Classification rationale
1
2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, corresponding to an observed population frequency of 0, which is below the 0.1% PM2 threshold and supports rarity in the general population.
gnomad_v2 ↗ gnomad_v4 ↗3
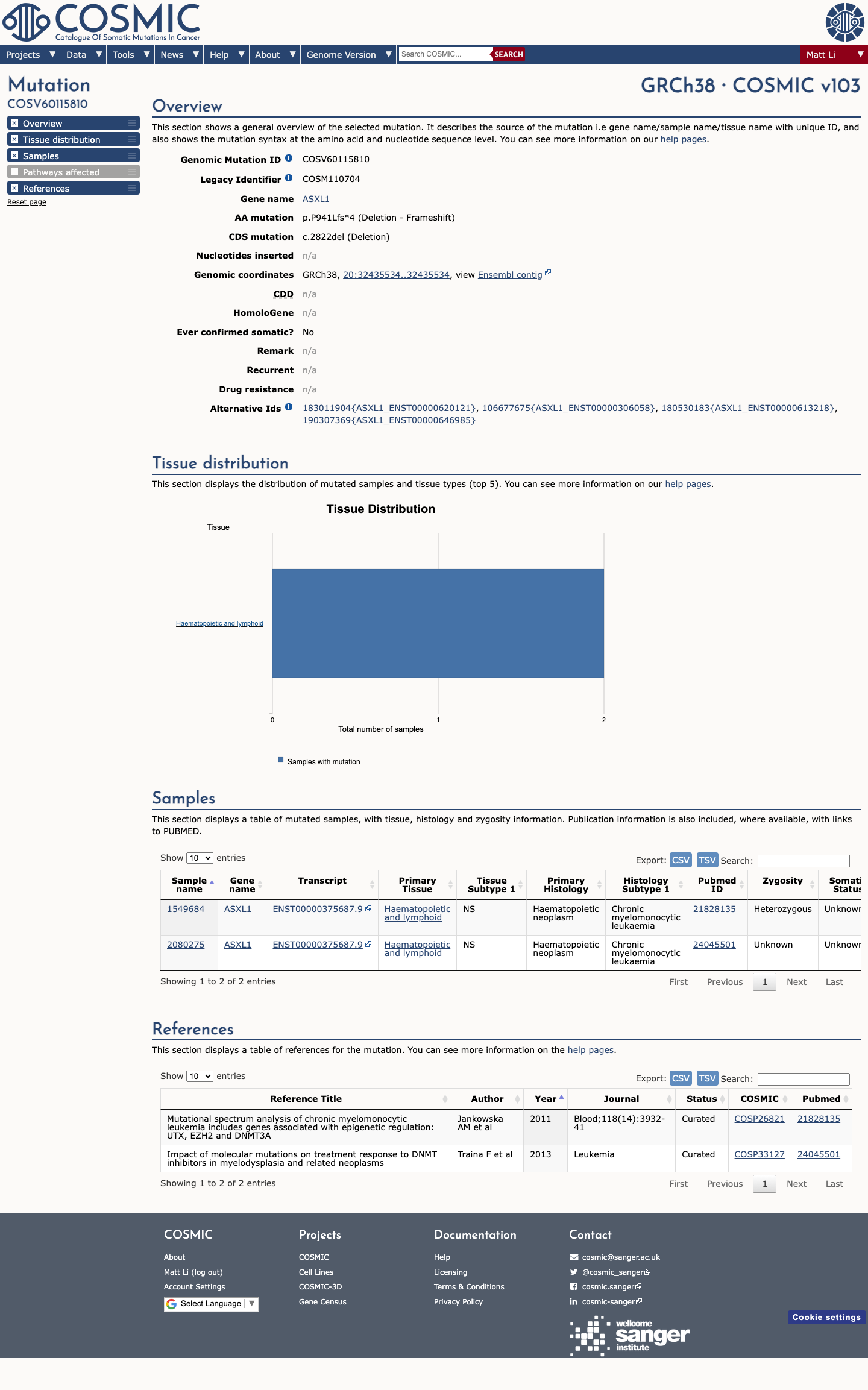
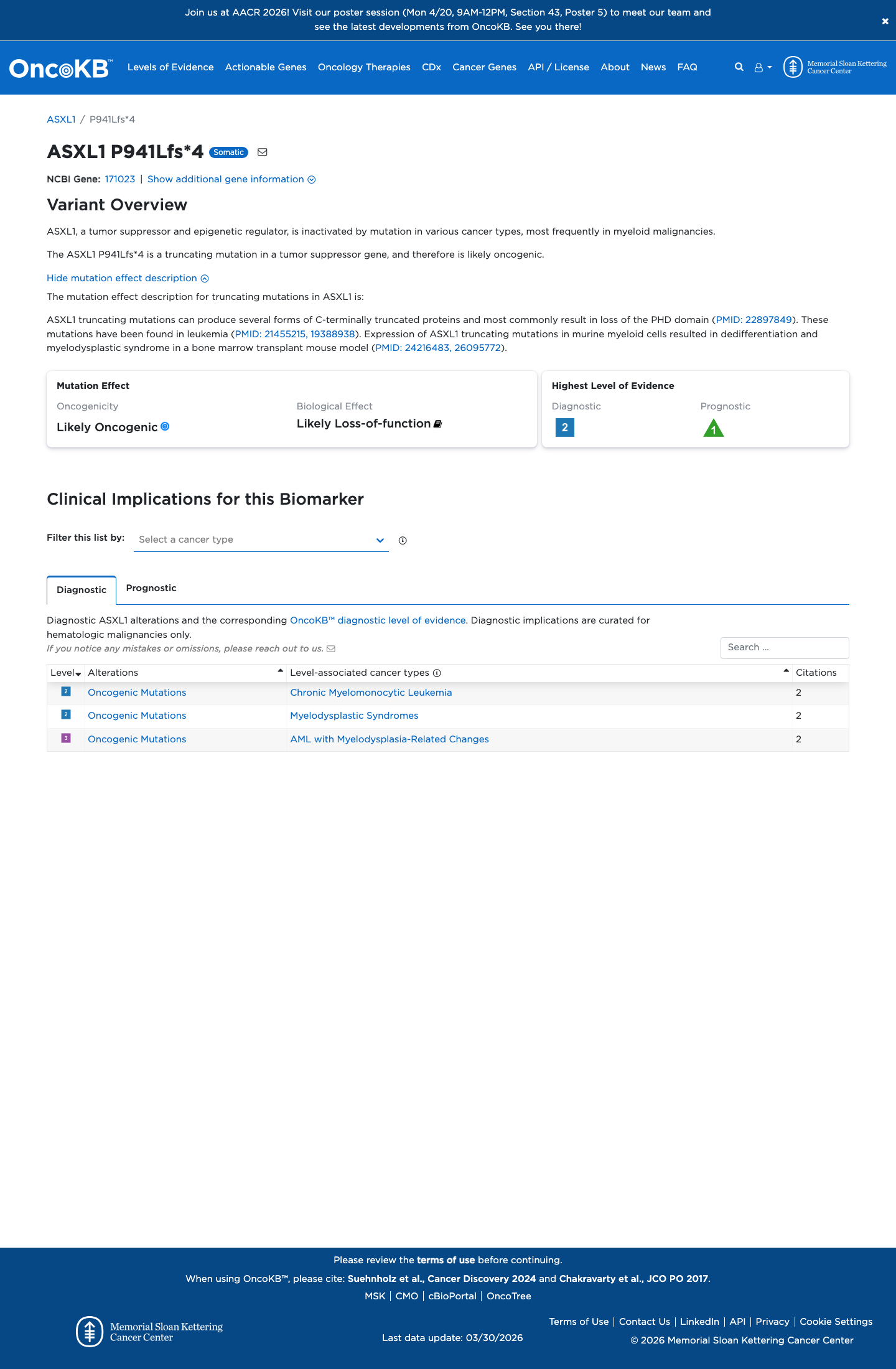
Published studies and OncoKB support damaging loss-of-function biology for ASXL1 truncating variants, but available data do not provide a validated germline functional assay for this specific variant.
oncokb ↗ PMID:19388938 ↗ PMID:21455215 ↗ PMID:22897849 ↗ PMID:24216483 ↗ PMID:26095772 ↗4
This single-base deletion causes a frameshift predicted to truncate the protein after four altered amino acids, removing 598 of 1542 residues (38.8%), and SpliceAI predicts no significant additional splice effect with a maximum delta score of 0.02.
spliceai ↗