Classification rationale
1
2
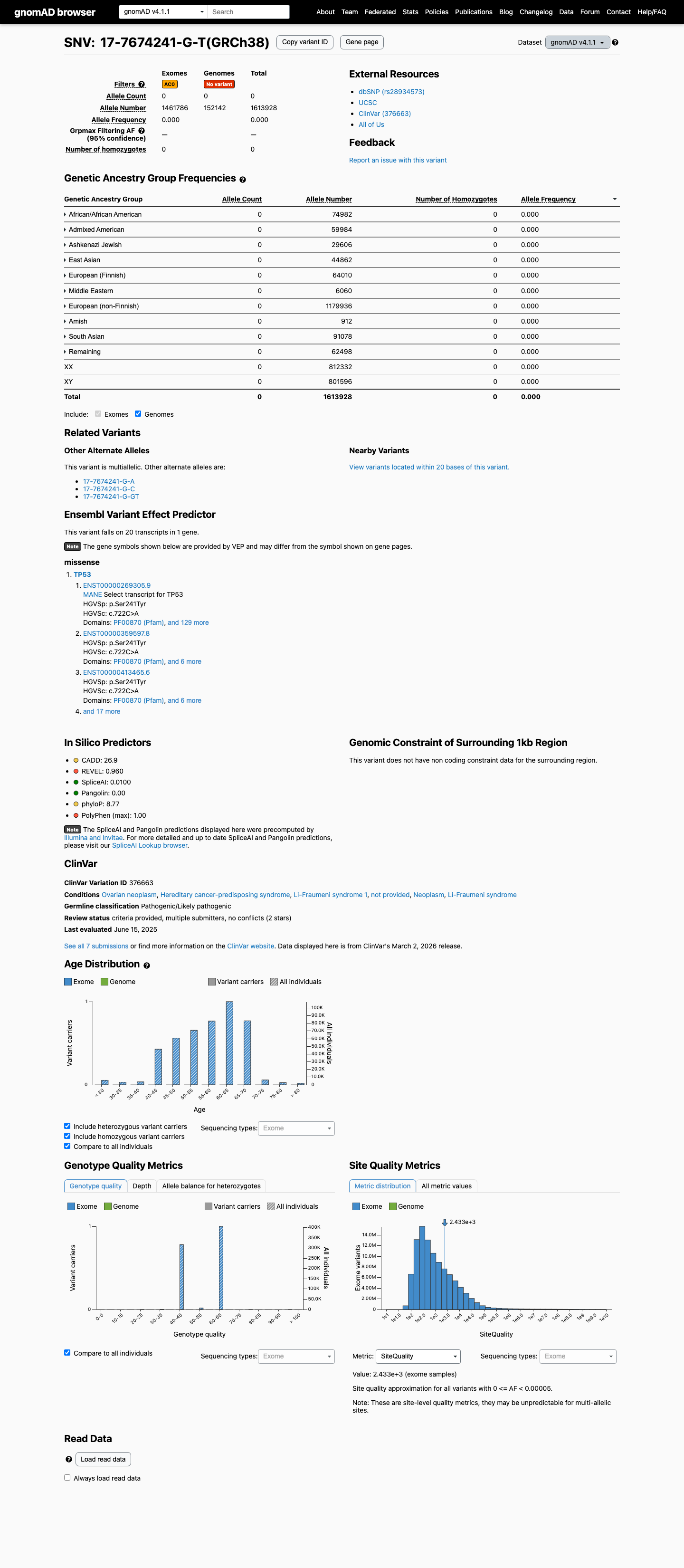
This variant is absent from gnomAD v2.1 and has 0/1,613,928 alleles in gnomAD v4.1, which is below the TP53 VCEP PM2_Supporting threshold of 0.00003.
gnomad_v2 ↗ gnomad_v4 ↗3
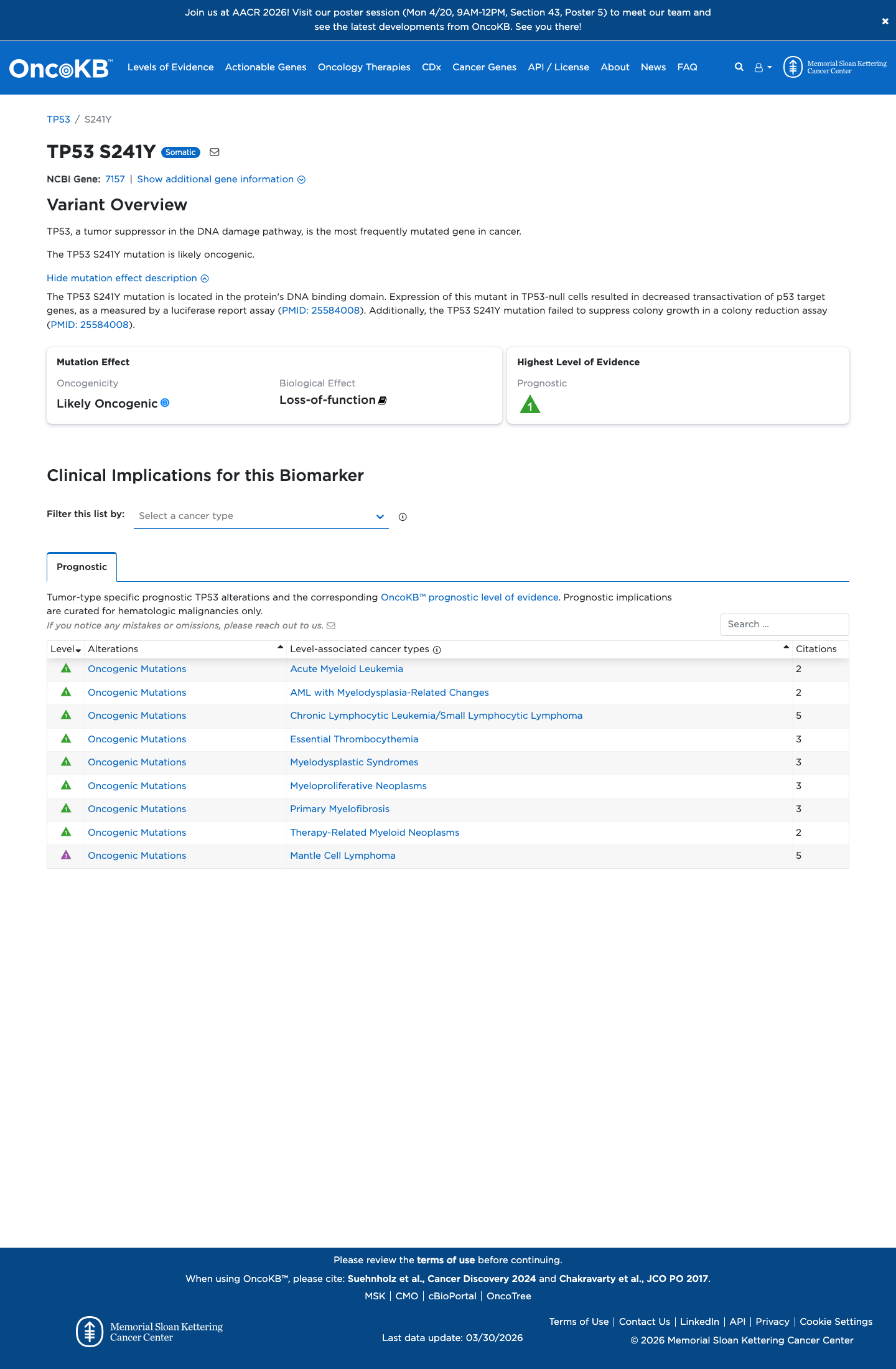
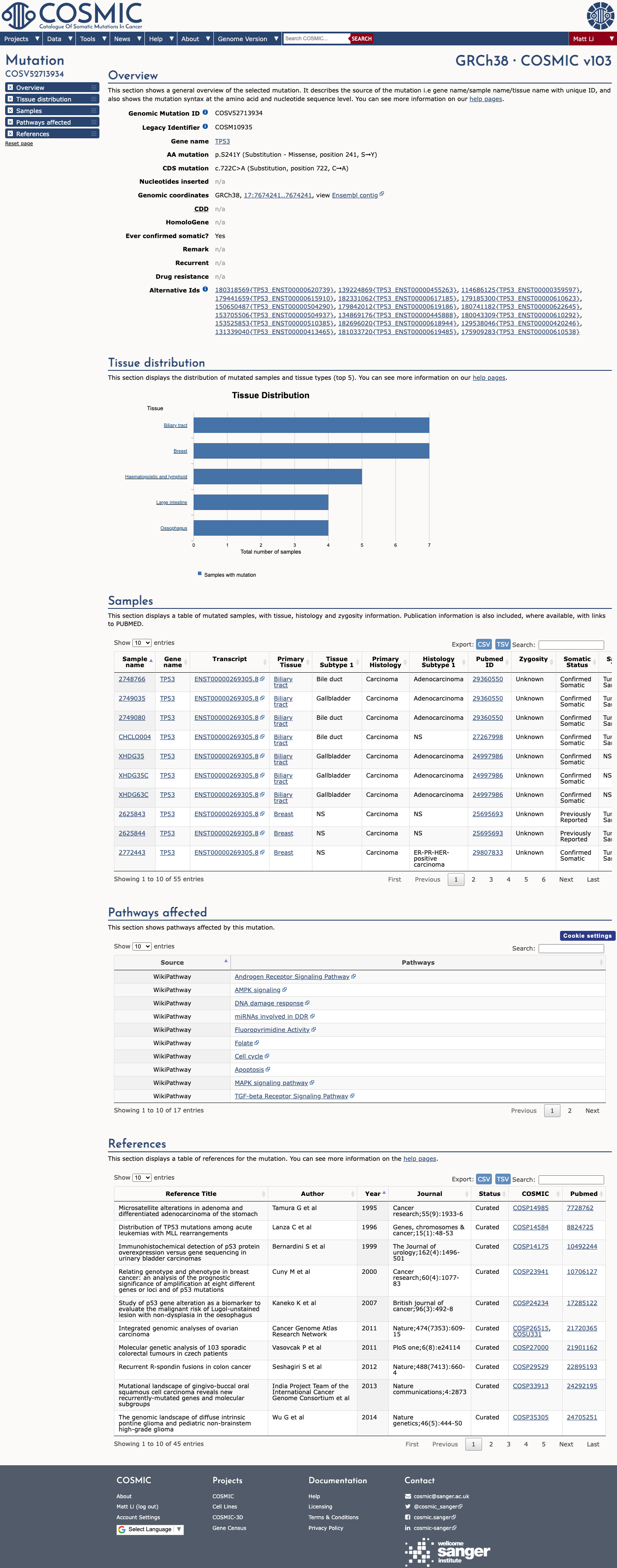
TP53 VCEP functional data support loss of p53 function for p.Ser241Tyr, with non-functional Kato results and loss of function in the majority of other eligible assays, supporting PS3.
oncokb ↗ PMID:12826609 ↗ PMID:25584008 ↗4
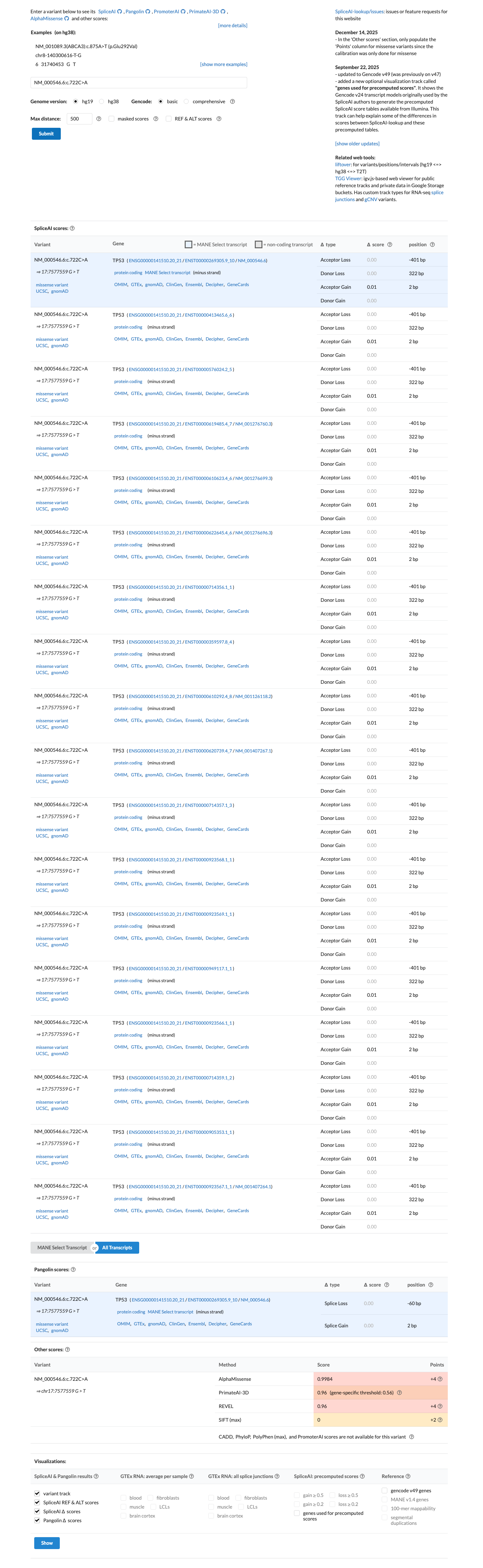
TP53-specific in silico evidence supports a deleterious effect, with Align-GVGD Class C65, BayesDel 0.544682, a pre-assigned PP3_moderate code, and SpliceAI showing no significant predicted splice impact (max delta score 0.01).
spliceai ↗