Classification rationale
1
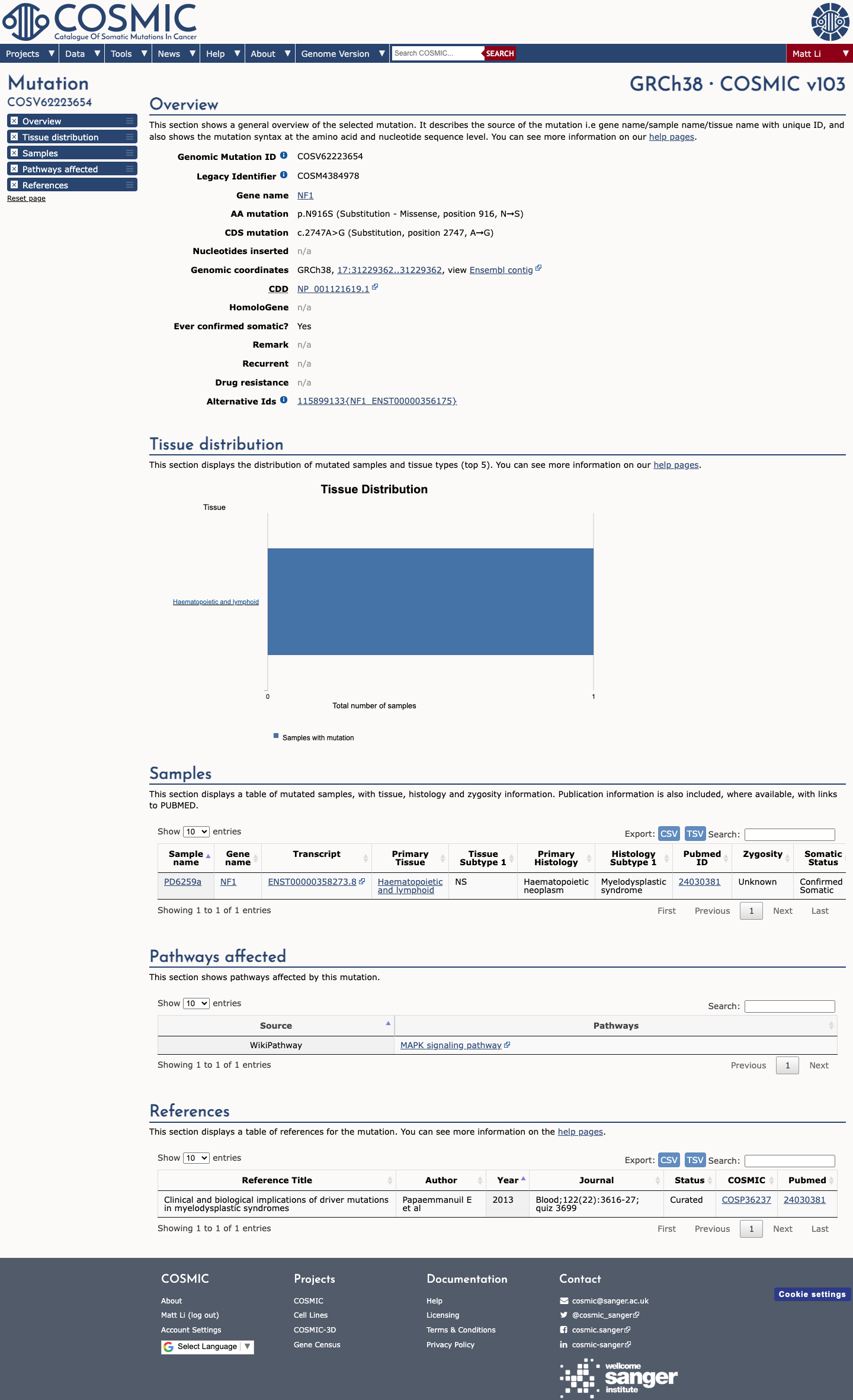
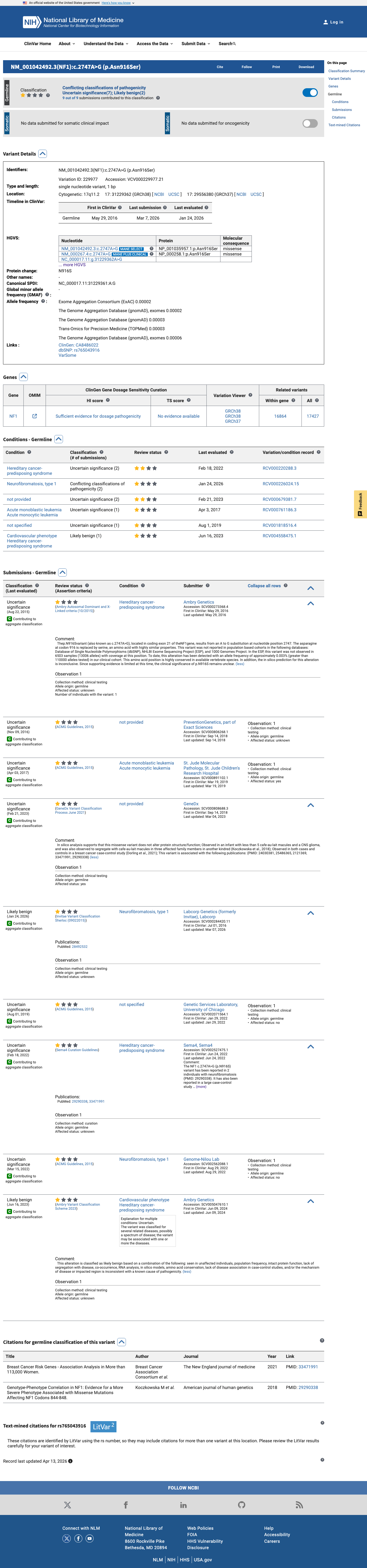
The NF1 c.2747A>G (p.Asn916Ser) variant has been reported in ClinVar with conflicting germline classifications, including uncertain significance and likely benign, and without an expert panel assertion.
clinvar ↗2
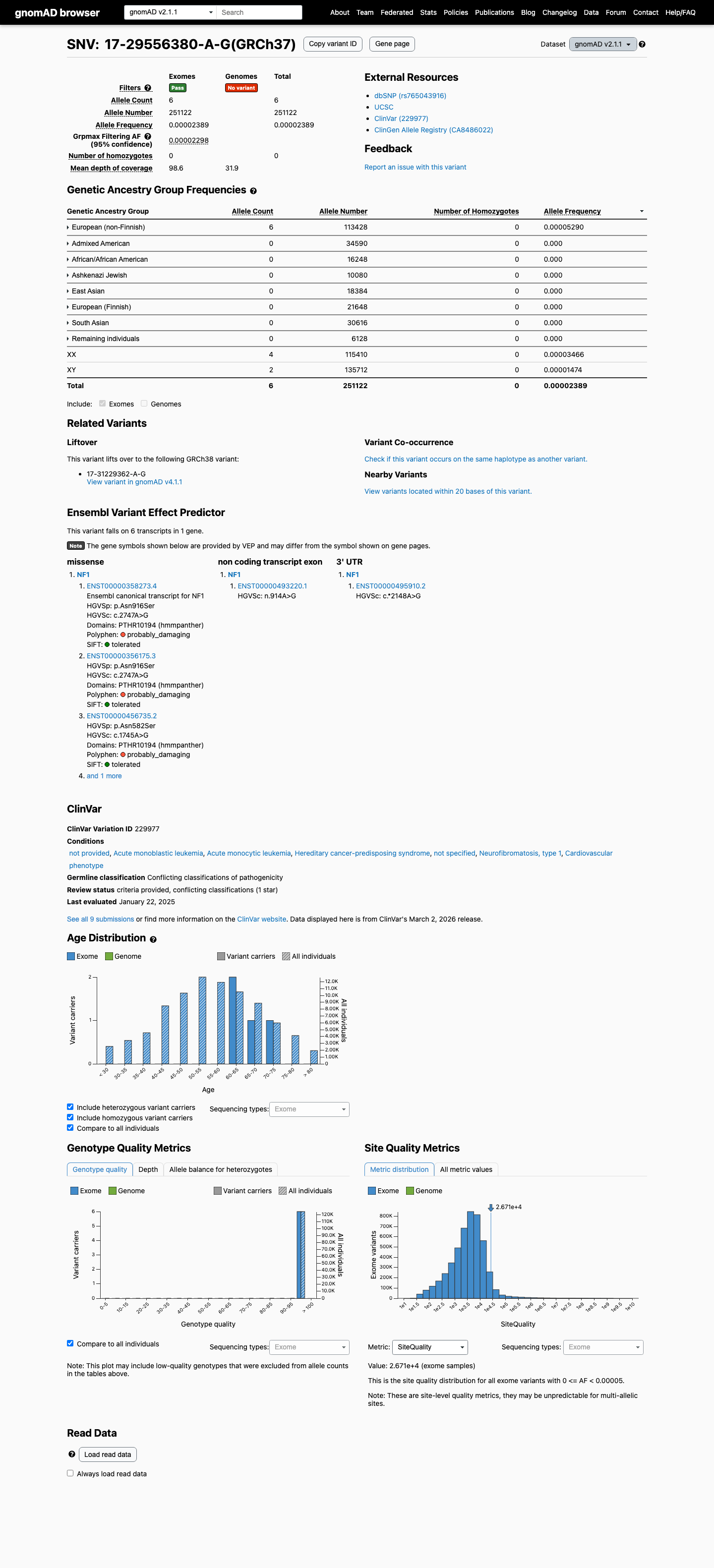
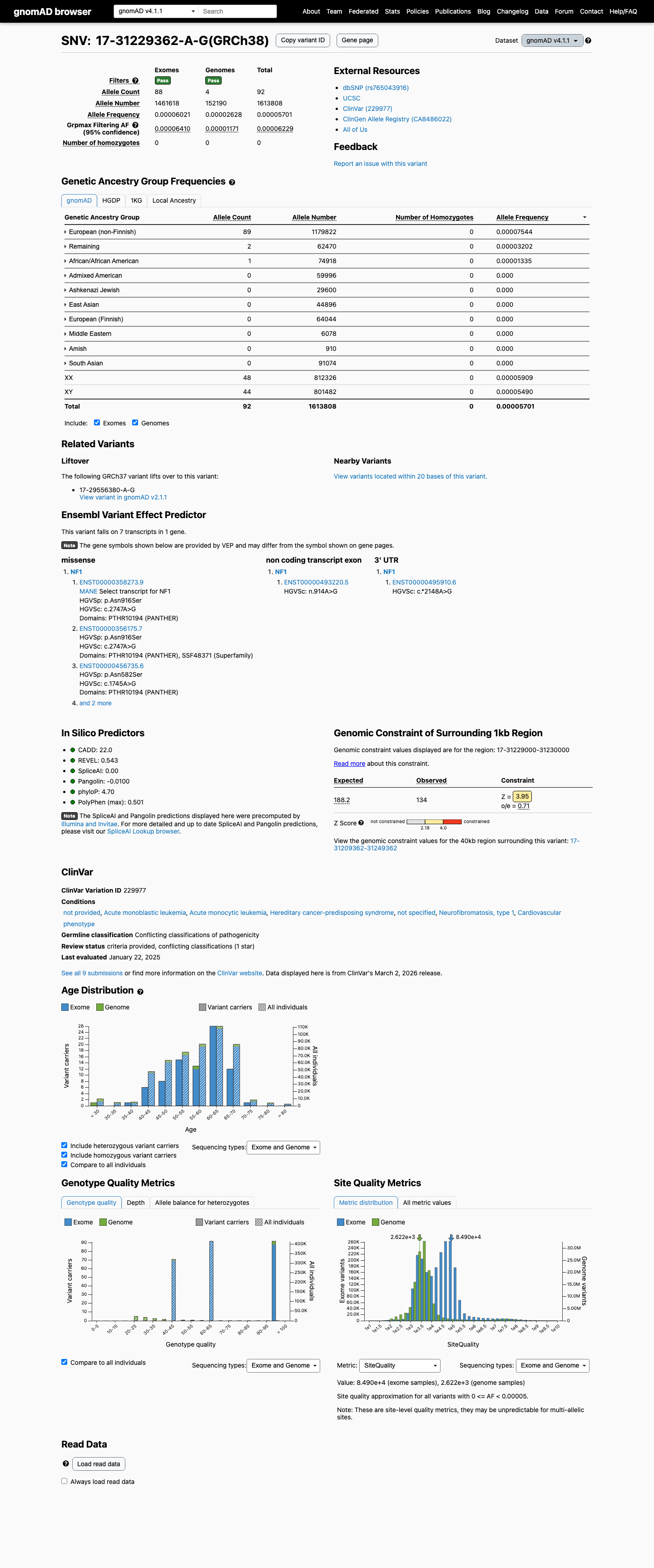
This variant is present at low frequency in population databases: gnomAD v4.1 shows an allele frequency of 0.00570% (92/1613808 alleles; grpmax FAF 0.006229%) and gnomAD v2.1 shows an allele frequency of 0.00239% (6/251122 alleles), which is below benign frequency thresholds but means the variant is not absent from controls.
gnomad_v2 ↗ gnomad_v4 ↗3
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01, so in silico splicing evidence does not independently support pathogenicity.
spliceai ↗