Classification rationale
1
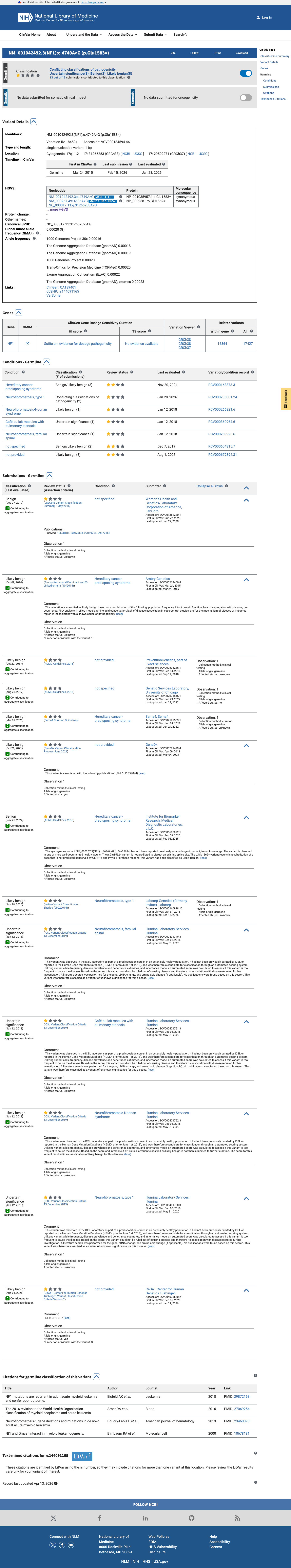
The NF1 c.4686A>G (p.Glu1562=) variant has been reported in ClinVar predominantly as likely benign or benign, with some uncertain significance submissions.
clinvar ↗2
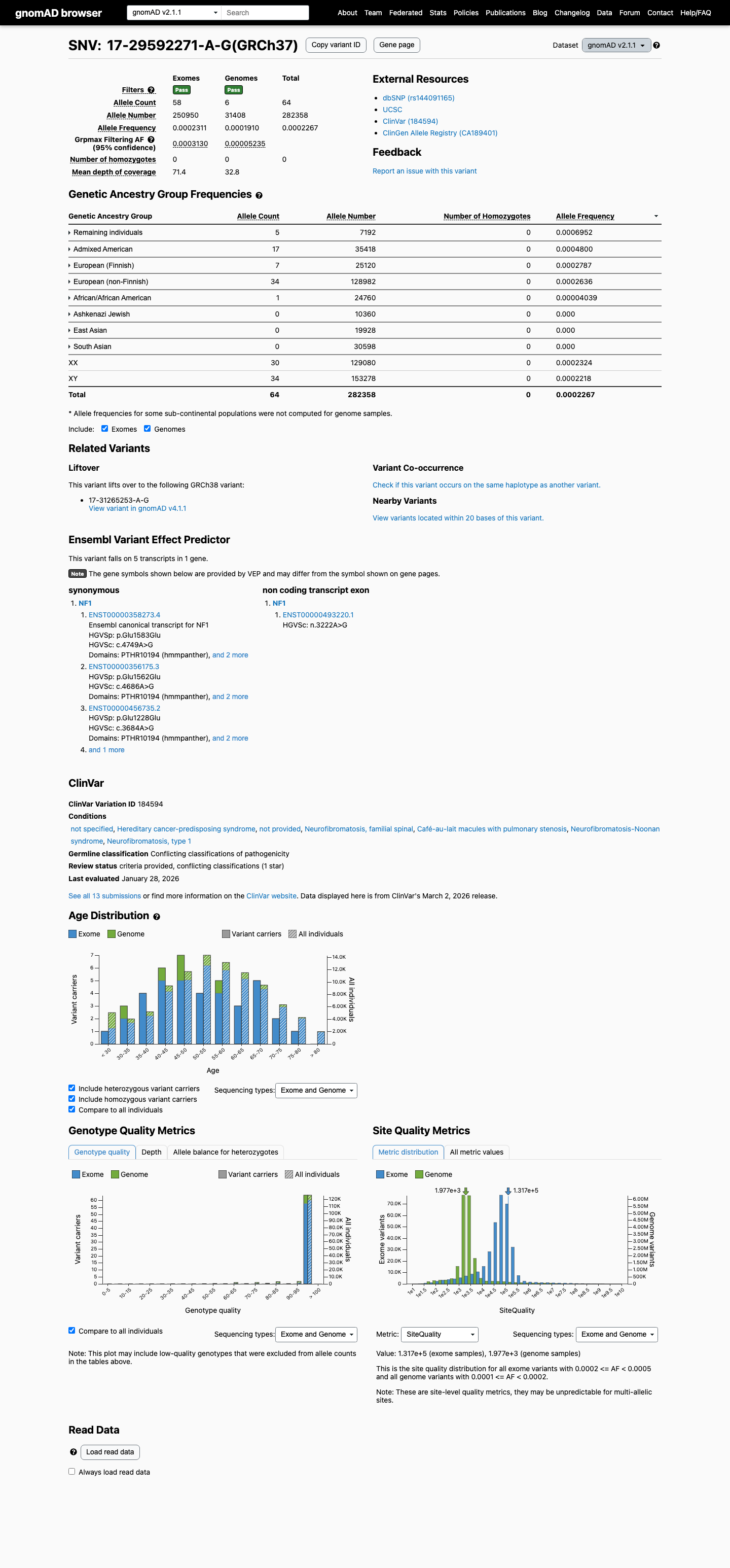
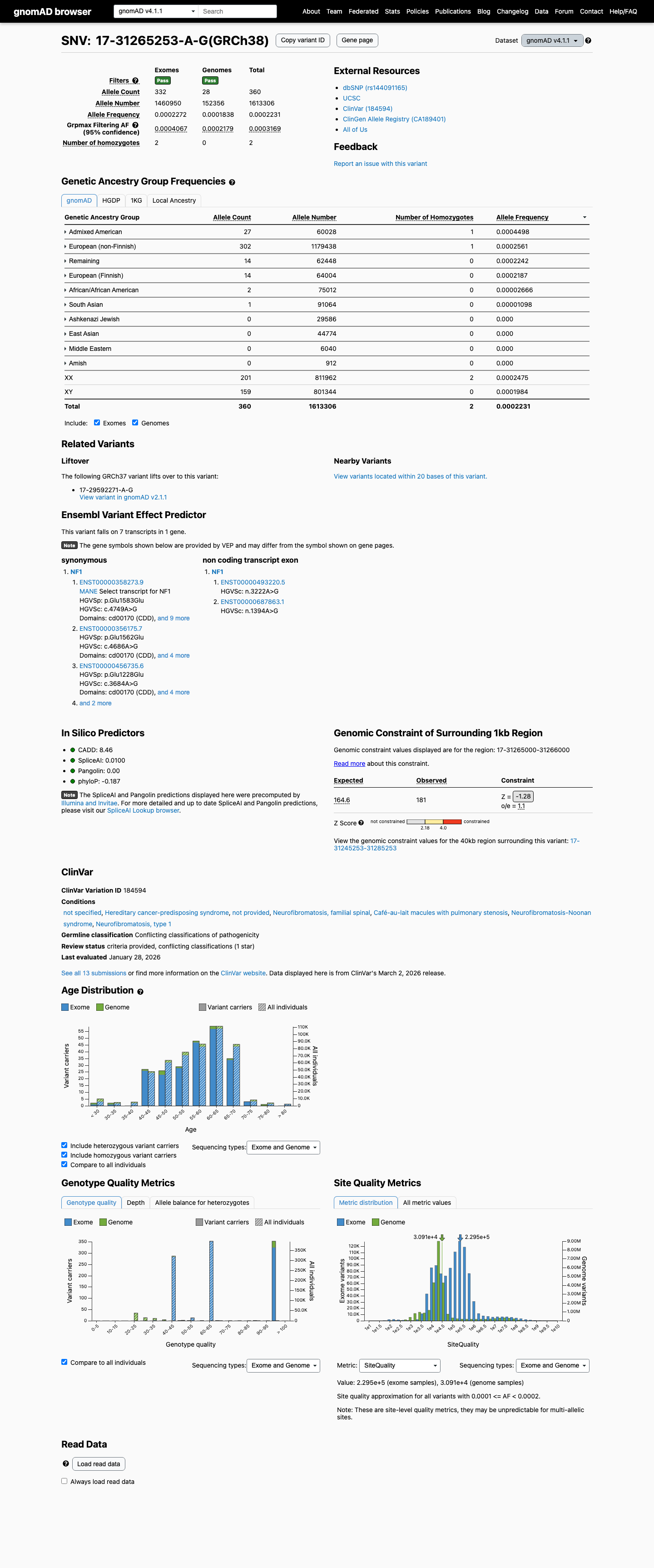
This variant is present in population databases, including gnomAD v2.1 at an allele frequency of 0.000226663 (64/282358) and gnomAD v4.1 at an allele frequency of 0.000223144, which is below common benign strong thresholds but inconsistent with absence from controls.
gnomad_v2 ↗ gnomad_v4 ↗3
In silico splice prediction does not support a splice-disrupting effect, with SpliceAI showing a maximum delta score of 0.01.
spliceai ↗