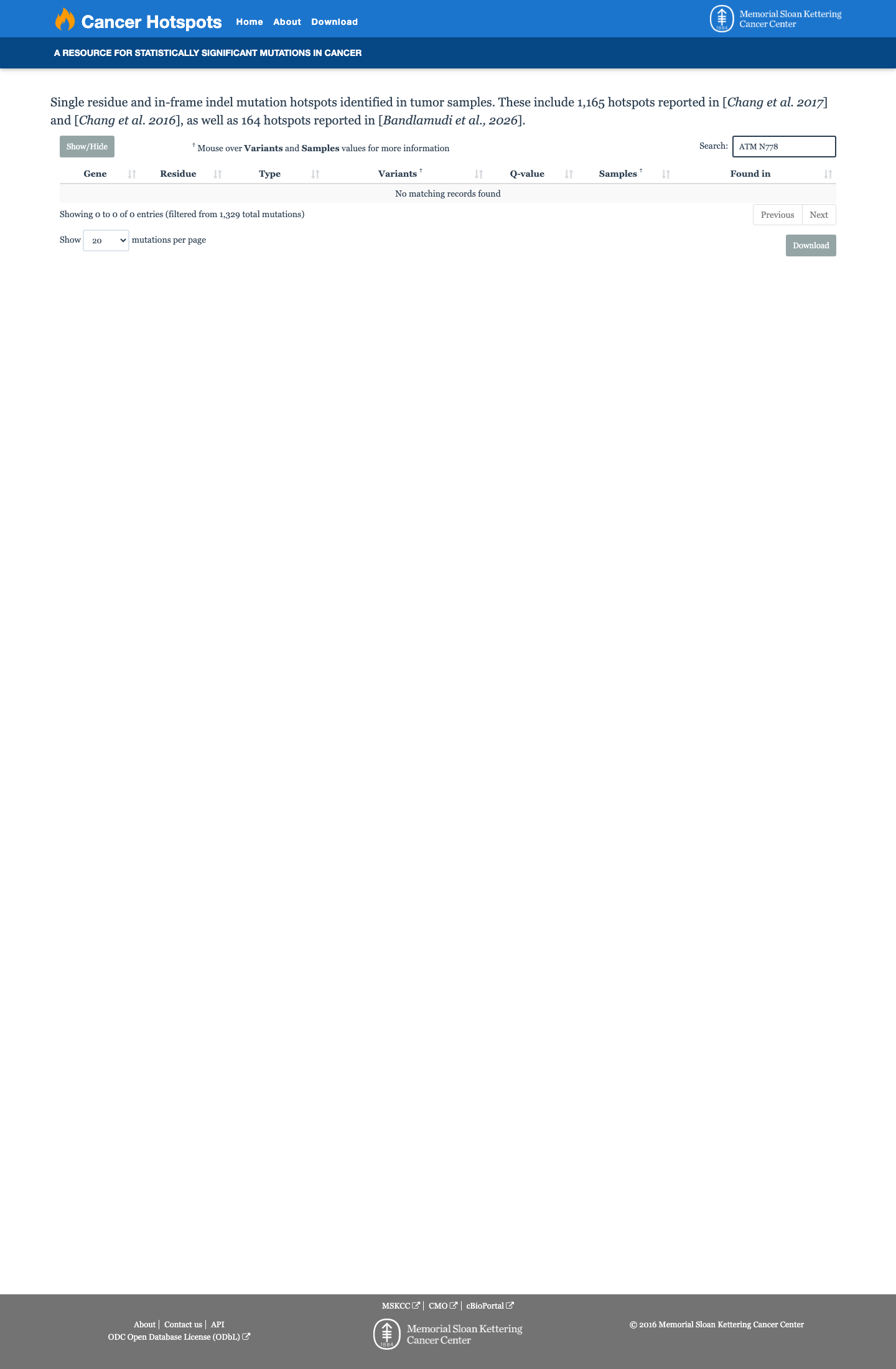
The ATM c.2333A>G (p.Asn778Ser; p.N778S) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar with mixed germline classifications, including 8 submissions as uncertain significance and 2 as likely benign.
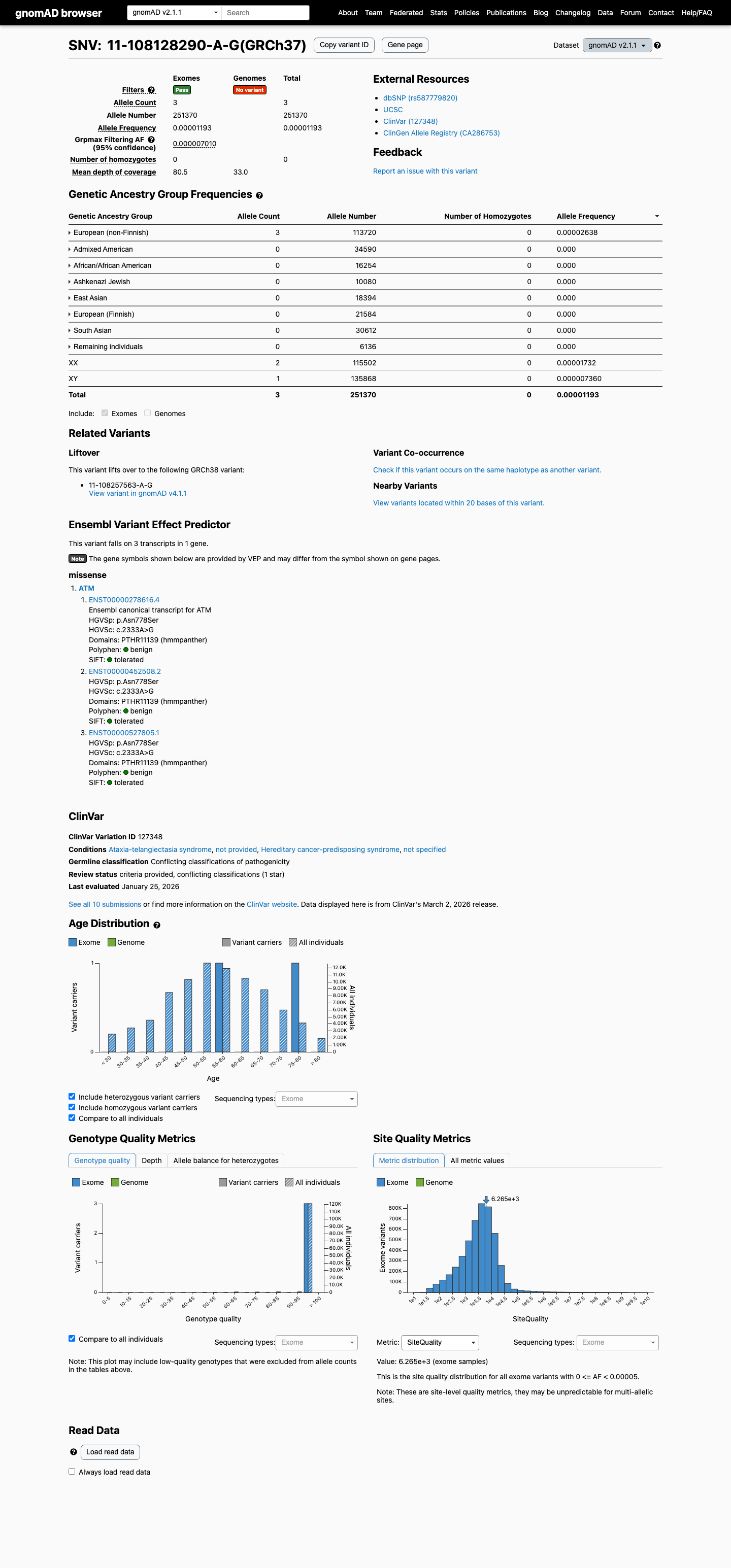
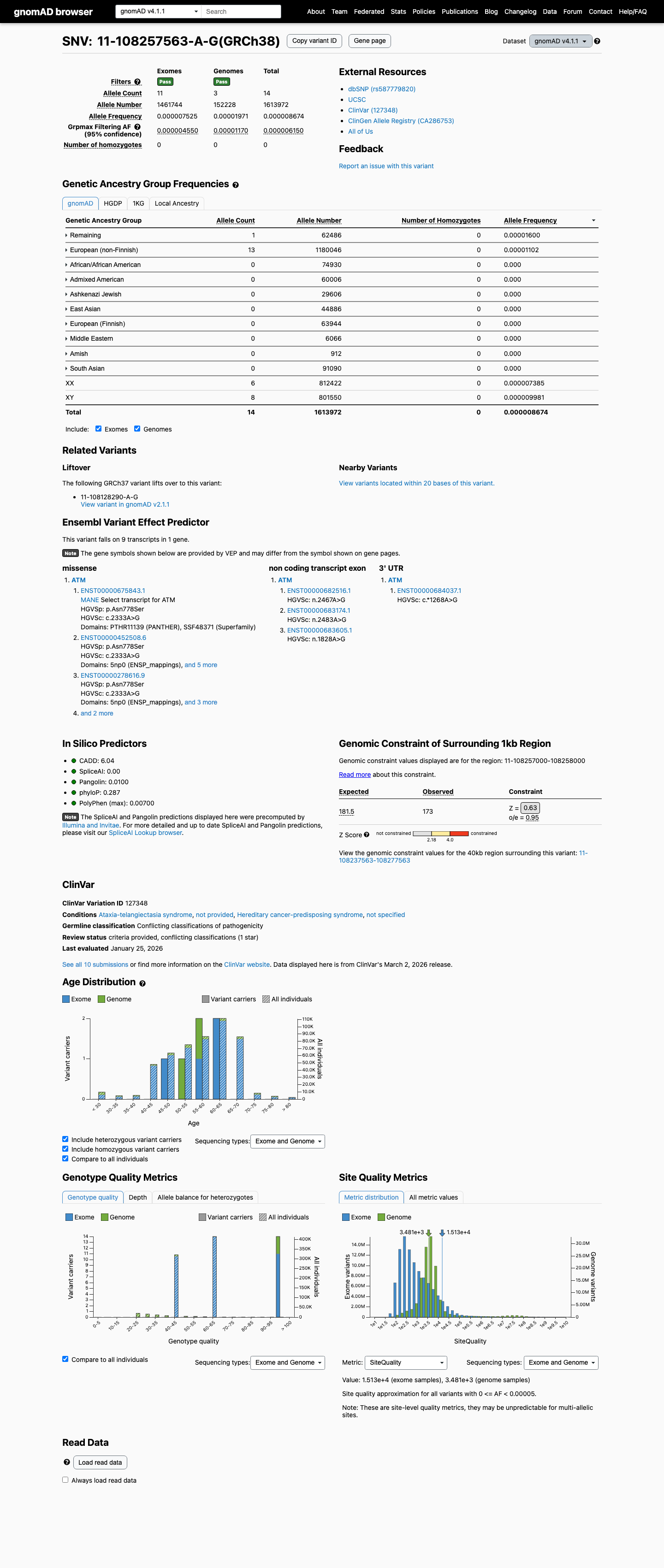
clinvar ↗This variant is present in gnomAD v4.1 at 0.000867% (14/1,613,972 alleles; 0 homozygotes), and this is below the ATM PM2_Supporting threshold of less than or equal to 0.001%.
gnomad_v4 ↗ cspec ↗A high-confidence ATM supplementary functional table classified this variant as functional, but the reviewed ATM-specific materials did not provide a direct mapping of that result to BS3 or PS3 strength.
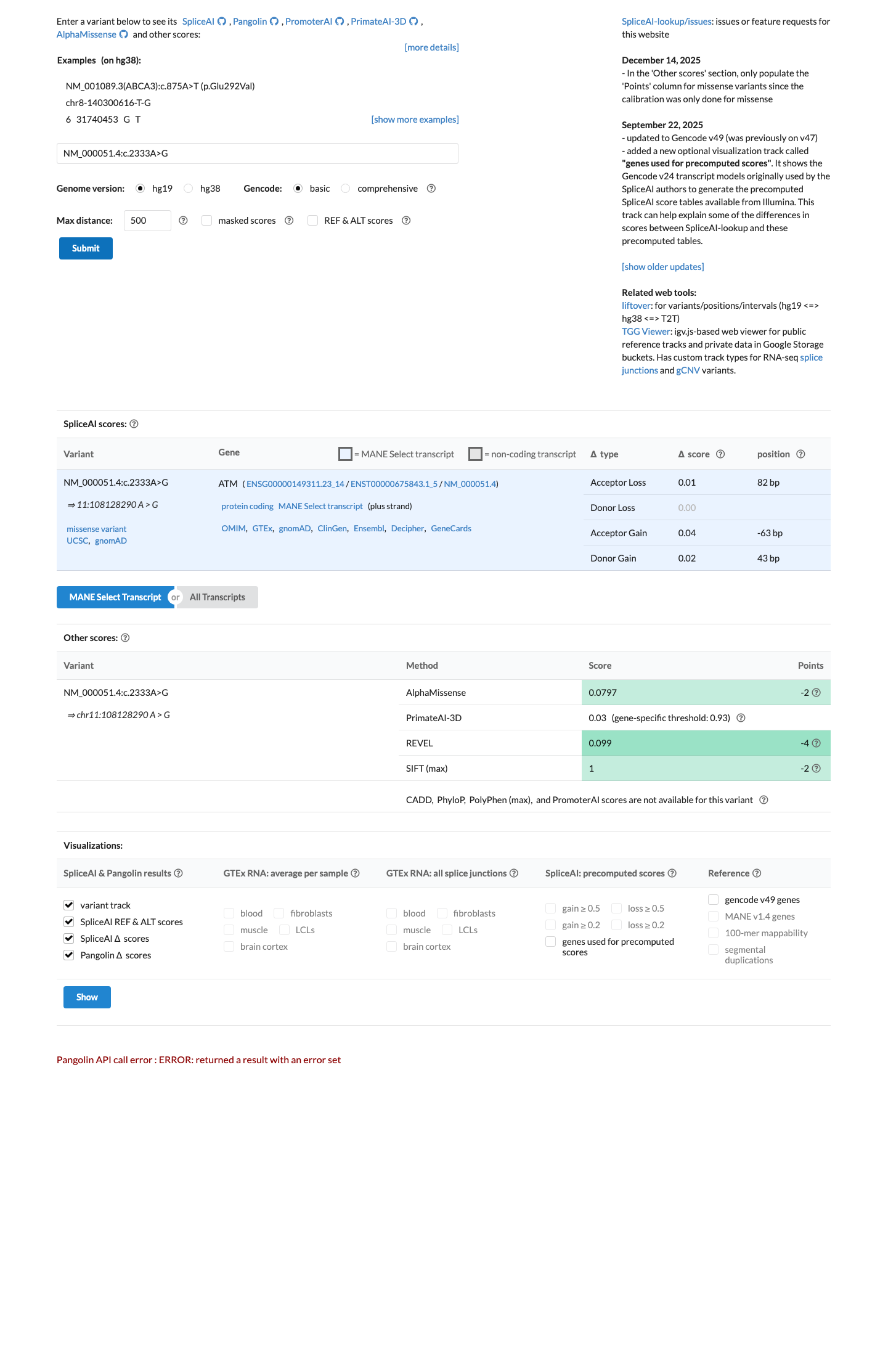
cspec ↗The REVEL score is 0.099, which is below the ATM PP3 threshold of greater than 0.7333 and within the BP4 missense range of less than or equal to 0.249, but no SpliceAI score was returned, so computational evidence was insufficient to apply BP4 and does not support PP3.
spliceai ↗ cspec ↗