Classification rationale
1
2
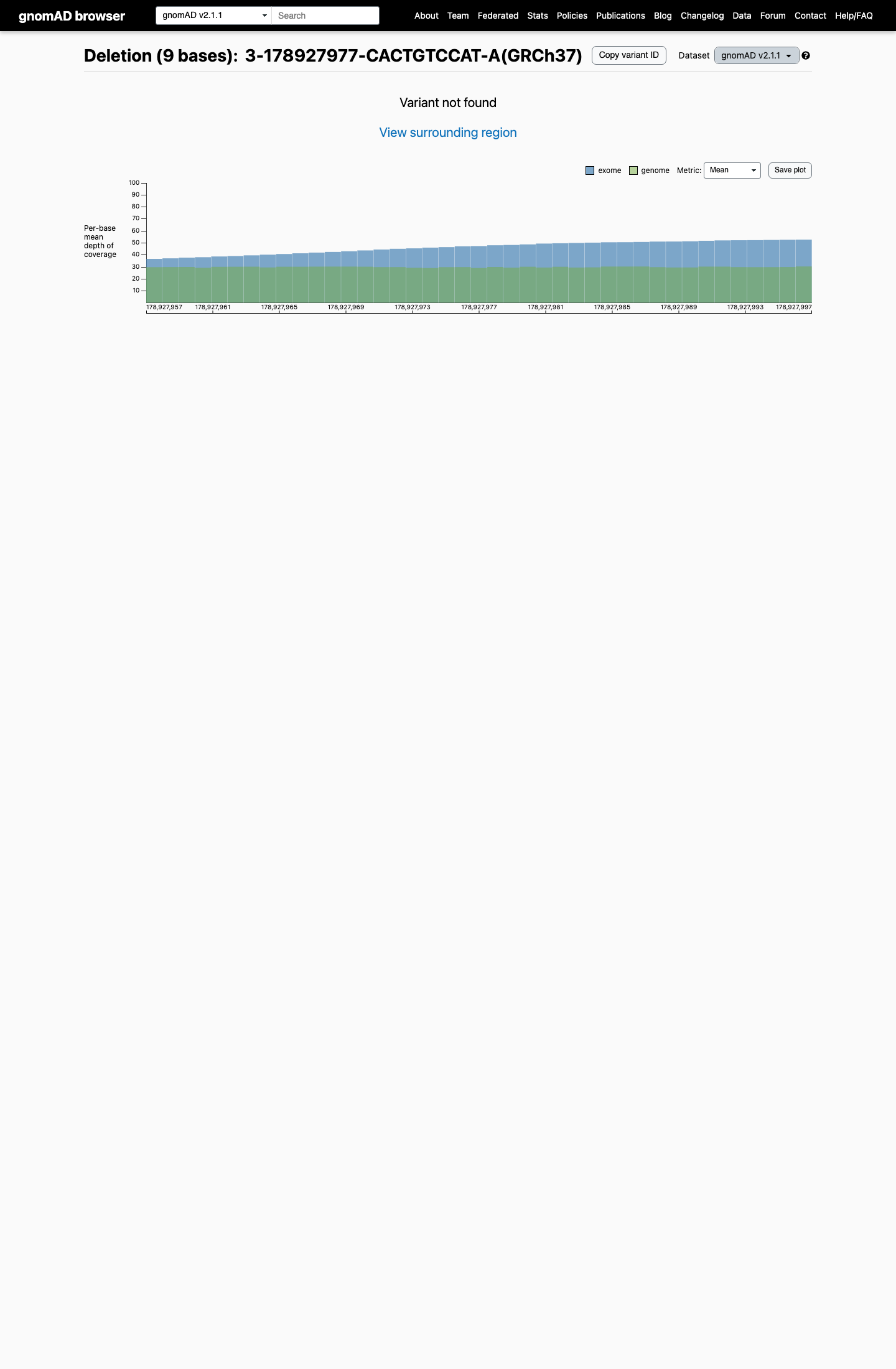
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting that it is absent or extremely rare in population controls.
gnomad_v2 ↗ gnomad_v4 ↗3
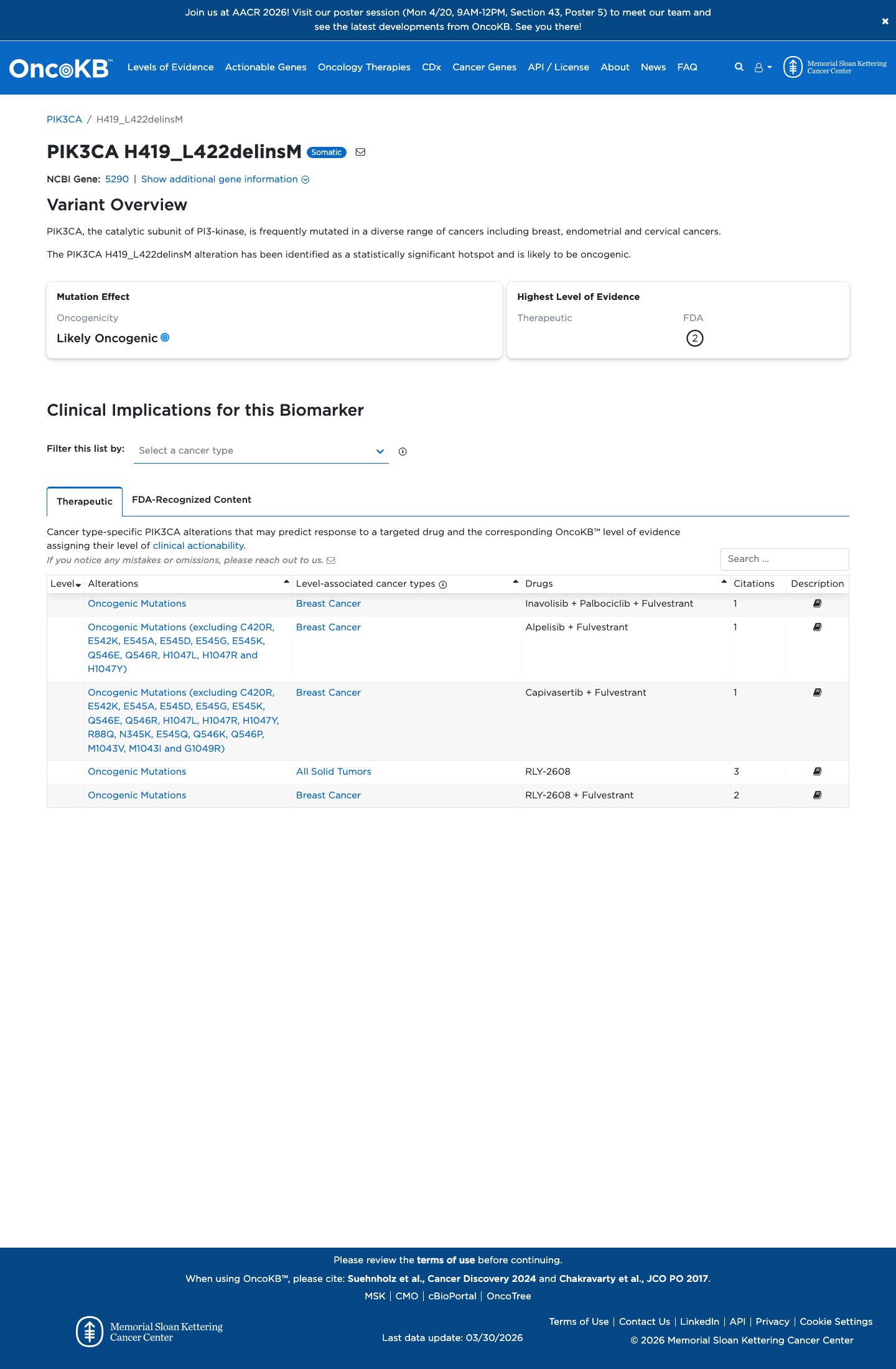
The altered residues fall within the PIK3CA kinase domain spanning amino acids 322 to 483, which is an approved Brain Malformations VCEP critical functional domain and supports PM1 at Supporting strength.
cspec ↗4
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01.
spliceai ↗