Classification rationale
1
The DNMT3A c.2116G>C (p.Gly706Arg; p.G706R) variant has not been reported in ClinVar.
clinvar ↗2
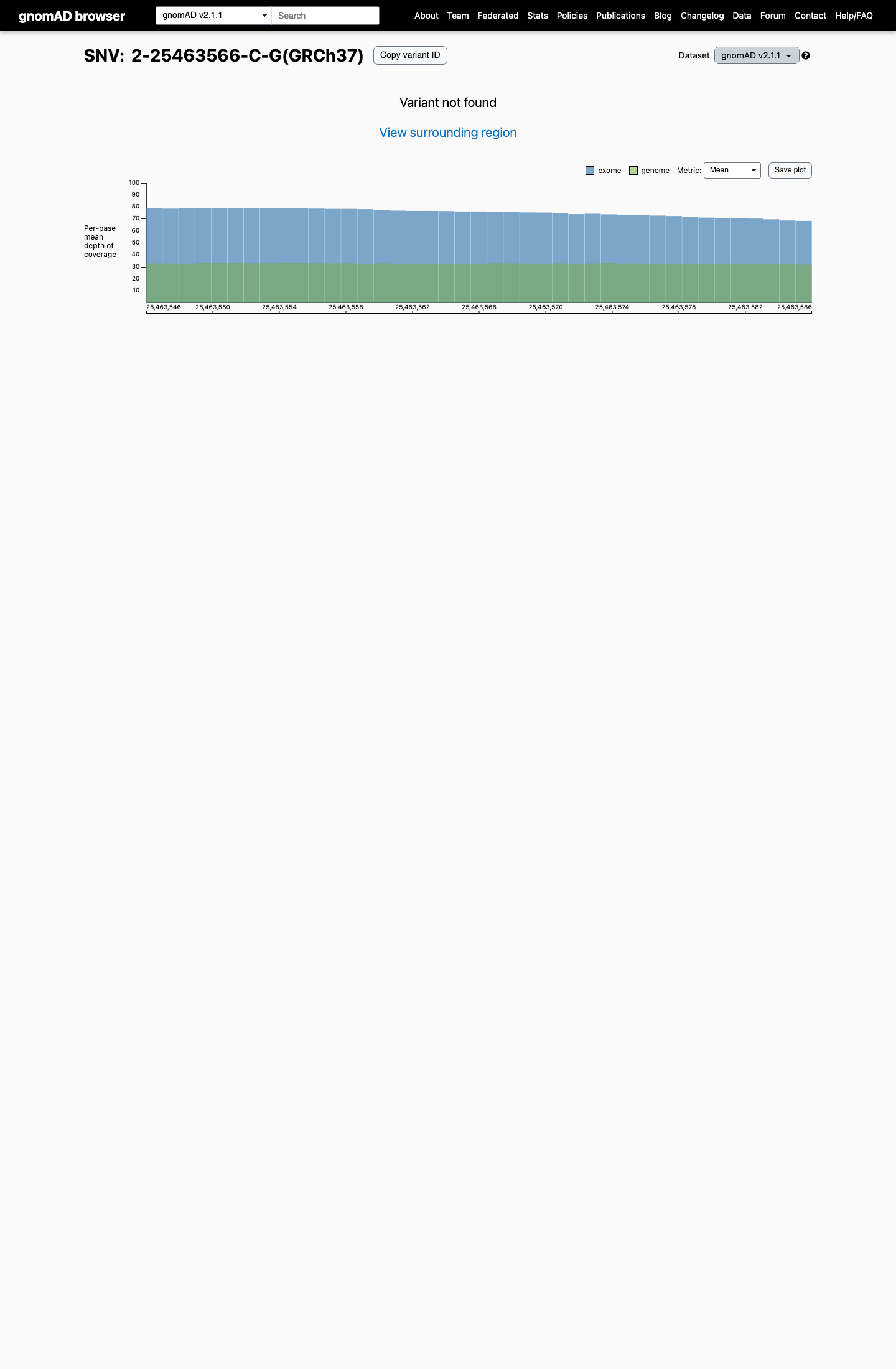
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in the general population.
gnomad_v2 ↗ gnomad_v4 ↗3
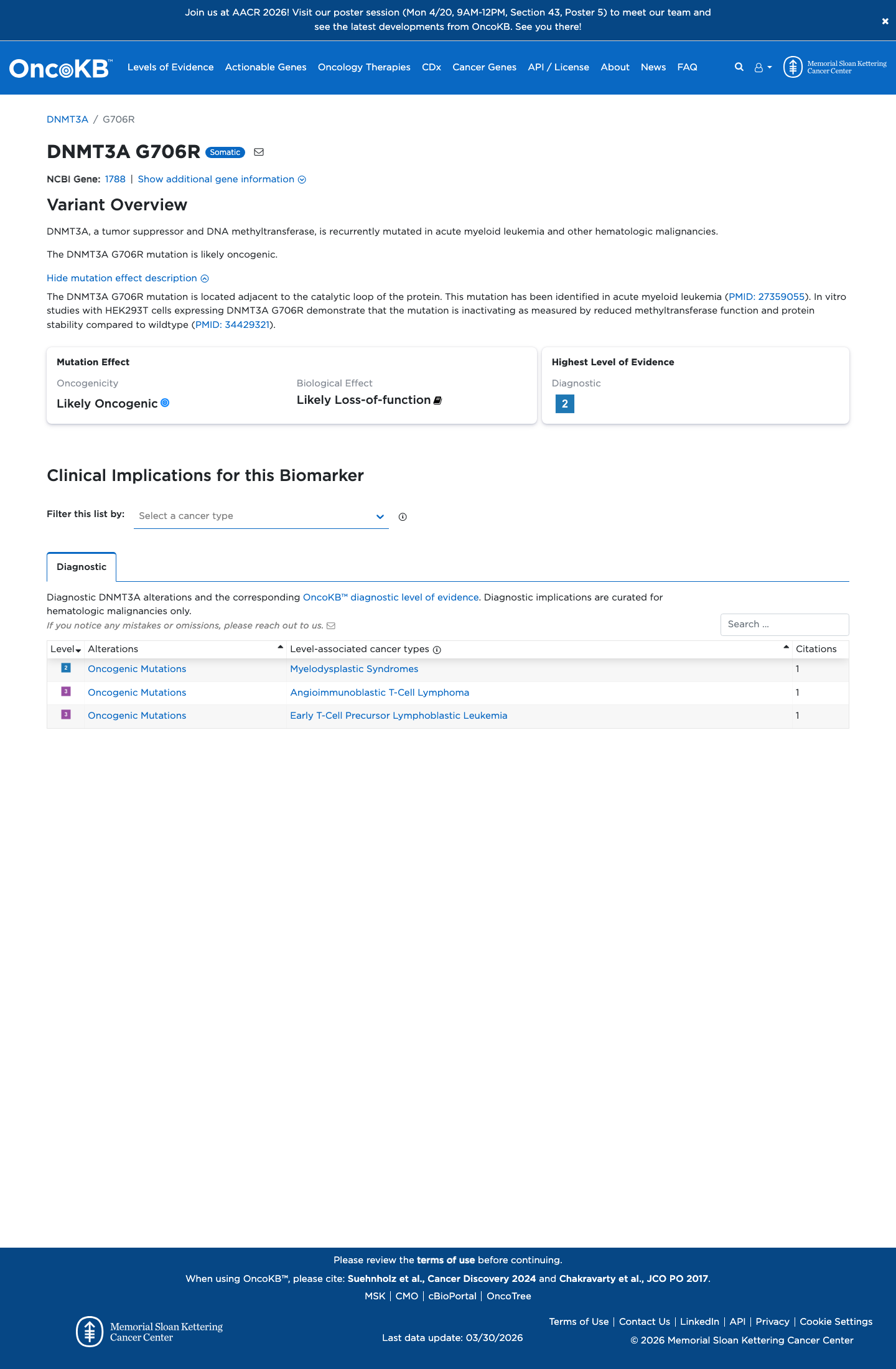
OncoKB classifies this variant as Likely Oncogenic with a likely loss-of-function biological effect, but the available evidence does not provide a well-established variant-specific functional study result sufficient for ACMG PS3 or BS3.
oncokb ↗ PMID:34429321 ↗4
Computational evidence supports a deleterious protein effect, with REVEL 0.971, while SpliceAI predicts no significant splice impact with a maximum delta score of 0.04.
spliceai ↗