Classification rationale
1
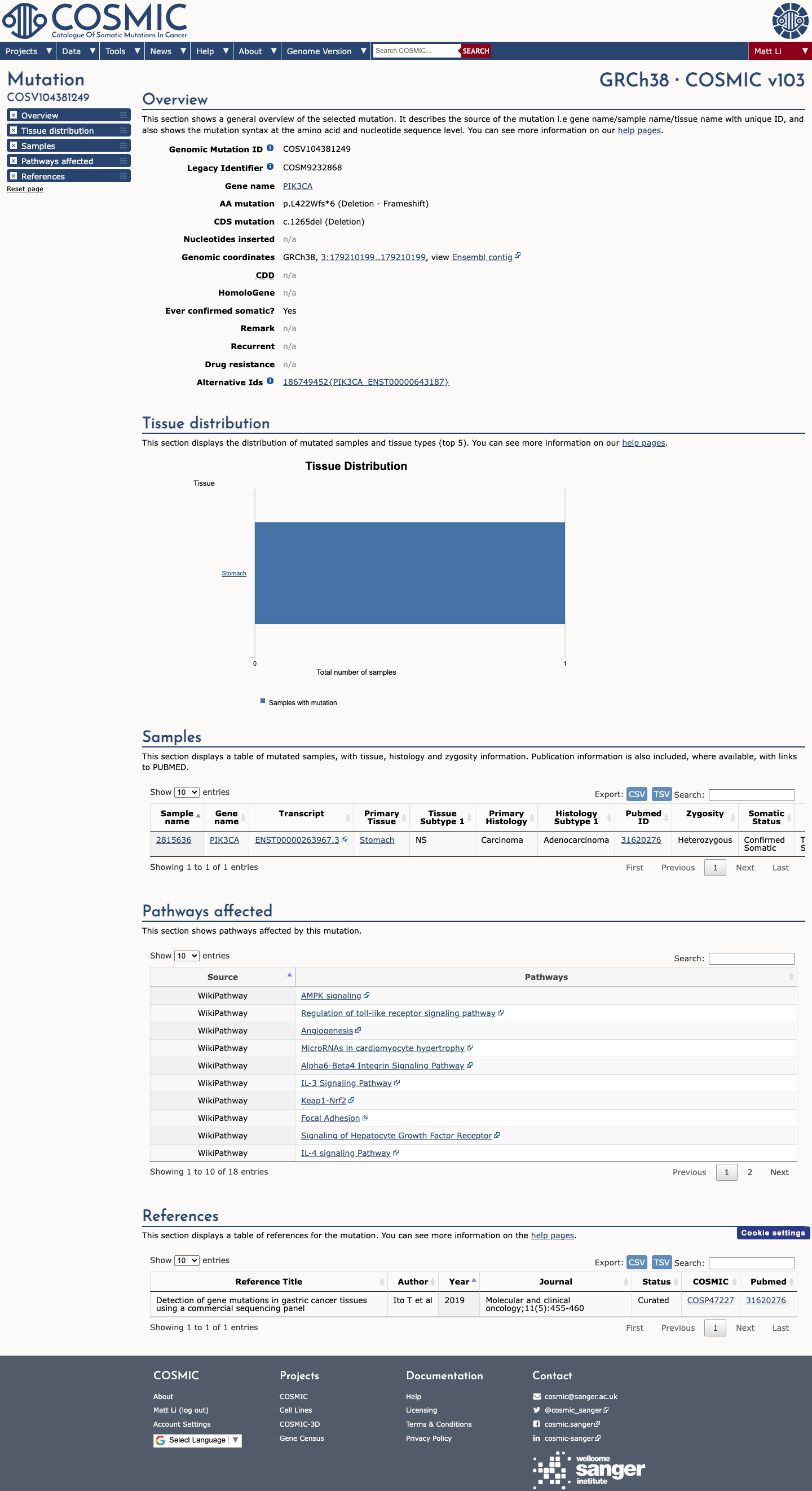
The PIK3CA c.1265del (p.(Leu422TrpfsTer6)) variant has not been reported in ClinVar.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting PM2 at Supporting strength, and its observed population frequency of 0% is below the VCEP BS1 threshold of >0.0185% and BA1 threshold of >0.0926%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
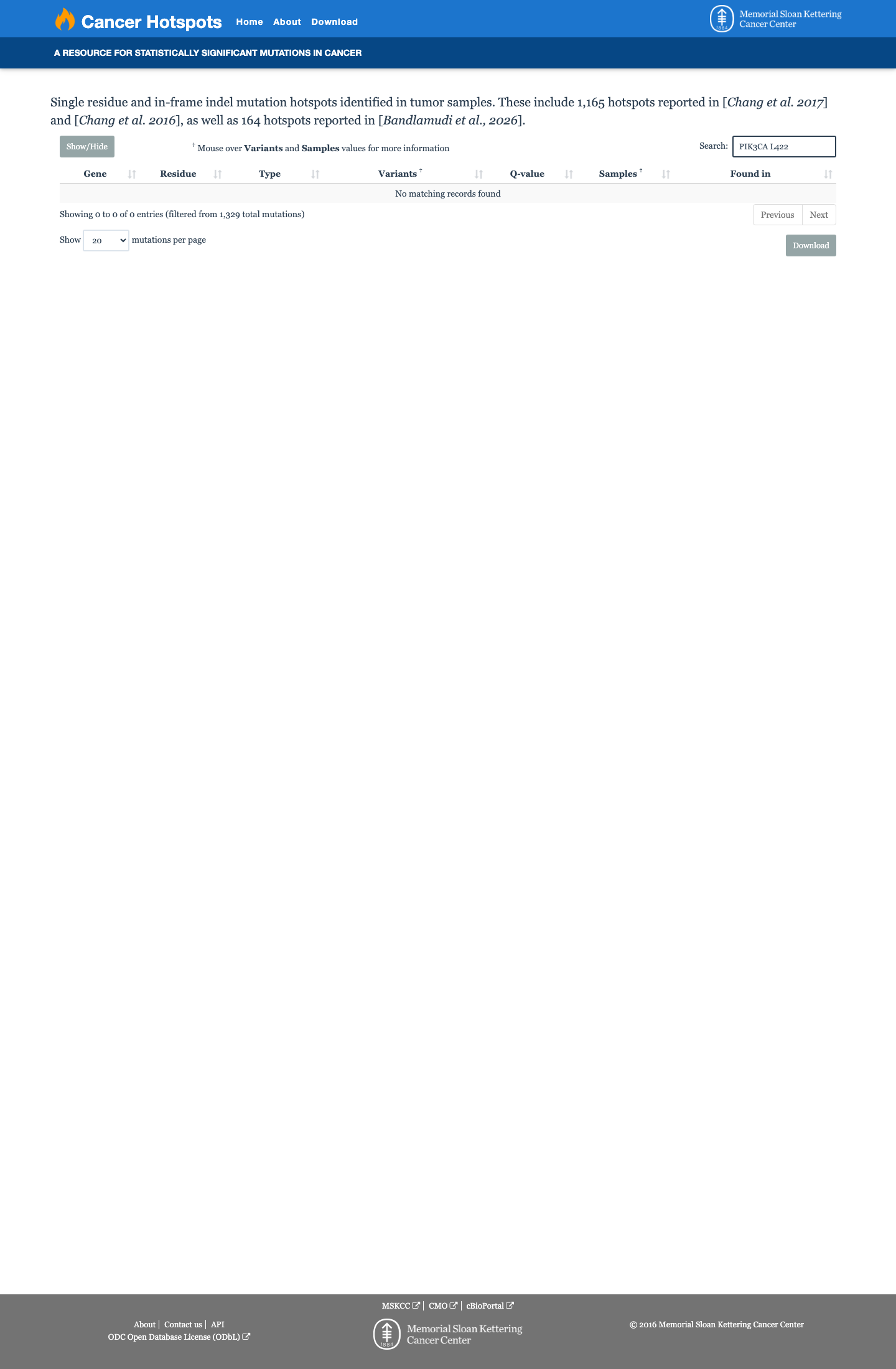
The altered residue lies within the approved PIK3CA kinase-domain interval spanning amino acids 322-483, which is a Brain Malformations VCEP critical functional domain and supports PM1_Supporting.
cspec ↗4
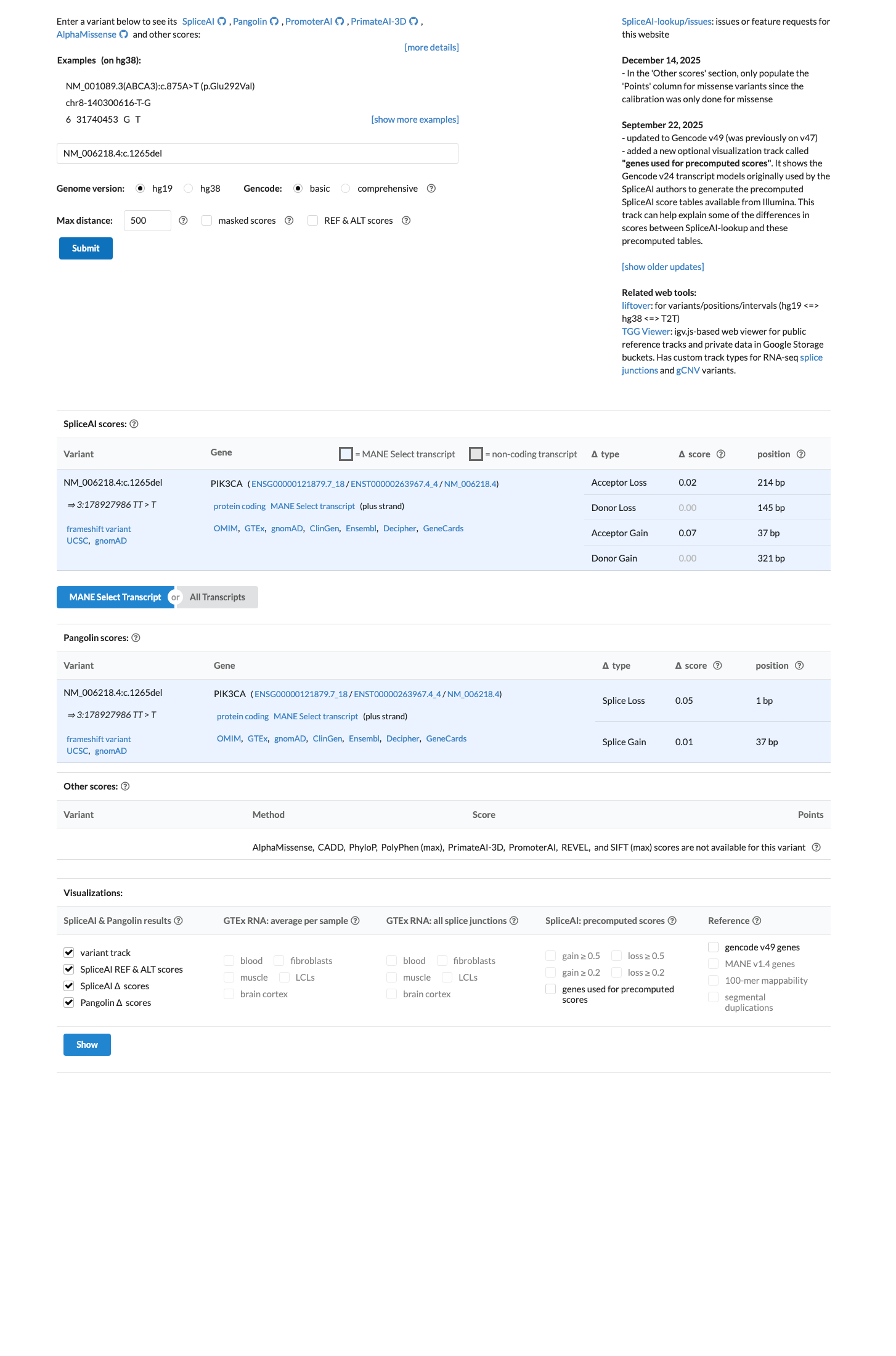
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.07, although computational pathogenicity criteria are not applied for PIK3CA gain-of-function interpretation in this VCEP framework.
spliceai ↗ cspec ↗