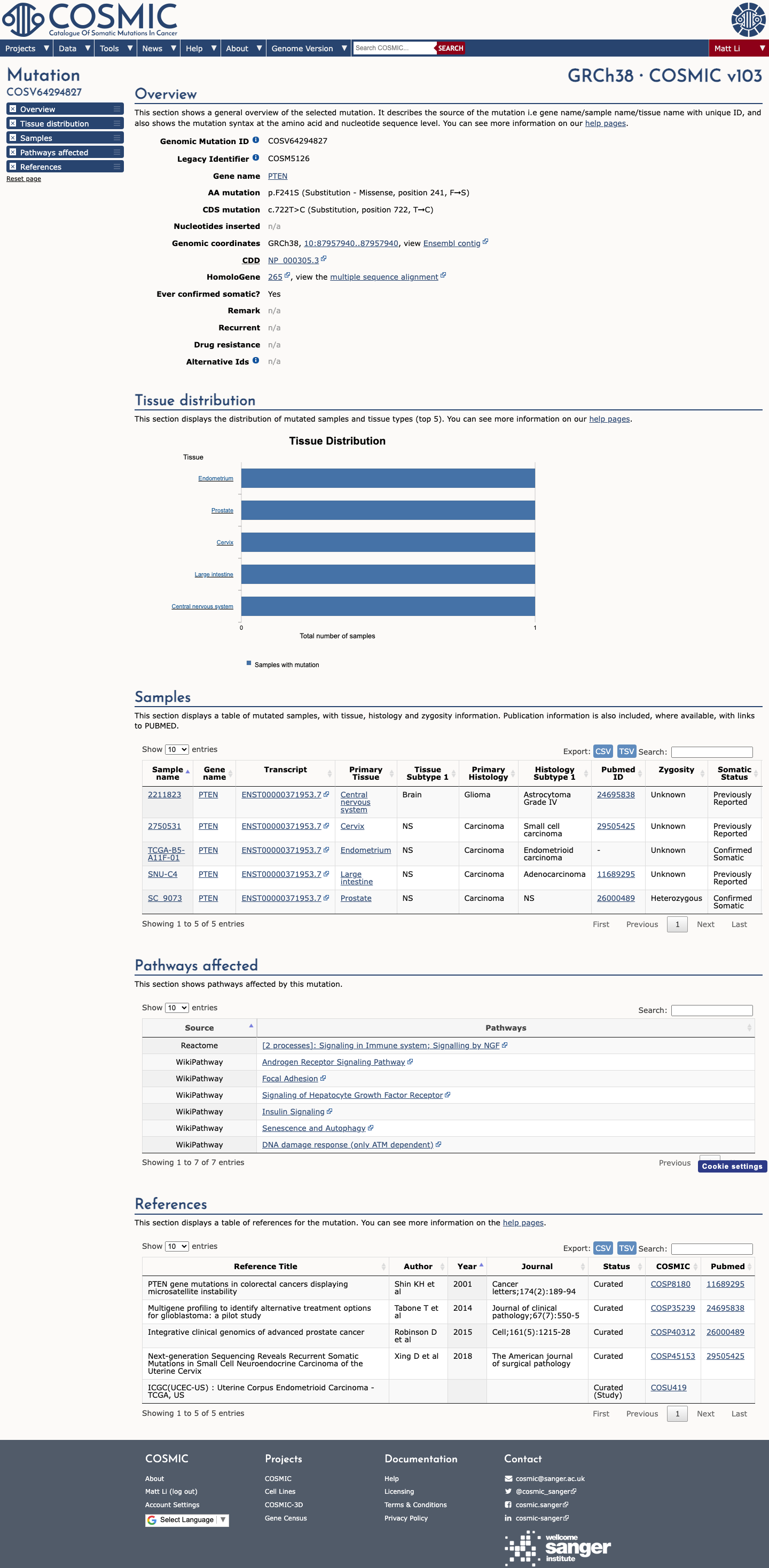
This variant is absent from gnomAD v2.1 and gnomAD v4.1, with an observed allele frequency of 0, which is below the PTEN Expert Panel PM2_Supporting threshold of <0.00001 (0.001%).
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
Functional studies support a damaging effect on PTEN, including a Mighell et al. phosphatase assay result of Cum_score -2.399 with High_conf=True, which is below the PTEN Expert Panel PS3_Moderate threshold of <= -1.11; OncoKB also describes the variant as Likely Oncogenic with a likely loss-of-function effect.
oncokb ↗4
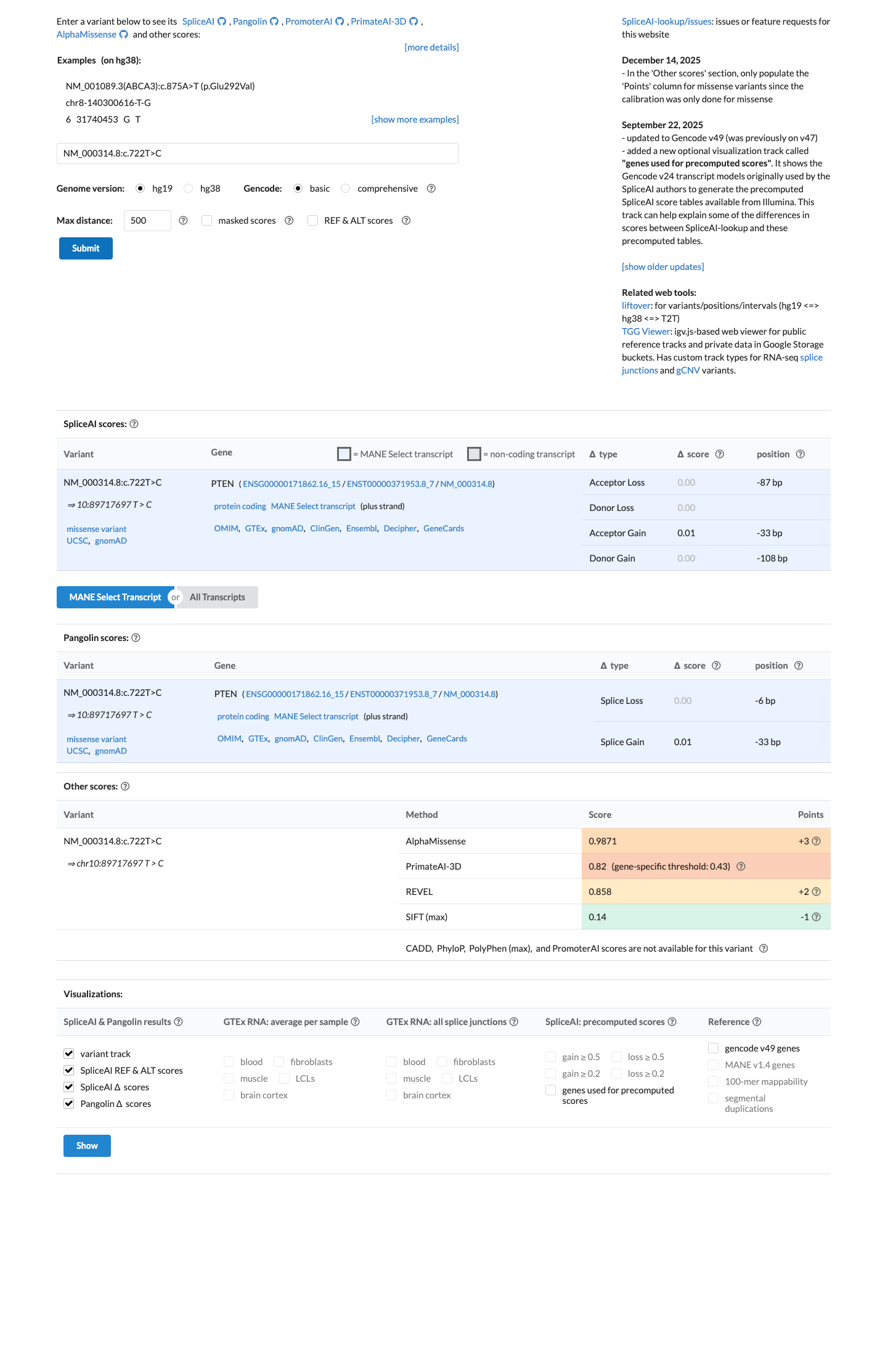
Computational evidence supports a deleterious effect for the missense change because the REVEL score is 0.858, above the PTEN Expert Panel PP3 threshold of >0.7, while SpliceAI predicts no significant splice effect with a maximum delta score of 0.01.
spliceai ↗ cspec ↗